MONOLIXSUITE 2020 Demonstration Monolix Suite 2020 R 1












![Monolix – writing models [LONGITUDINAL] input = {Cl, V 1, Q 2, V 2, Monolix – writing models [LONGITUDINAL] input = {Cl, V 1, Q 2, V 2,](https://slidetodoc.com/presentation_image_h/04ad9c5627924131e8bd8cd785280fb0/image-34.jpg)



![Mlxtran: using macros iv V k 12 k 21 k [LONGITUDINAL] input = {V, Mlxtran: using macros iv V k 12 k 21 k [LONGITUDINAL] input = {V,](https://slidetodoc.com/presentation_image_h/04ad9c5627924131e8bd8cd785280fb0/image-40.jpg)

![Mlxtran: using ODEs [LONGITUDINAL] input = {V, k, k 12, k 21} PK: depot(adm=1, Mlxtran: using ODEs [LONGITUDINAL] input = {V, k, k 12, k 21} PK: depot(adm=1,](https://slidetodoc.com/presentation_image_h/04ad9c5627924131e8bd8cd785280fb0/image-42.jpg)





- Slides: 52

MONOLIXSUITE 2020 Demonstration

Monolix. Suite 2020 R 1 2

Case study: Remifentanil PK-PD Remifentanil - analgesic drug used for sedation and to relieve pain during surgery Dataset: q 65 healthy adults q constant infusion rate between 1 and 8 µg. kg-1. min-1 for 4 to 20 minutes q Measurements: • PK: concentration measurements during and after infusion • PD: dense electroencephalogram (EEG) measurements (depth of sedation) q Covariates: AGE, SEX, TINFCAT 2020 R 1 Monolix. Suite 3

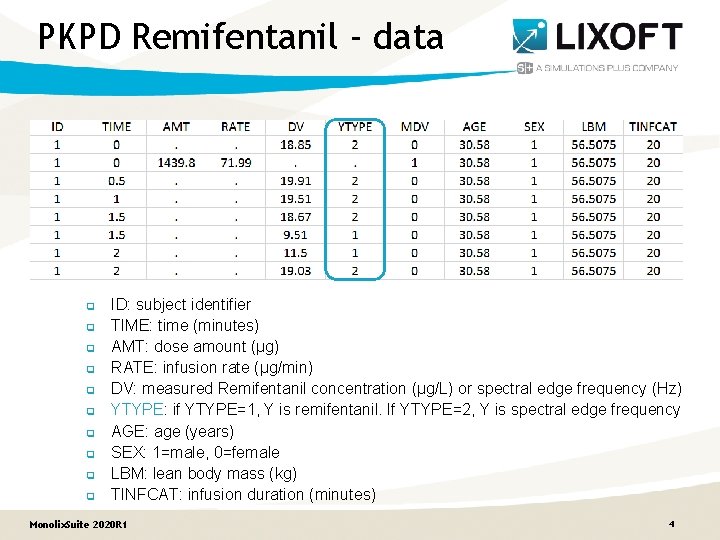
PKPD Remifentanil - data q q q q q ID: subject identifier TIME: time (minutes) AMT: dose amount (µg) RATE: infusion rate (µg/min) DV: measured Remifentanil concentration (µg/L) or spectral edge frequency (Hz) YTYPE: if YTYPE=1, Y is remifentanil. If YTYPE=2, Y is spectral edge frequency AGE: age (years) SEX: 1=male, 0=female LBM: lean body mass (kg) TINFCAT: infusion duration (minutes) Monolix. Suite 2020 R 1 4

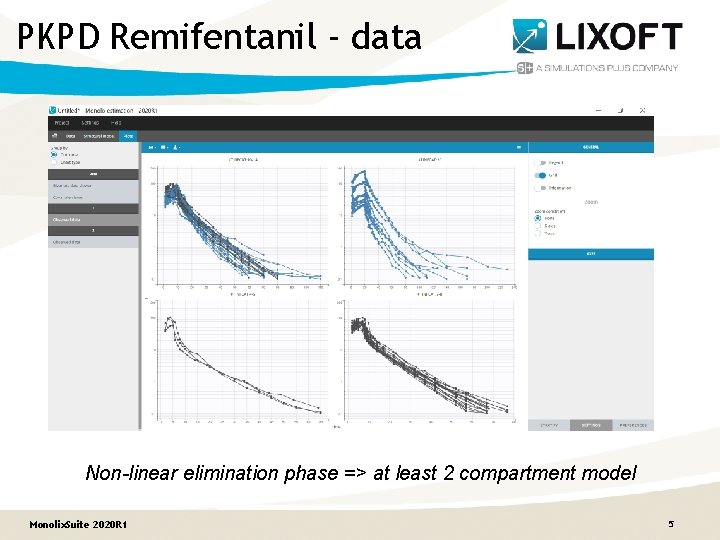
PKPD Remifentanil - data Non-linear elimination phase => at least 2 compartment model Monolix. Suite 2020 R 1 5

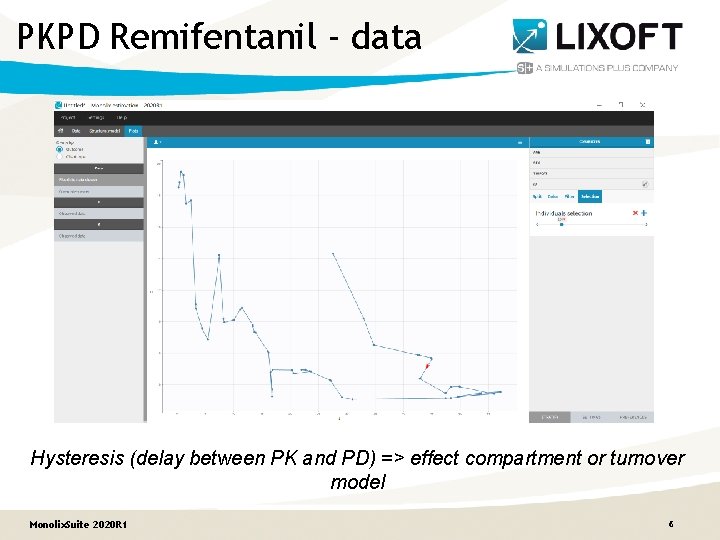
PKPD Remifentanil - data Hysteresis (delay between PK and PD) => effect compartment or turnover model Monolix. Suite 2020 R 1 6

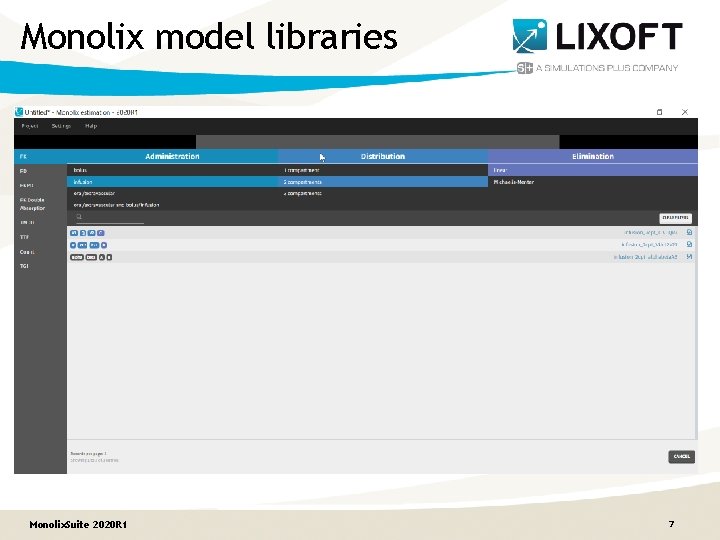
Monolix model libraries Monolix. Suite 2020 R 1 7

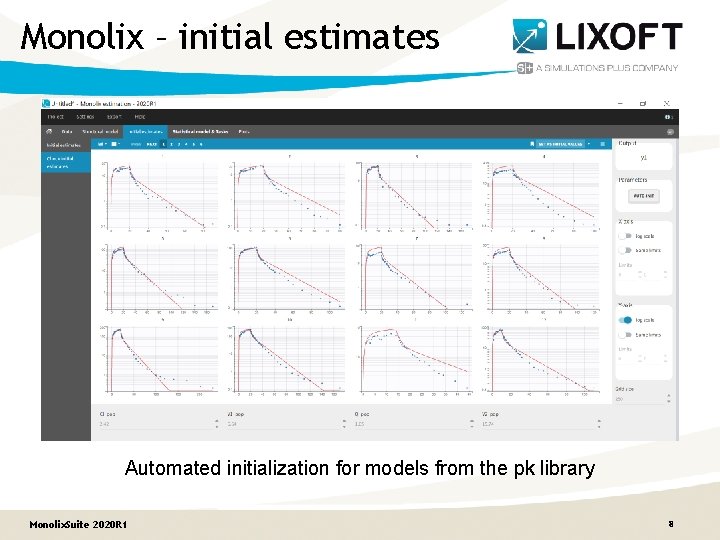
Monolix – initial estimates Automated initialization for models from the pk library Monolix. Suite 2020 R 1 8

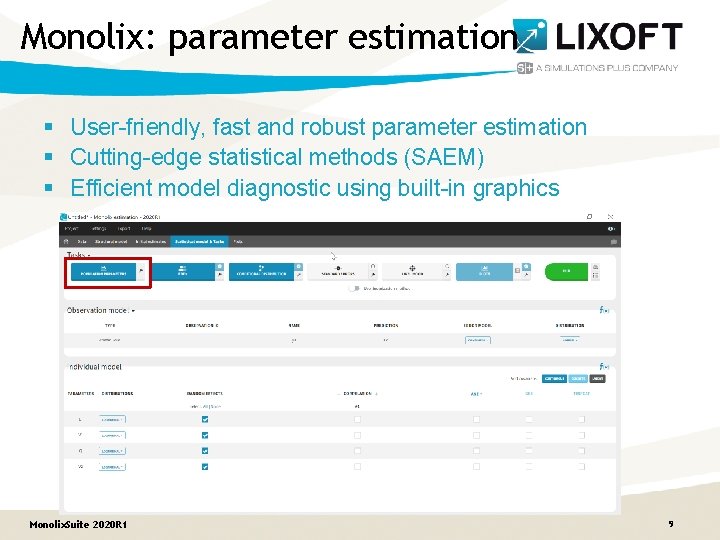
Monolix: parameter estimation § User-friendly, fast and robust parameter estimation § Cutting-edge statistical methods (SAEM) § Efficient model diagnostic using built-in graphics Monolix. Suite 2020 R 1 9

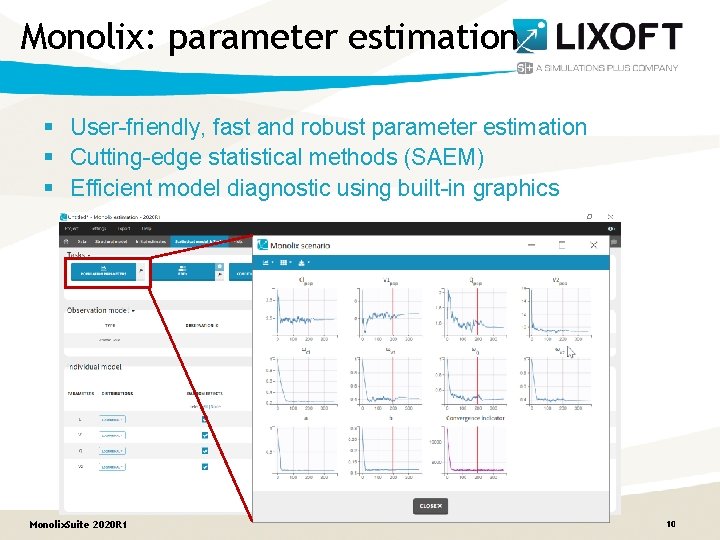
Monolix: parameter estimation § User-friendly, fast and robust parameter estimation § Cutting-edge statistical methods (SAEM) § Efficient model diagnostic using built-in graphics Monolix. Suite 2020 R 1 10

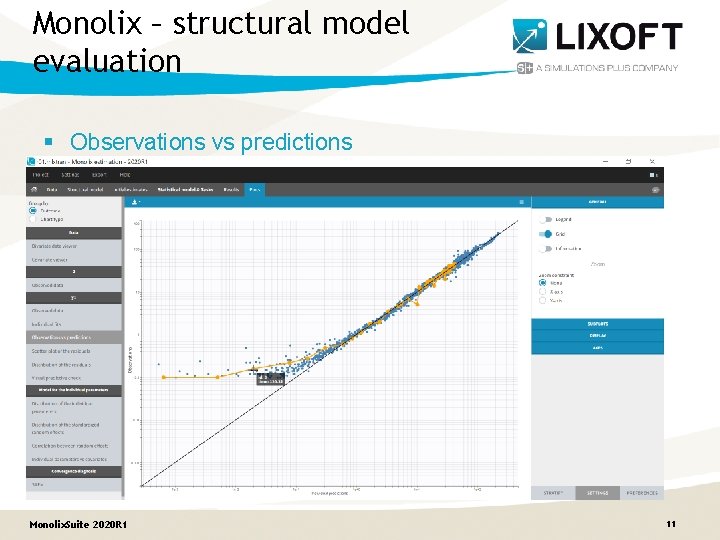
Monolix – structural model evaluation § Observations vs predictions Monolix. Suite 2020 R 1 11

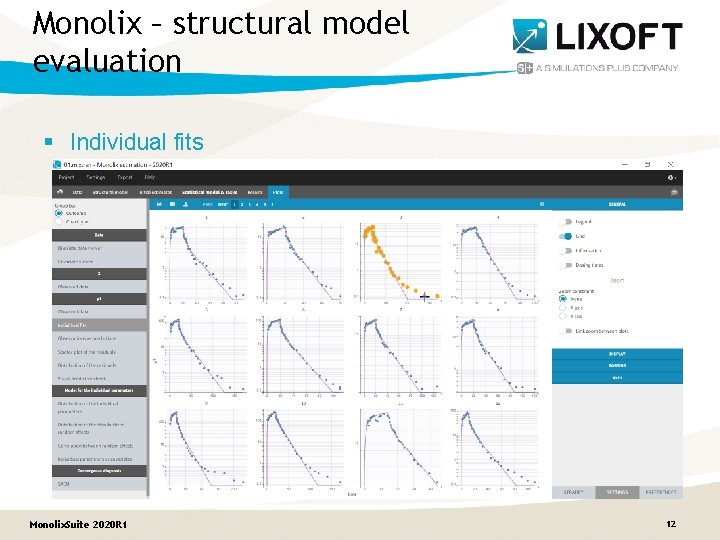
Monolix – structural model evaluation § Individual fits Monolix. Suite 2020 R 1 12

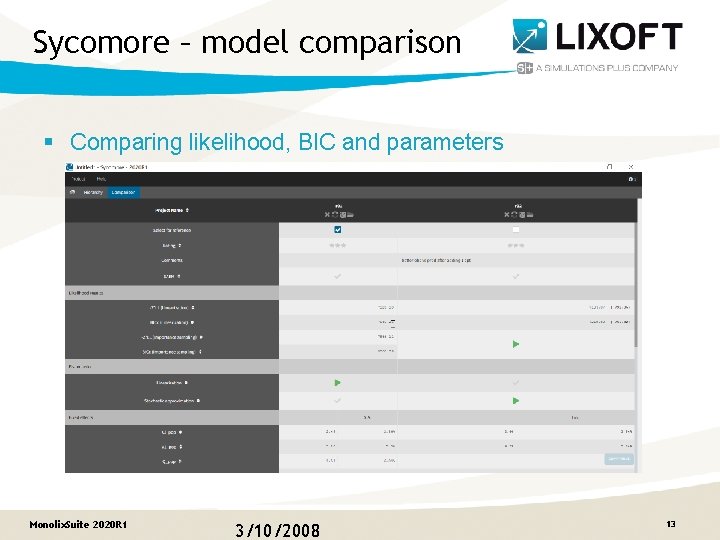
Sycomore – model comparison § Comparing likelihood, BIC and parameters Monolix. Suite 2020 R 1 3/10/2008 13

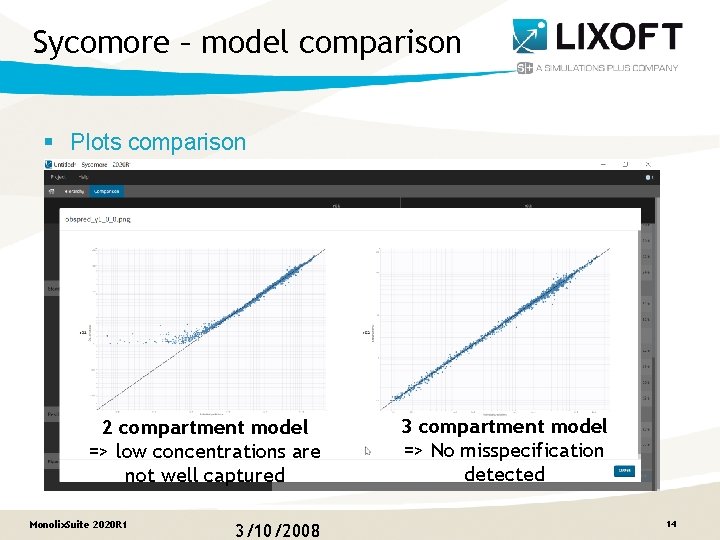
Sycomore – model comparison § Plots comparison 2 compartment model => low concentrations are not well captured Monolix. Suite 2020 R 1 3/10/2008 3 compartment model => No misspecification detected 14

Population approach Data: § several individuals § longitudinal data (repeated measurements over time) Þ yij = observation for individual i at time j Goal: characterize the typical biological phenomena for the population, but also the inter-individual variability hierarchical model for both the observations and the individual parameters Monolix. Suite 2020 R 1 15

Population approach Model for the observations: observation for individual i at time j structural model parameters of individual i residual error model standardized random variable Model for the individual parameters: parameters of individual i Monolix. Suite 2020 R 1 typical parameters in population random effect covariate of individual i 16

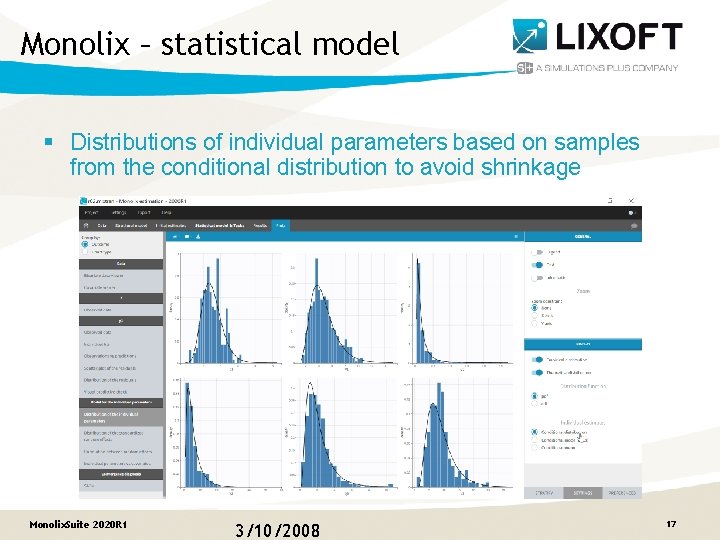
Monolix – statistical model § Distributions of individual parameters based on samples from the conditional distribution to avoid shrinkage Monolix. Suite 2020 R 1 3/10/2008 17

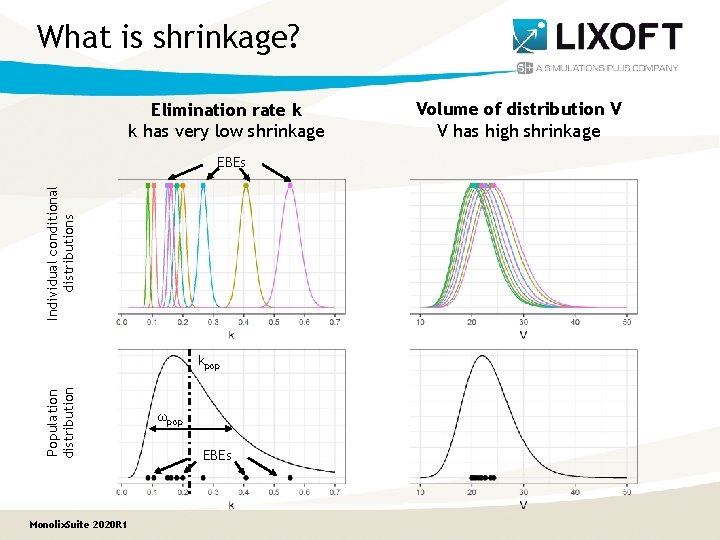
What is shrinkage? Elimination rate k k has very low shrinkage Individual conditional distributions EBEs Population distribution kpop Monolix. Suite 2020 R 1 ωpop EBEs Volume of distribution V V has high shrinkage

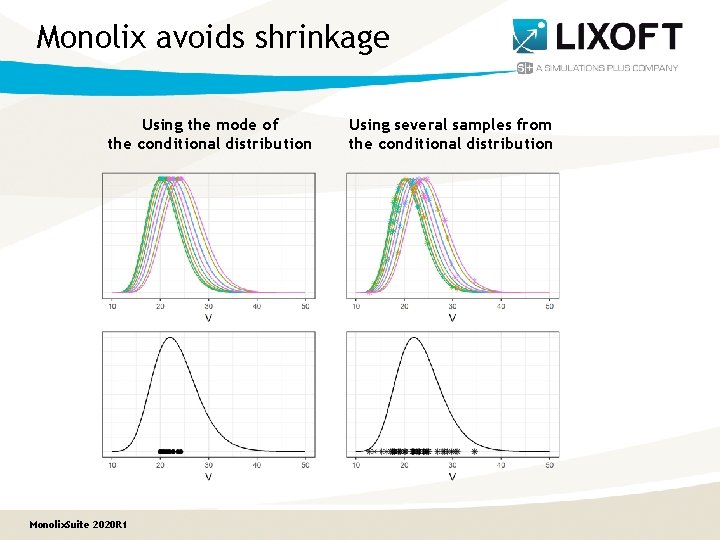
Monolix avoids shrinkage Using the mode of the conditional distribution Monolix. Suite 2020 R 1 Using several samples from the conditional distribution

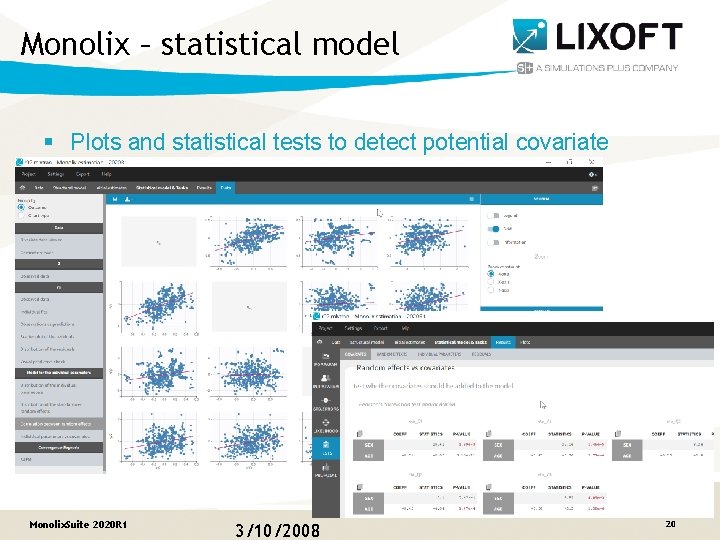
Monolix – statistical model § Plots and statistical tests to detect potential covariate effects Monolix. Suite 2020 R 1 3/10/2008 20

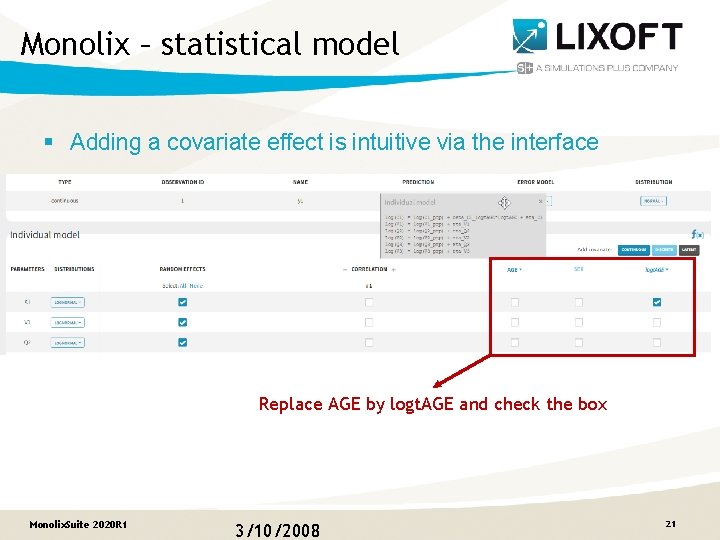
Monolix – statistical model § Adding a covariate effect is intuitive via the interface Replace AGE by logt. AGE and check the box Monolix. Suite 2020 R 1 3/10/2008 21

Monolix – statistical model § Transforming covariates Exponential relationship Monolix. Suite 2020 R 1 Power law relationship 22

Transforming covariates § Transforming categorical covariates Typical value for reference group (SEX=1): Typical value for SEX=2: Categorical covariates can also be transformed with reorganized modalities Monolix. Suite 2020 R 1 23

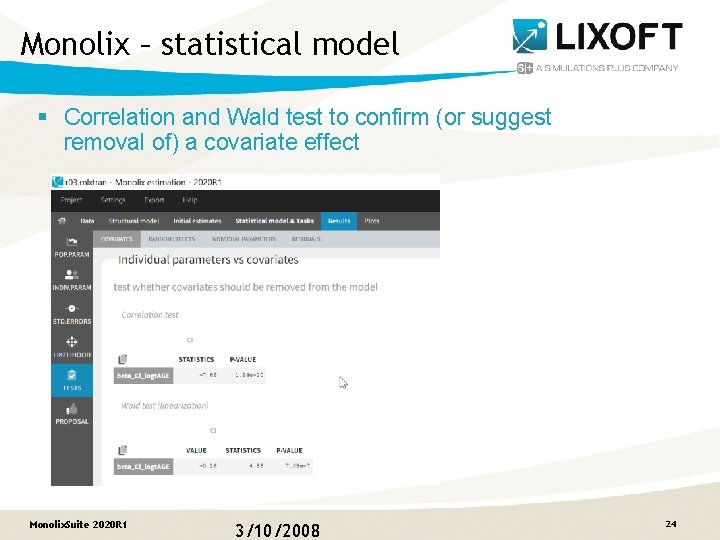
Monolix – statistical model § Correlation and Wald test to confirm (or suggest removal of) a covariate effect Monolix. Suite 2020 R 1 3/10/2008 24

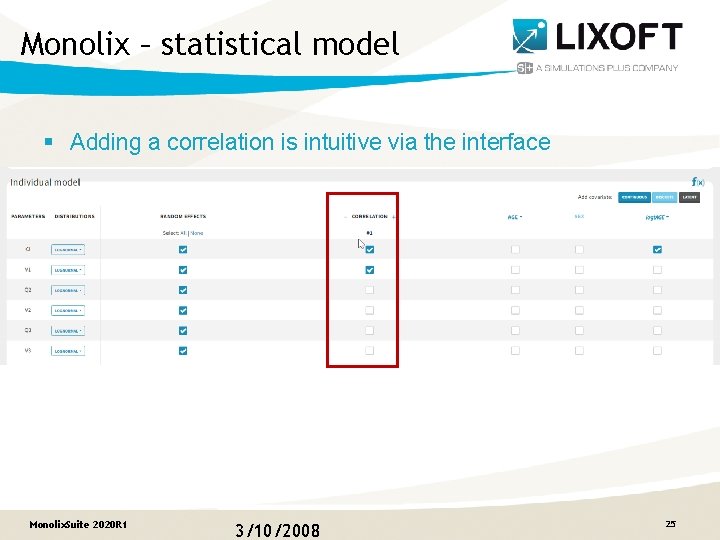
Monolix – statistical model § Adding a correlation is intuitive via the interface Monolix. Suite 2020 R 1 3/10/2008 25

Correlations between random effects no correlation : variance-covariance matrix is diagonal Monolix. Suite 2020 R 1 adds 1 parameter pair of correlated random effects 26

Monolix – automatic model building § Automatic statistical and covariate model building § Possibility to lock-in/lock-out relationships Monolix. Suite 2020 R 1 3/10/2008 27

Monolix – automatic model building § SCM: all covariate-parameter relationships are tested at each step § COSSAC: only the most promising (given the statistical tests) covariate-parameter relationship is tested at each step Monolix. Suite 2020 R 1 28

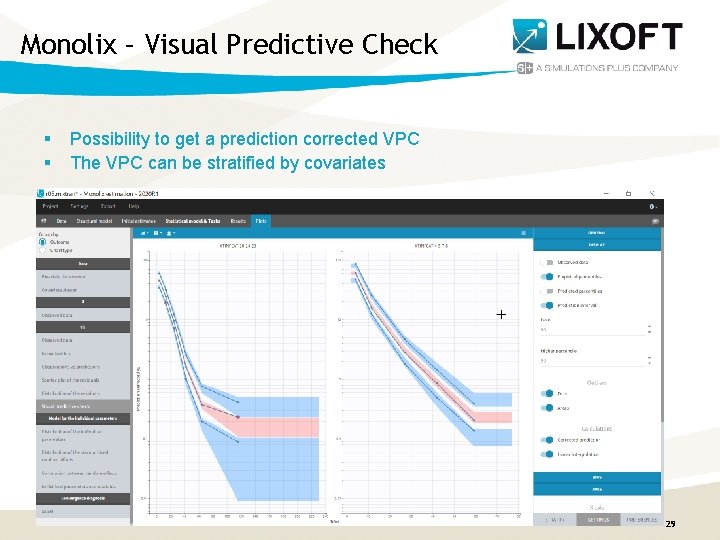
Monolix – Visual Predictive Check § § Possibility to get a prediction corrected VPC The VPC can be stratified by covariates Monolix. Suite 2020 R 1 29

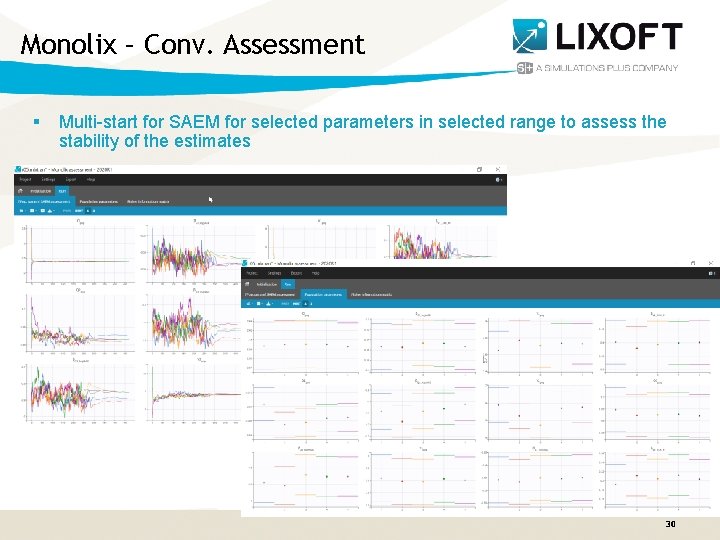
Monolix – Conv. Assessment § Multi-start for SAEM for selected parameters in selected range to assess the stability of the estimates Monolix. Suite 2020 R 1 30

Monolix - PK-PD model § Approaches for joint modelling Population PK parameters Individual PK parameters PD parameters (population + individual) Joint estimated Intermediate fixed (from PK analysis) estimated Sequential - fixed (from PK analysis) estimated Monolix. Suite 2020 R 1 + Computing speed Accuracy - + 31

Monolix - PK-PD model Joint approach Intermediate approach PK-PD dataset PK-PD model Sequential approach PK-PD dataset + PK parameters PK-PD model + regressors Cl = {use = regressor} V 1 = {use = regressor} Q 2 = {use = regressor} V 2 = {use = regressor} Q 3 = {use = regressor} V 3 = {use = regressor} Monolix. Suite 2020 R 1 PK parameters PD parameters 32

Monolix – writing models Mlxtran: human-readable language § Very natural syntax § Unified language for all applications § Support of continuous, categorical, count and time-to-event models (and any combination of them) § Model definition via macros or equations (ODEs, DDEs) § Extensive documentation (online or PDF) Monolix. Suite 2020 R 1 33
![Monolix writing models LONGITUDINAL input Cl V 1 Q 2 V 2 Monolix – writing models [LONGITUDINAL] input = {Cl, V 1, Q 2, V 2,](https://slidetodoc.com/presentation_image_h/04ad9c5627924131e8bd8cd785280fb0/image-34.jpg)
Monolix – writing models [LONGITUDINAL] input = {Cl, V 1, Q 2, V 2, Q 3, V 3, EEG 0, Imax, IC 50, gam, ksyn} EQUATION: V = V 1 k = Cl/V 1 k 12 = Q 2/V 1 k 21 = Q 2/V 2 k 13 = Q 3/V 1 k 31 = Q 3/V 3 kdeg = ksyn/EEG 0 using macros {Cc} = pkmodel(V, k, k 12, k 21, k 13, k 31) t_0=0 EEG_0=EEG 0 using ODEs ddt_EEG = ksyn * (1 - Imax * max(Cc, 0)^gam/(max(Cc, 0)^gam + IC 50^gam)) - kdeg*EEG OUTPUT: output = {Cc, EEG} Monolix. Suite 2020 R 1 34

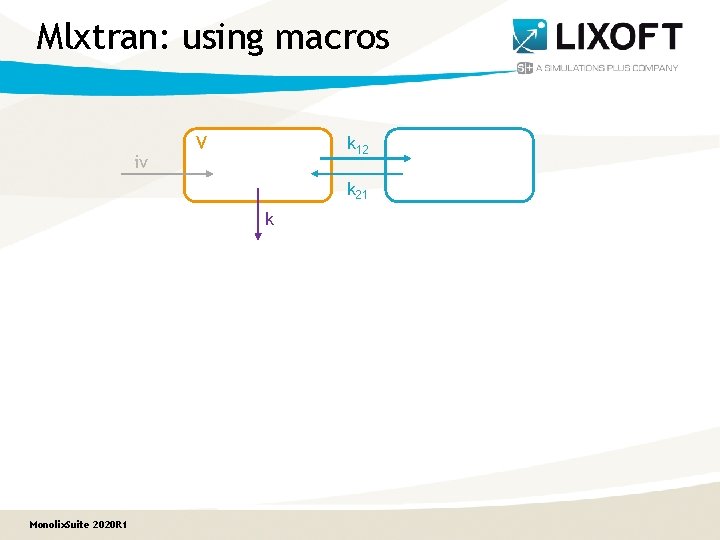
Mlxtran: using macros iv V k 12 k 21 k Monolix. Suite 2020 R 1

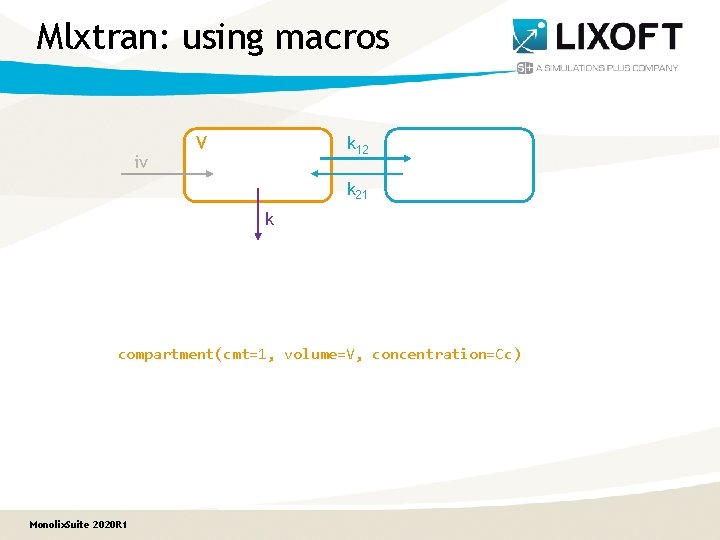
Mlxtran: using macros iv V k 12 k 21 k compartment(cmt=1, volume=V, concentration=Cc) Monolix. Suite 2020 R 1

Mlxtran: using macros iv V k 12 k 21 k compartment(cmt=1, volume=V, concentration=Cc) iv(adm=1, cmt=1) Monolix. Suite 2020 R 1

Mlxtran: using macros iv V k 12 k 21 k compartment(cmt=1, volume=V, concentration=Cc) iv(adm=1, cmt=1) peripheral(k 12, k 21) Monolix. Suite 2020 R 1

Mlxtran: using macros iv V k 12 k 21 k compartment(cmt=1, volume=V, concentration=Cc) iv(adm=1, cmt=1) peripheral(k 12, k 21) elimination(cmt=1, k) Monolix. Suite 2020 R 1
![Mlxtran using macros iv V k 12 k 21 k LONGITUDINAL input V Mlxtran: using macros iv V k 12 k 21 k [LONGITUDINAL] input = {V,](https://slidetodoc.com/presentation_image_h/04ad9c5627924131e8bd8cd785280fb0/image-40.jpg)
Mlxtran: using macros iv V k 12 k 21 k [LONGITUDINAL] input = {V, k, k 12, k 21} PK: compartment(cmt=1, volume=V, concentration=Cc) iv(adm=1, cmt=1) peripheral(k 12, k 21) elimination(cmt=1, k) OUTPUT: output = Cc Monolix. Suite 2020 R 1

Mlxtran: using macros § peripheral(k 12, k 21) implicitly defines the second compartment § for a second peripheral compartment, with compartment number 3: peripheral(k 13, k 31) § k 12 and k 21 are recognized keywords. To use other names or other parameterizations: peripheral(k 12=Q/V, k 21=Q/V 2) § iv(adm=1, cmt=1): adm refers to the ADM column of the data set, which permits to distinguish several types of administrations § in case of several outputs, output={Cc, Effect}, the match is done by order with the YTYPE column of the data set Monolix. Suite 2020 R 1
![Mlxtran using ODEs LONGITUDINAL input V k k 12 k 21 PK depotadm1 Mlxtran: using ODEs [LONGITUDINAL] input = {V, k, k 12, k 21} PK: depot(adm=1,](https://slidetodoc.com/presentation_image_h/04ad9c5627924131e8bd8cd785280fb0/image-42.jpg)
Mlxtran: using ODEs [LONGITUDINAL] input = {V, k, k 12, k 21} PK: depot(adm=1, target=Ac) EQUATION: t_0 = 0 Ac_0 = 0 Ap_0 = 0 ddt_Ac = -k*Ac – k 12*Ac + k 21*Ap ddt_Ap = k 12*Ac - k 21*Ap Cc = Ac/V OUTPUT: output = Cc Monolix. Suite 2020 R 1 iv cmt=1 V k 12 k 21 k cmt=2

Mlxtran syntax for ODEs § depot macro to link data set and model. q q q Bolus, zero-order and first-order absorptions can be used. Target must be an amount (not a concentration). For first-order absorption, write depot(target=Ac, ka). No need for a depot compartment. § define inital time and initial values with “_0”, for instance “t_0=“ or “Ac_0=“ § define ODEs with “ddt_” for instance “ddt_Ac=“ § ODEs in block EQUATION: (while macros are in block PK: ) Monolix. Suite 2020 R 1 43

Simulx - GUI § § § § Advanced simulator of clinical trials and decision-making tool Easy-to-use and interactive interface Intuitive workflow Interconnected with Monolix. Suite applications Flexible in building simulation scenarios Advanced computational capabilities with C++ engine Immediate visual feedback Export of plots and results Monolix. Suite 2020 R 1 44

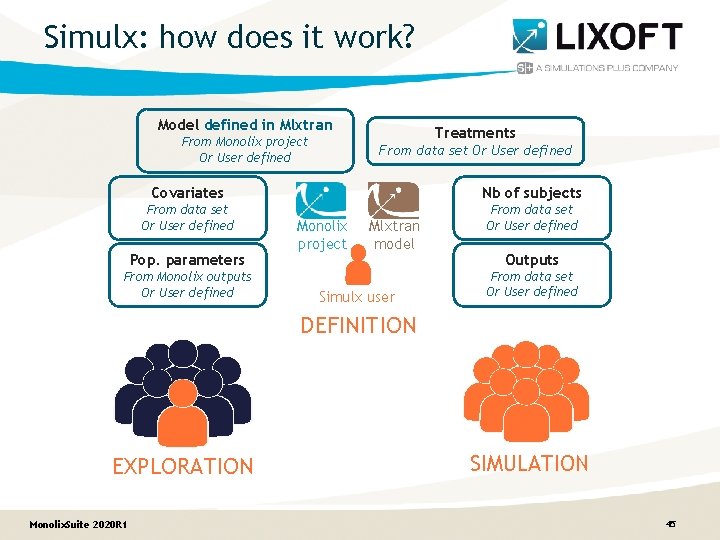
Simulx: how does it work? Model defined in Mlxtran From Monolix project Or User defined Treatments From data set Or User defined Covariates Nb of subjects From data set Or User defined Pop. parameters From Monolix outputs Or User defined Monolix project Mlxtran model Simulx user Outputs From data set Or User defined DEFINITION EXPLORATION Monolix. Suite 2020 R 1 SIMULATION 45

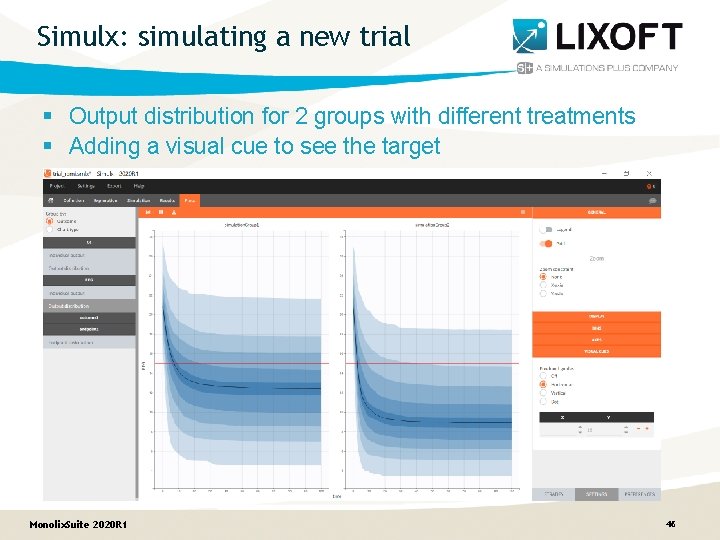
Simulx: simulating a new trial § Output distribution for 2 groups with different treatments § Adding a visual cue to see the target Monolix. Suite 2020 R 1 46

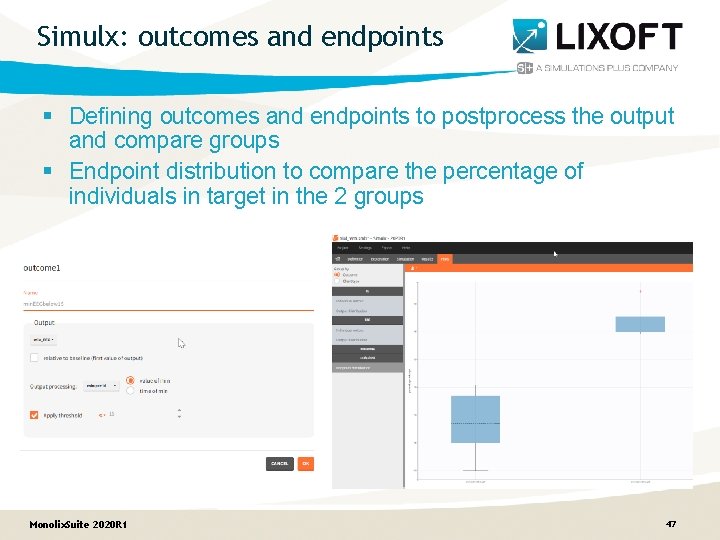
Simulx: outcomes and endpoints § Defining outcomes and endpoints to postprocess the output and compare groups § Endpoint distribution to compare the percentage of individuals in target in the 2 groups Monolix. Suite 2020 R 1 47

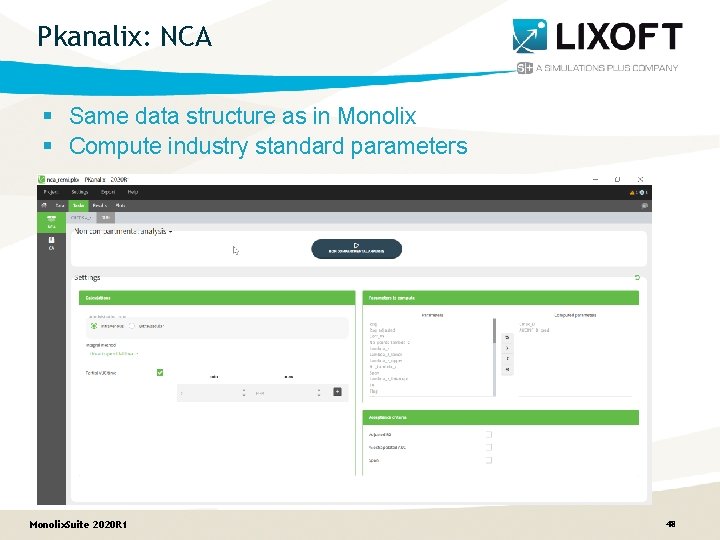
Pkanalix: NCA § Same data structure as in Monolix § Compute industry standard parameters Monolix. Suite 2020 R 1 48

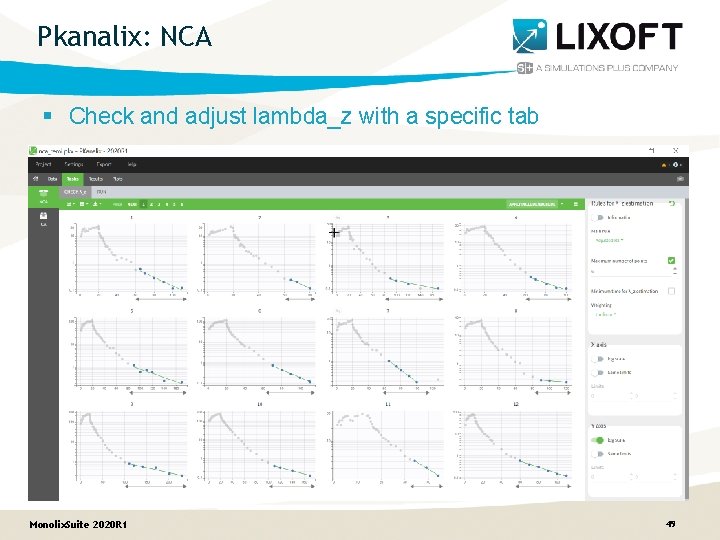
Pkanalix: NCA § Check and adjust lambda_z with a specific tab Monolix. Suite 2020 R 1 49

Pkanalix: CA § Simple PK model library and auto-init available § Easy export to Monolix to model whole populations Monolix. Suite 2020 R 1 50

Summary Interconnected applications that cover full model-based drug development workflow in a simple and efficient GUI environment. Plus: § Evaluation license § Technical support § Online documentation, videos, case studies § API to run projects from command line § … Monolix. Suite 2020 R 1 51

Documentation § § § § Data set: http: //dataset. lixoft. com Mlxtran language: http: //mlxtran. lixoft. com Datxplore: http: //datxplore. lixoft. com Monolix: http: //monolix. lixoft. com Sycomore: http: //sycomore. lixoft. com Simulx: http: //simulx. lixoft. com PKanalix: http: //pkanalix. lixoft. com § Technical support: support@lixoft. com § Case studies and videos: http: //lixoft. com/lixoft-university Monolix. Suite 2020 R 1 52