Monogenic Versus Polygenic Disease Monogenic One base change

Monogenic Versus Polygenic Disease • Monogenic: One base change = disease. Relatively easy to detect and analyze. • Polygenic/complex trait: A set of base changes affect the probability of disease. Subtle – hard to detect and analysis

Questions about Monogenic Disease ● What kinds of mutations?

Monogenic Disease • • • About 1500 proteins Most (73%) single base changes Average of 13 base changes per protein 84% missense/nonsense 14% splicing 2% regulatory Source: Human Gene Mutation Database (HGMD)

Questions about Monogenic Disease ● What kinds of mutations? Predominantly non-synonymous single base changes.

Questions about Monogenic Disease ● What kinds of mutations? Predominantly non-synonymous single base changes. ● In what ways do these SNPs affect protein function?
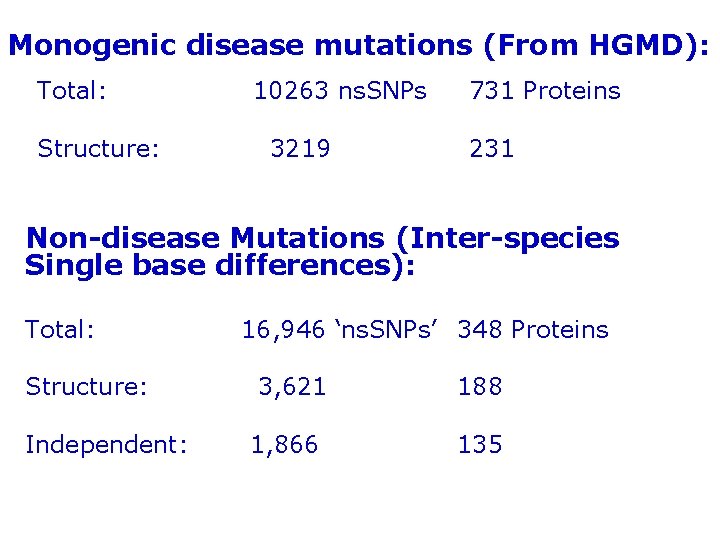
Monogenic disease mutations (From HGMD): Total: Structure: 10263 ns. SNPs 3219 731 Proteins 231 Non-disease Mutations (Inter-species Single base differences): Total: Structure: Independent: 16, 946 ‘ns. SNPs’ 348 Proteins 3, 621 1, 866 188 135

Structure Modeling • • High throughput >40% sequence ID Fixed environment Identify unreliable regions
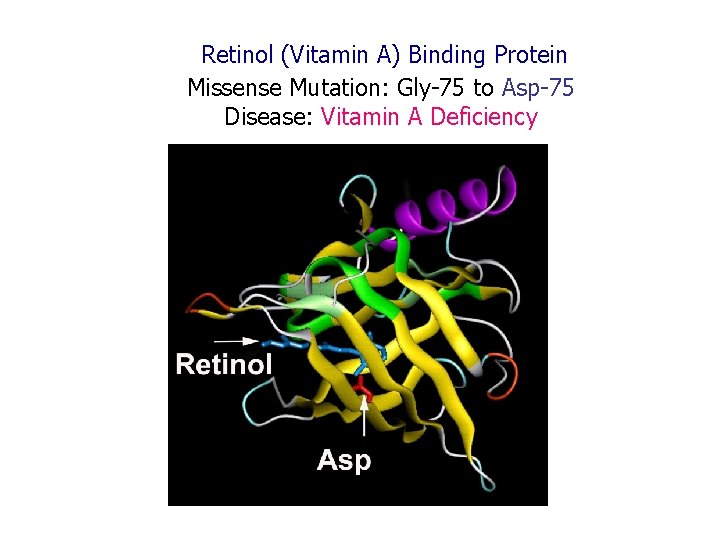
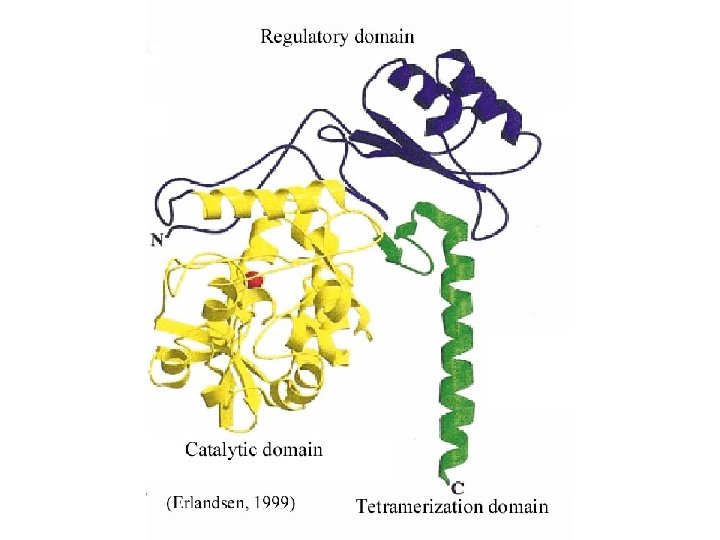
Retinol (Vitamin A) Binding Protein Missense Mutation: Gly-75 to Asp-75 Disease: Vitamin A Deficiency
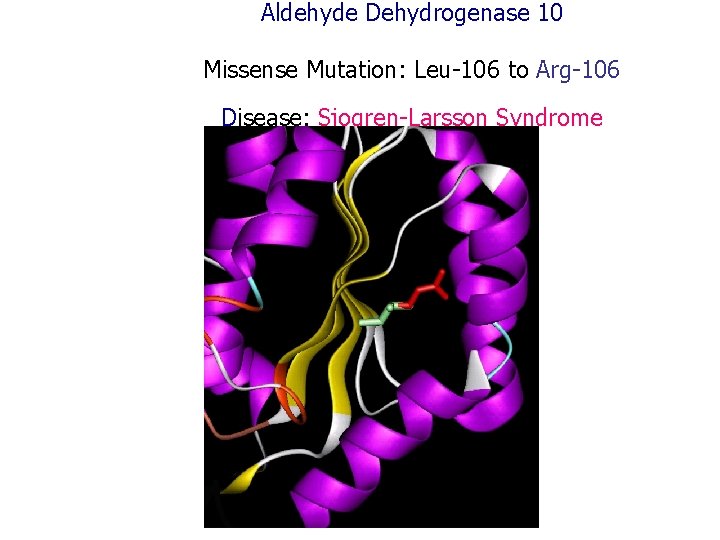
Aldehyde Dehydrogenase 10 Missense Mutation: Leu-106 to Arg-106 Disease: Sjogren-Larsson Syndrome
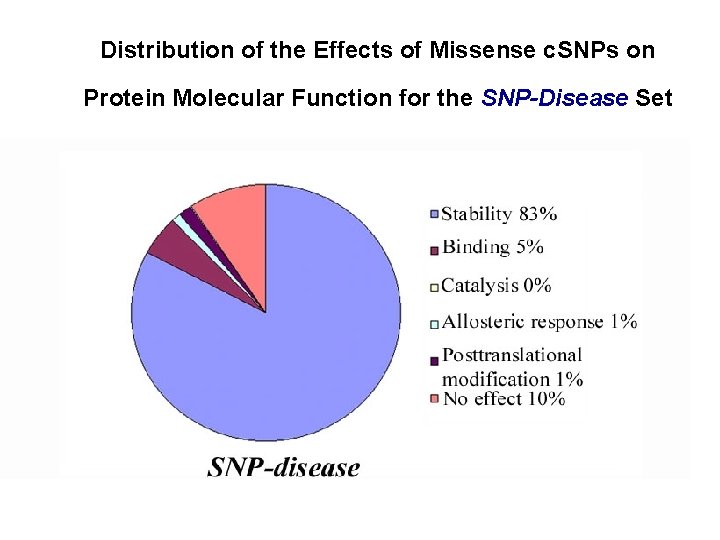
Distribution of the Effects of Missense c. SNPs on Protein Molecular Function for the SNP-Disease Set
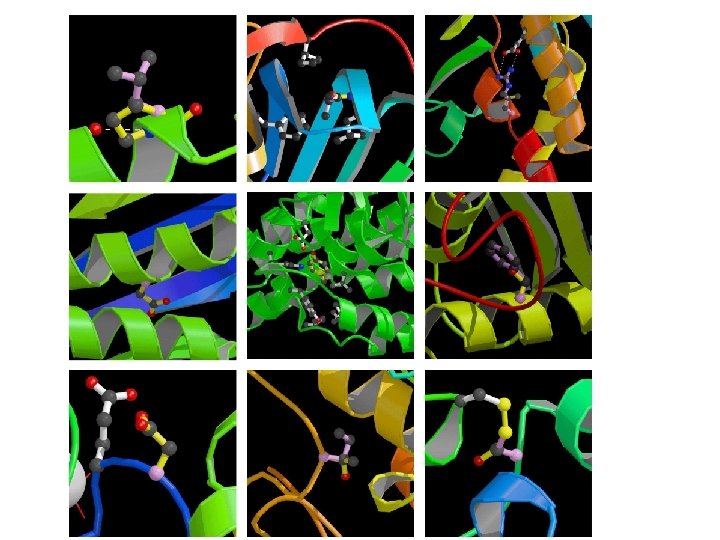
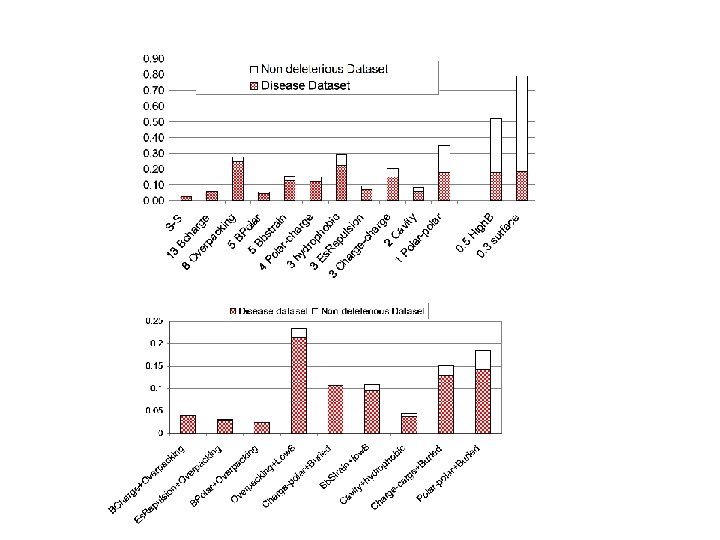
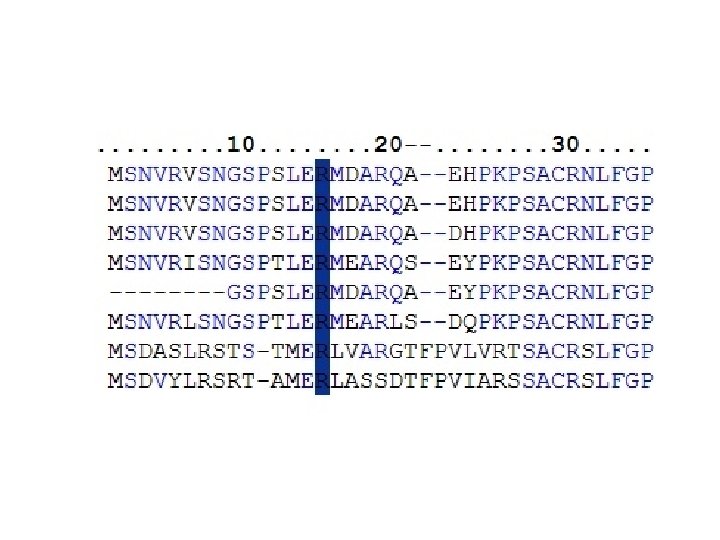

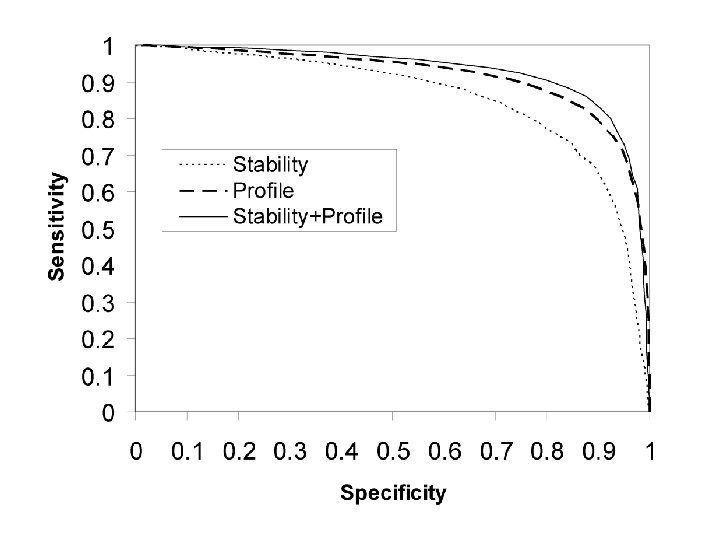
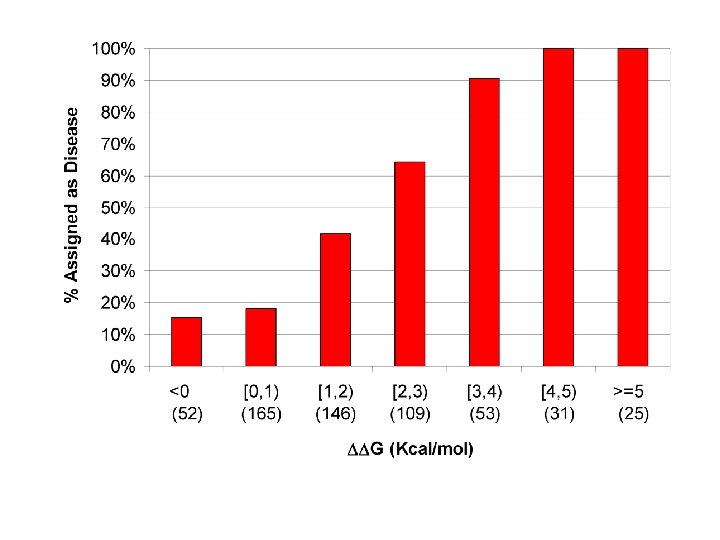

Questions about Monogenic Disease ● What kinds of mutations? Predominantly non-synonymous single base changes. ● In what ways do these SNPs affect protein function? Mild destabilization of protein structure dominates.

Questions about Monogenic Disease ● What kinds of mutations? Predominantly non-synonymous single base changes. ● In what ways do these SNPs affect protein function? Mild destabilization of protein structure dominates. ● Why does mild destabilization matter?

Protein Stability and Unfolding
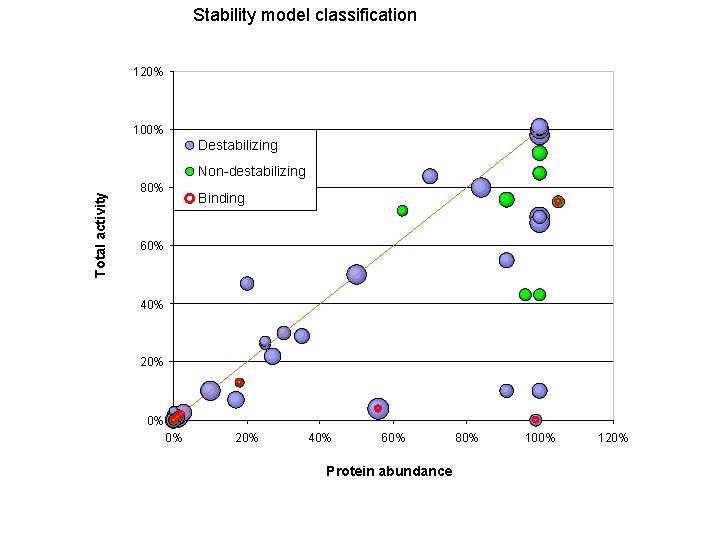
Stability model classification 120% 100% Destabilizing Total activity Non-destabilizing 80% Binding 60% 40% 20% 0% 0% 20% 40% 60% Protein abundance 80% 100% 120%
![Steady State Abundance d[P]/dt = 0 = C – k 1. [P] – k Steady State Abundance d[P]/dt = 0 = C – k 1. [P] – k](http://slidetodoc.com/presentation_image/4bc1f20aa50f54f5a19f7faee0e59336/image-24.jpg)
Steady State Abundance d[P]/dt = 0 = C – k 1. [P] – k 2. [P]e-∆G/k. T

Questions about Monogenic Disease ● What kinds of mutations? Predominantly non-synonymous single base changes. ● In what ways do these SNPs affect protein function? Mild destabilization of protein structure dominates. ● Why does mild destabilization matter? Increased rate of protein removal.

Questions about Complex Trait Disease ● Are there non-disease causing deleterious SNPs?
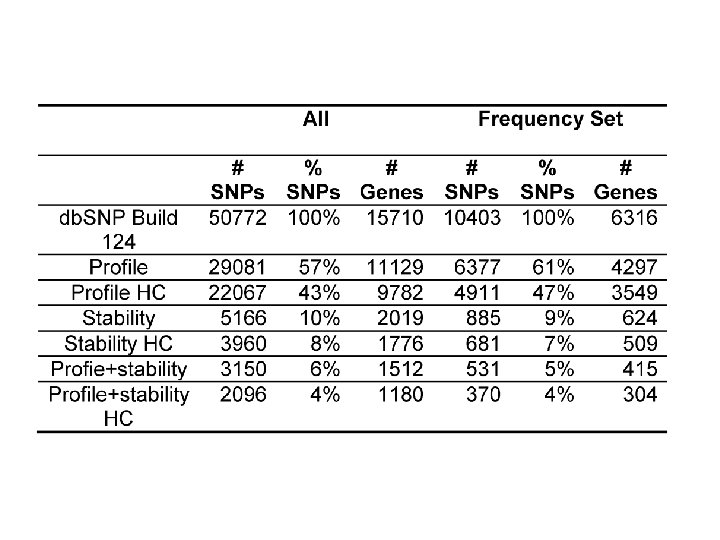
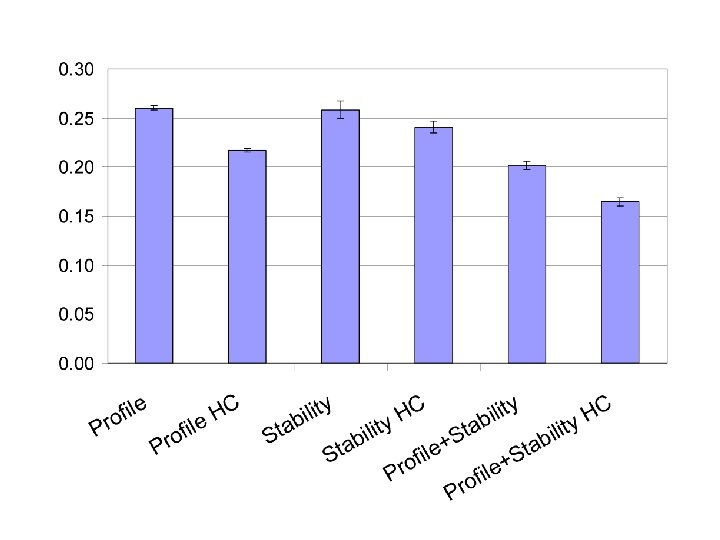
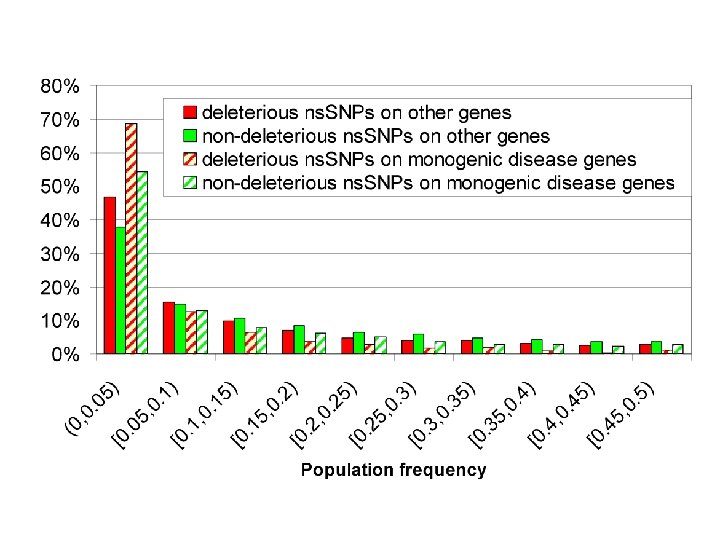
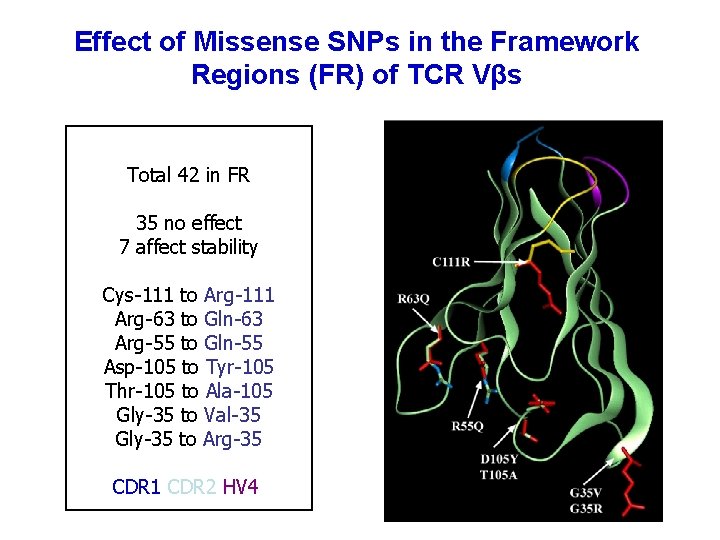
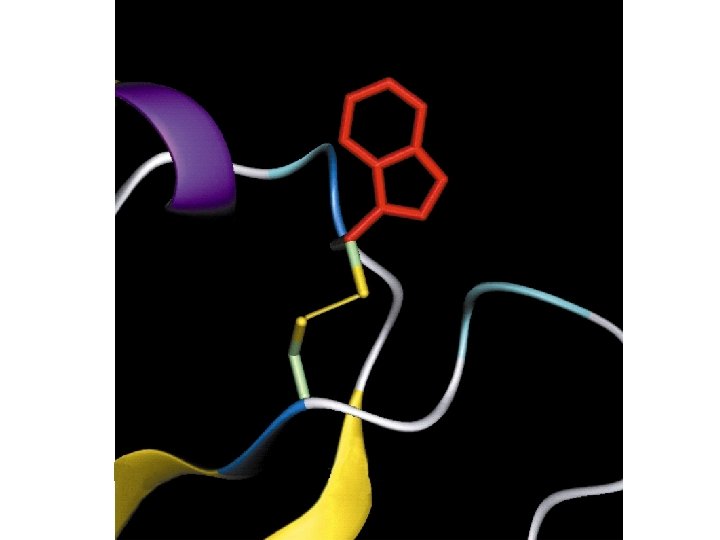
Effect of Missense SNPs in the Framework Regions (FR) of TCR Vβs Total 42 in FR 35 no effect 7 affect stability Cys-111 to Arg-111 Arg-63 to Gln-63 Arg-55 to Gln-55 Asp-105 to Tyr-105 Thr-105 to Ala-105 Gly-35 to Val-35 Gly-35 to Arg-35 CDR 1 CDR 2 HV 4
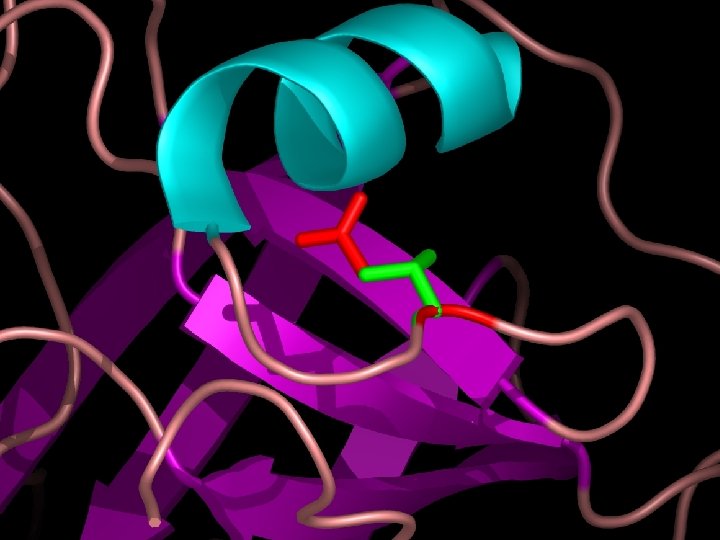

Questions about Complex Trait Disease ● Are there non-disease causing deleterious SNPs? Yes, about 25% of population SNPs are deleterious.

Questions about Complex Trait Disease ● Are there non-disease causing deleterious SNPs? Yes, about 25% of population SNPs are deleterious. ● What is the difference between the ~1000 monogenic disease proteins and the rest?
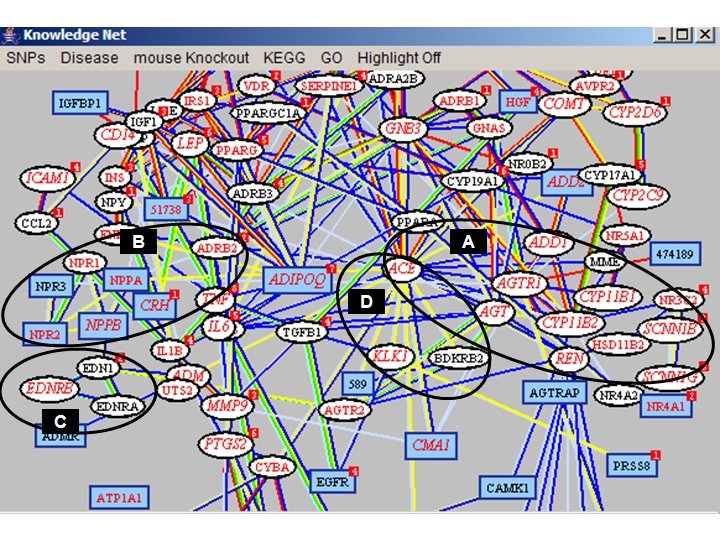

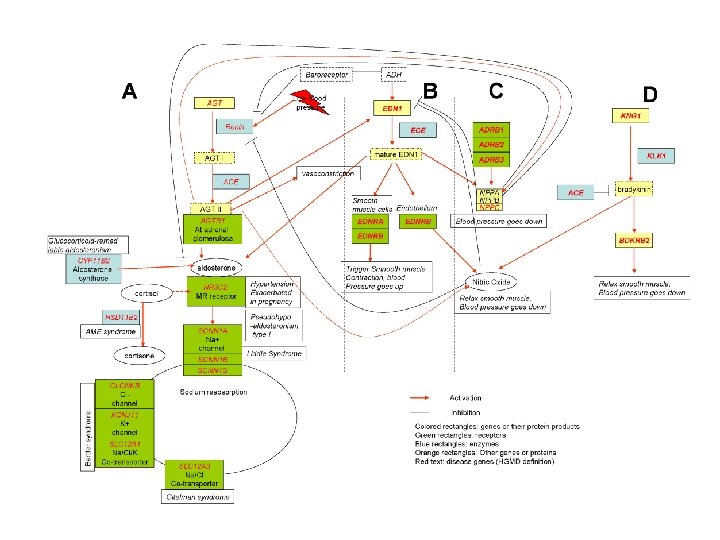

Questions about Complex Trait Disease ● Are there non-disease causing deleterious SNPs? Yes, about 25% of population SNPs are deleterious. ● What is the difference between the ~1000 monogenic disease proteins and the rest? Network environment provides buffering against component defects.

Questions about Complex Trait Disease ● Are there non-disease causing deleterious SNPs? Yes, about 25% of population SNPs are deleterious. ● What is the difference between the ~1000 monogenic disease proteins and the rest? Network environment provides buffering against component defects. ● Are deleterious SNPs good markers for disease susceptibility?
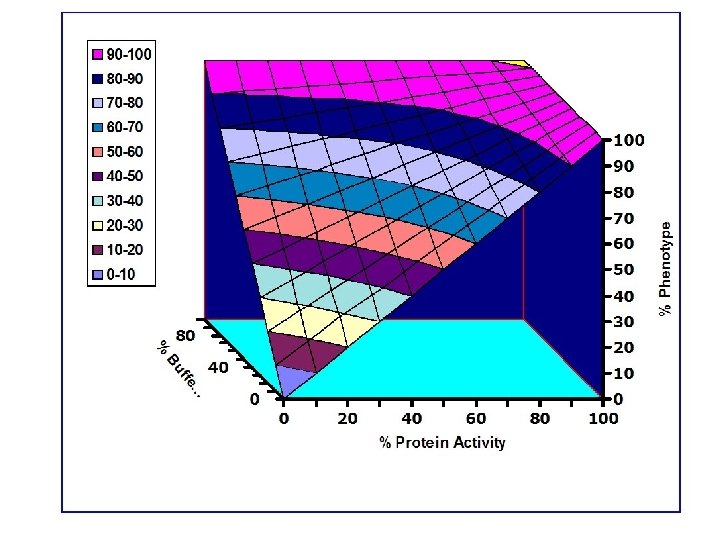

Questions about Complex Trait Disease ● Are there non-disease causing deleterious SNPs? Yes, about 25% of population SNPs are deleterious. ● What is the difference between the ~1000 monogenic disease proteins and the rest? Network environment provides buffering against component defects. ● Are deleterious SNPs good markers for disease susceptibility? Don’t know…

Zhen Wang Peng Yue Eugene Melamud

http: //www. SNPS 3 D. org
- Slides: 43