Molecular Techniques in Molecular Systematics DNADNA hybridisation Measures

Molecular Techniques in Molecular Systematics

DNA-DNA hybridisation - Measures the degree of genetic similarity between pools of DNA sequences. - Normally used to determine the genetic distance between two species. - The method compares the melting of a labeled sample after it is hybridized to iteself vs its melting after hybridized to unlabeled DNA of another organism.

RAPD, AFLP, Minisatellites - Data is scored as presence (1) or absence (0) of a DNA band/fragment. - Suitable for studies involving closely related taxa because the variation detected by these markers is high. - Homology problem may arise due to comigration of DNA bands/fragments. RADP data

PCR-RFLP / CAP - Data scored as presence (1) or absence (0) of a restriction site. - 4 -base and 6 -base cutters are used to generate data. - Data scoring becomes difficult if the variations involve length mutations (deletions or insertions / indels)

CAPS PCR products of the trn. S-trn. M region (cp. DNA) restricted by Taq. I 580 bp 290 bp 180 bp

PCR-RFLP / CAP - The probability of restriction site loss is higher than restriction site gain. - A site loss could be due to various characterstates (conditions). Restricted by Dra. I (TTTAAA) Sequence Taxon Species Species A B C D E CGTATTTAAACCGCTC CGTATTCAAACCGCTC CGTATTGAAACCGCTC CGTATTTCAACCGCTC Restriction Cut No Cut

DNA Sequencing - DNA data is multiple state data. It normally exist in 4 different bases (A, T, C and G). - DNA data must be aligned (multiple sequence alignment) in order to be scored. - CLUSTAL software is used to align DNA sequences.
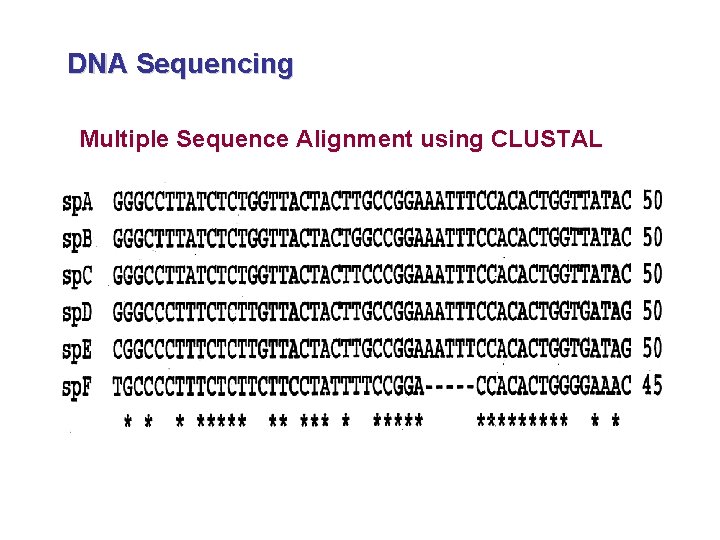
DNA Sequencing Multiple Sequence Alignment using CLUSTAL
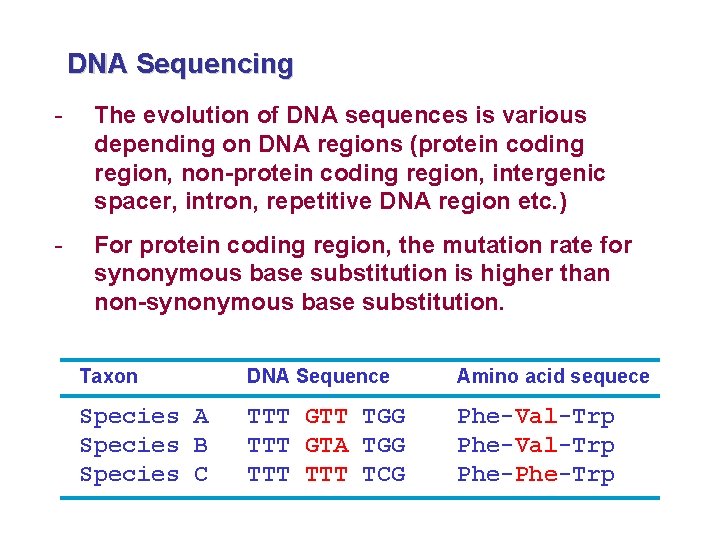
DNA Sequencing - The evolution of DNA sequences is various depending on DNA regions (protein coding region, non-protein coding region, intergenic spacer, intron, repetitive DNA region etc. ) - For protein coding region, the mutation rate for synonymous base substitution is higher than non-synonymous base substitution. Taxon DNA Sequence Amino acid sequece Species A Species B Species C TTT GTT TGG TTT GTA TGG TTT TCG Phe-Val-Trp Phe-Trp

Amino Acid Sequencing - Amino acid sequencing was practiced before the establishment of DNA sequencing methods. - Amino acid data is a multiple state data. It consists of 20 states (20 different amino acids).

Single-Letter Amino Acid Code G - Glycine (Gly) P - Proline (Pro) A - Alanine (Ala) V - Valine (Val) L - Leucine (Leu) I - Isoleucine (Ile) M - Methionine (Met) C - Cysteine (Cys) F - Phenylalanine (Phe) Y - Tyrosine (Tyr) W - Tryptophan (Trp) H - Histidine (His) K - Lysine (Lys) R - Arginine (Arg) Q - Glutamine (Gln) N - Asparagine (Asn) E - Glutamic Acid (Glu) D - Aspartic Acid (Asp) S - Serine (Ser) T - Threonine (Thr)

Amino Acid Sequencing - Amino acid sequencing is time consuming. It is done by HPLC approach. - At present the amino acid sequences are generated mainly from the inference of protein coding DNA sequences. - Amino acid of hemoglobin (in animals) and amino acid of large subunit ribulose-1, 5 -bisphophate carboxylase (rbc. L; in plants) are widely used for phylogenetic studies.
- Slides: 12