Molecular Subtypes Not Quite Ready for Prime Time

Molecular Subtypes: Not Quite Ready for Prime Time Scott Kopetz, MD, Ph. D. Department of GI Medical Oncology MD Anderson Cancer Center

Individual Biomarkers versus Molecular Subtypes YES • Individual biomarkers: – Microsatellite instability in all patients • For adjuvant decisions in Stage II and screening for HNPCC – KRAS, NRAS, BRAF in all metastatic patients • For consideration of EGFR sensitivity and prognosis • Molecular subtypes – 200 gene… 400 gene…whole exome sequencing – Gene expression profiles – Proteomic panels NOT YET

Why not yet…. ? • We need studies to evaluate the benefit from extended molecular testing (beyond KRAS, NRAS, BRAF) • We need to define the molecular subtypes by gene expression • We need validated assays to move into the clinic 3

Why not yet…. ? • We need studies to evaluate the benefit from extended molecular testing (beyond KRAS, NRAS, BRAF) • We need to define the molecular subtypes by gene expression • We need validated assays to move into the clinic 4
“My panel is bigger than yours…” 5
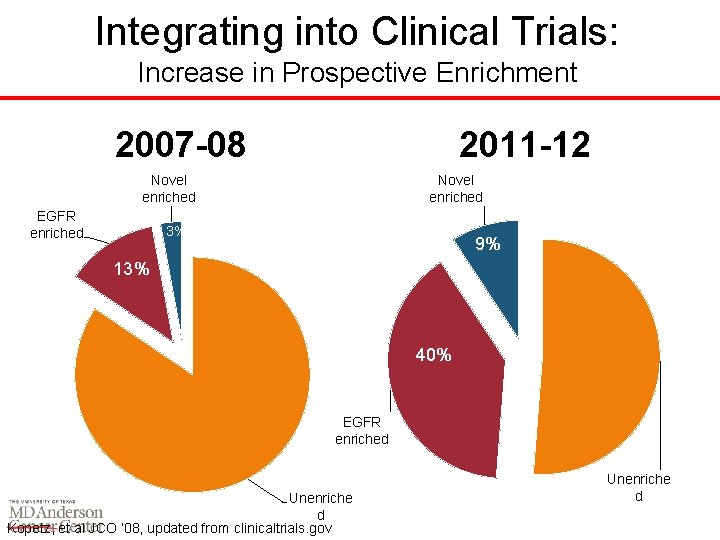
Integrating into Clinical Trials: Increase in Prospective Enrichment 2007 -08 2011 -12 Novel enriched EGFR enriched Novel enriched 3% 9% 13% 40% EGFR enriched Unenriche d Kopetz, et al JCO ‘ 08, updated from clinicaltrials. gov Unenriche d
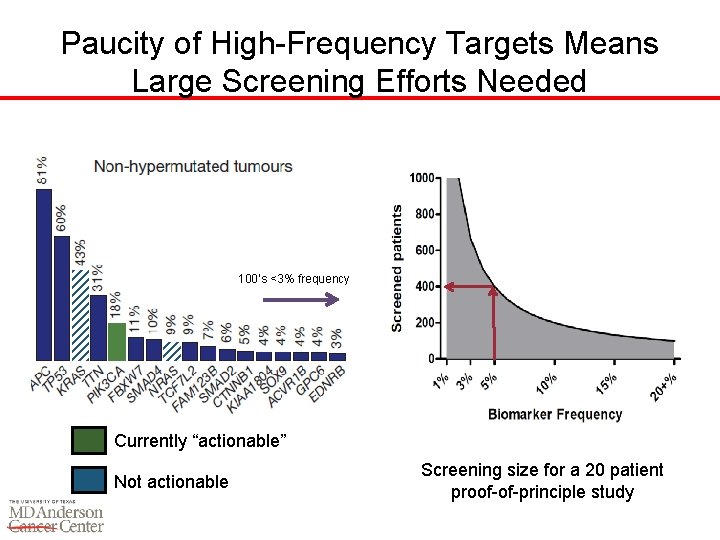
Paucity of High-Frequency Targets Means Large Screening Efforts Needed 100’s <3% frequency Currently “actionable” Not actionable Screening size for a 20 patient proof-of-principle study
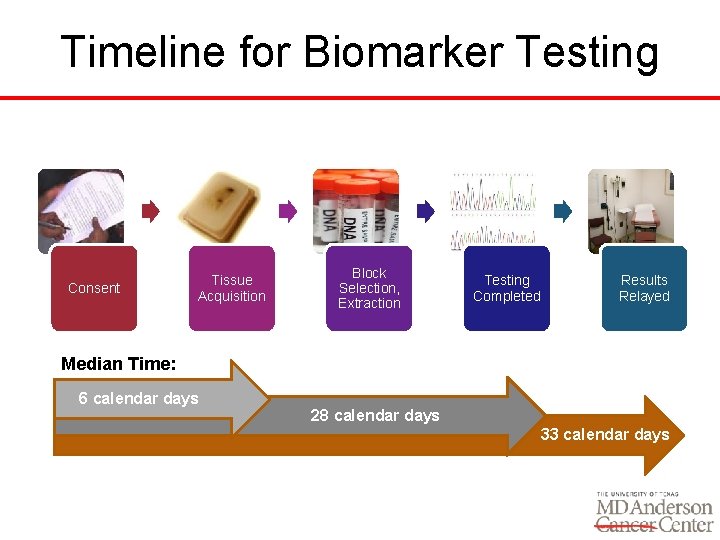
Timeline for Biomarker Testing Consent Tissue Acquisition Block Selection, Extraction Testing Completed Results Relayed Median Time: 6 calendar days 28 calendar days 33 calendar days

ATTACC Program: Assessment of Targeted Therapies Against Colorectal Cancer Eligibility • 5 -FU refractory disease • ECOG PS 0 -1 • Tissue available for molecular testing Screening • Patients undergo biomarker assessment • CLIA-certified assays Allocation • Based on biomarker, patients allocated to one of several treatment protocols • Patients not expressing biomarker of interest are treated in unenriched protocols S. Kopetz, PI
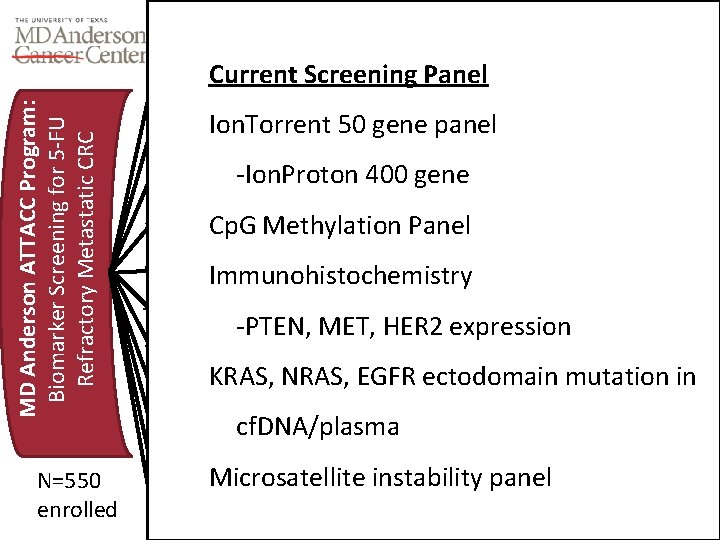
Enrichment Therapeutic Mechanism Akt inhibitor Current Screening Panel MD Anderson ATTACC Program: Biomarker Screening for 5 -FU Refractory Metastatic CRC PTEN Loss or PIK 3 CAmut MK-2206 Cp. G Island Methylation Demethylator Mitotic inhib Azacitadine + XELOX Nab-paclitaxel HER 2 overexpression HER 2 mutation HER 2 inhibition ERB family inhib Trastuzumab +/- EGFR TBA Ion. Torrent 50 gene panel -Ion. Proton 400 gene Exon 3, 4 KRAS or NRAS mutant RAF inhibition ERK inhibition Cp. G Methylation Panel LY 3009120 Biomed Valley BRAF+EGFR+irino Immunohistochemistry Vemurafenib +cetux+ irino BRAF Mutation PTEN Loss/KRAS WT PI 3 K-beta inhibitor SAR 26031 -PTEN, MET, HER 2 expression MEK + EGFR inhibition Panitum + Aquired RAS mutation Trametinib KRAS, NRAS, EGFR ectodomain mutation in Alternate EGFR Panitumumab EGFR ectodomain mutation cf. DNA/plasma Dual MEK, PI 3 K KRAS and PIK 3 CA mutation N=550 enrolled Agent(s) BYL 719 and MEK 162 CTLA 4 and PD 1 panel Nivolumumab, Ipilumumb Microsatellite instability MSI High Triple KRAS/BRAF/NRAS WT EGFR+HER 2 Cetuximab + trastuzumab
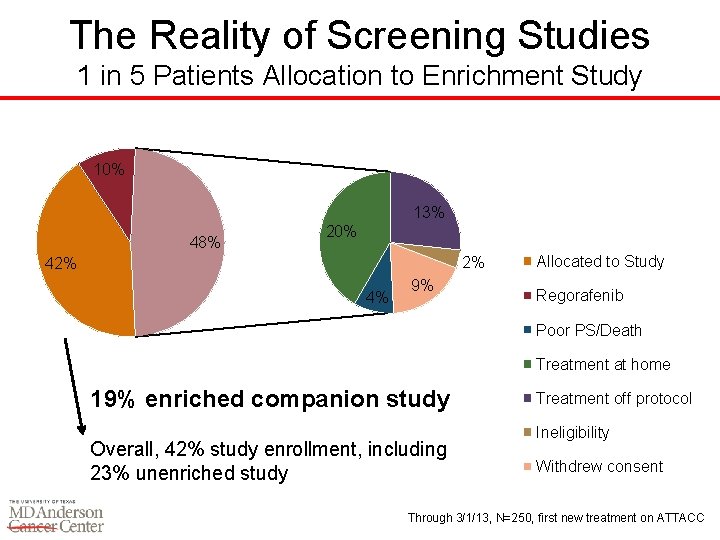
The Reality of Screening Studies 1 in 5 Patients Allocation to Enrichment Study 10% 13% 48% 20% 2% 4% 9% Allocated to Study Regorafenib Poor PS/Death Treatment at home 19% enriched companion study Overall, 42% study enrollment, including 23% unenriched study Treatment off protocol Ineligibility Withdrew consent Through 3/1/13, N=250, first new treatment on ATTACC

Practical Considerations for Enrichment Studies • Enrichment strategies require… – Consenting patients for screening – Explaining the study – High research staff utilization per “screen failure” • Patient-satisfaction is very dependent on biomarker turn-around time – Obtaining outside paraffin blocks is rate-limiting step – How long should one delay treatment waiting for a 5% frequency biomarker? • Other experimental options need to be available – Enrichment study is hard to justify to patients in isolation
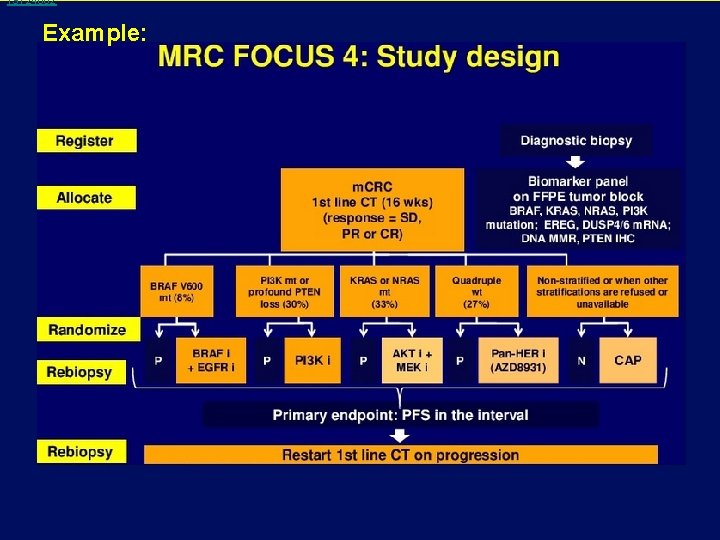
19124802 Example:
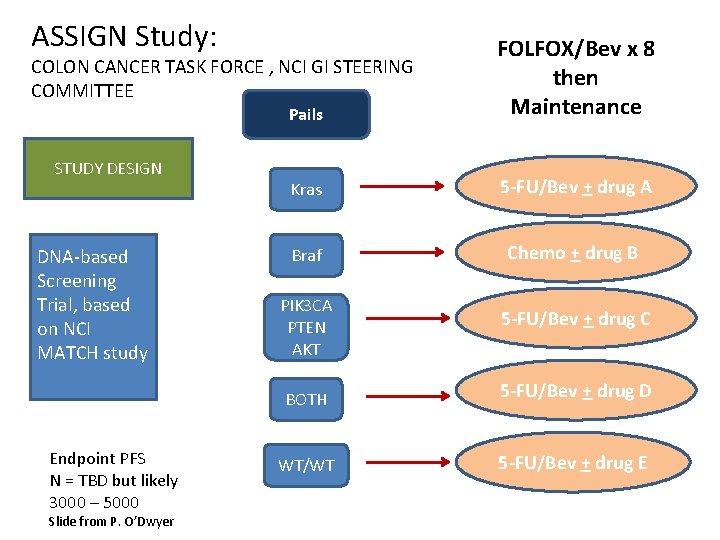
ASSIGN Study: Pails FOLFOX/Bev x 8 then Maintenance Kras 5 -FU/Bev + drug A Braf Chemo + drug B COLON CANCER TASK FORCE , NCI GI STEERING COMMITTEE STUDY DESIGN DNA-based Screening Trial, based on NCI MATCH study Endpoint PFS N = TBD but likely 3000 – 5000 Slide from P. O’Dwyer PIK 3 CA PTEN AKT 5 -FU/Bev + drug C BOTH 5 -FU/Bev + drug D WT/WT 5 -FU/Bev + drug E
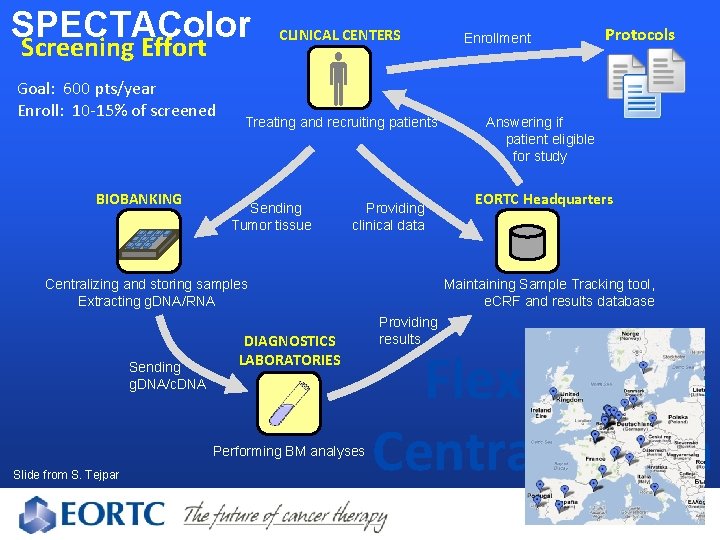
SPECTAColor Screening Effort Goal: 600 pts/year Enroll: 10 -15% of screened BIOBANKING CLINICAL CENTERS Treating and recruiting patients Sending Tumor tissue Providing clinical data Sending g. DNA/c. DNA Performing BM analyses Slide from S. Tejpar Protocols Answering if patient eligible for study EORTC Headquarters Maintaining Sample Tracking tool, e. CRF and results database Centralizing and storing samples Extracting g. DNA/RNA DIAGNOSTICS LABORATORIES Enrollment Providing results Flexibility & Centralization

To date… Limited Prospective Biomarker Success in CRC New or Anticipated Agents/Indications – – No new biomarkerdirected therapy Bevacizumab (2 nd line) Ziv-aflibercept Regorafenib TAS-102 Wrong premise, wrong implementation, or still too early?

Why not yet…. ? • We need studies to evaluate the benefit from extended molecular testing (beyond KRAS, NRAS, BRAF) • We need to define the molecular subtypes by gene expression • We need validated assays to move into the clinic 17
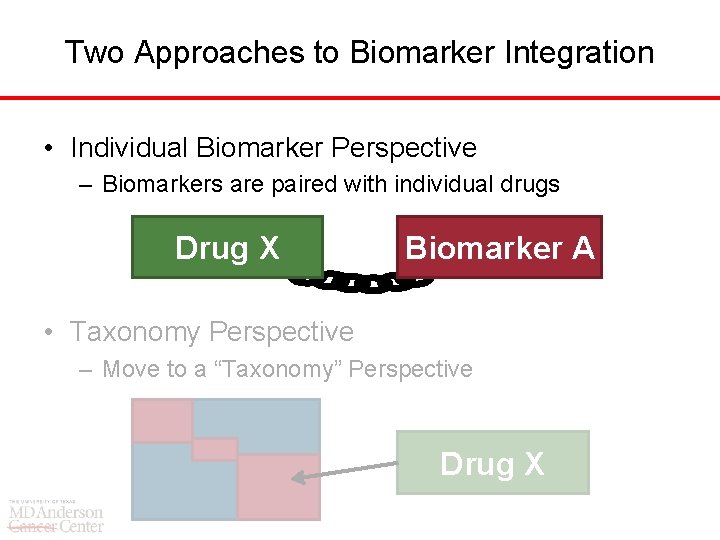
Two Approaches to Biomarker Integration • Individual Biomarker Perspective – Biomarkers are paired with individual drugs Drug X Biomarker A • Taxonomy Perspective – Move to a “Taxonomy” Perspective Drug X
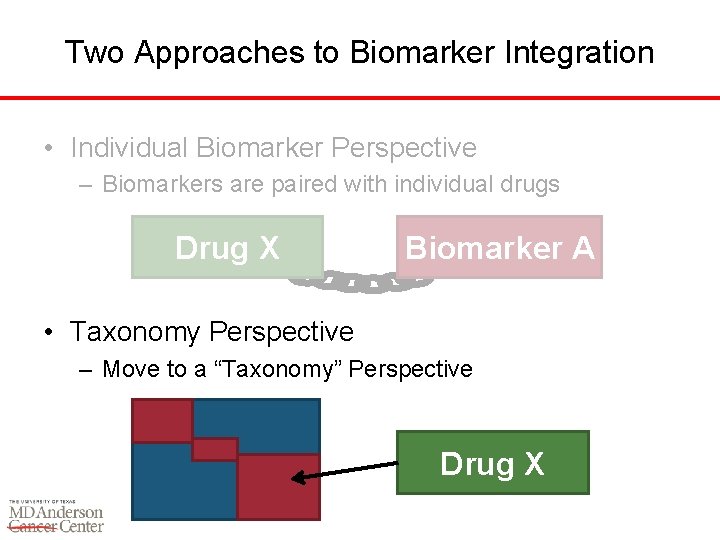
Two Approaches to Biomarker Integration • Individual Biomarker Perspective – Biomarkers are paired with individual drugs Drug X Biomarker A • Taxonomy Perspective – Move to a “Taxonomy” Perspective Drug X

Definitions Taxidermy = Stuffing Taxonomy = Grouping based on common patterns

CRC Taxonomy Hasn’t Been Defined To Date Lymphoma Her 2 -pos Luminal B Luminal A Basal Breast Cancer Colorectal Cancer ? Sotiriou et al NEJM, 2009; Alizadeh et al, Nature 2000
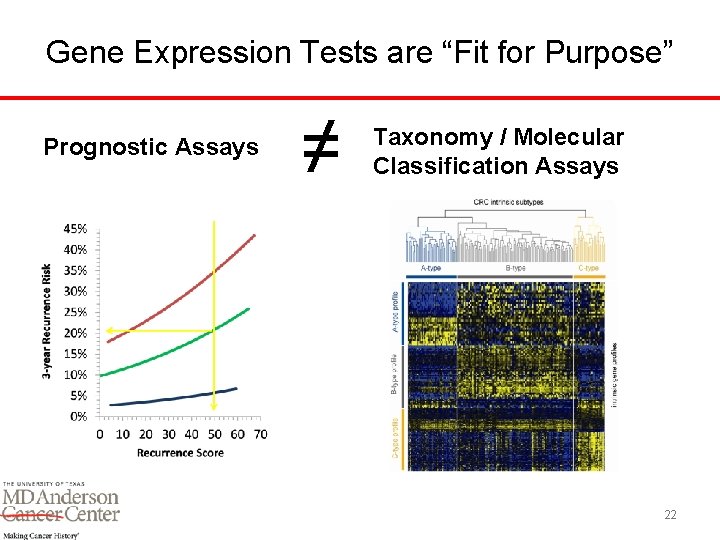
Gene Expression Tests are “Fit for Purpose” Prognostic Assays ≠ Taxonomy / Molecular Classification Assays 22
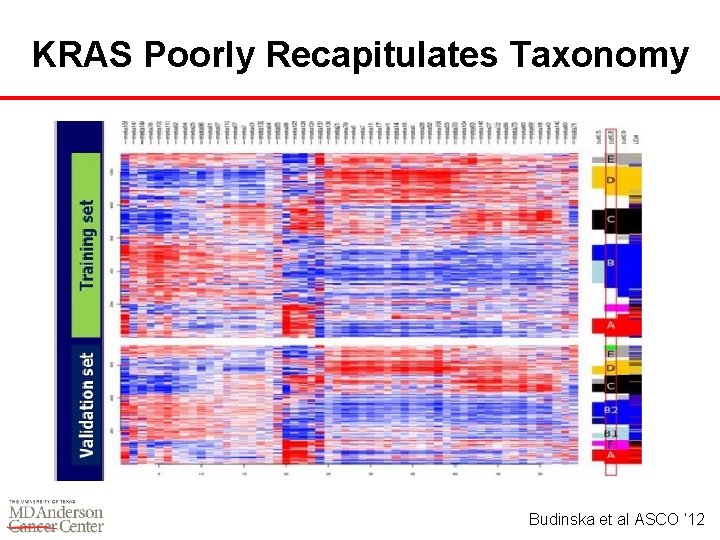
KRAS Poorly Recapitulates Taxonomy Budinska et al ASCO ‘ 12
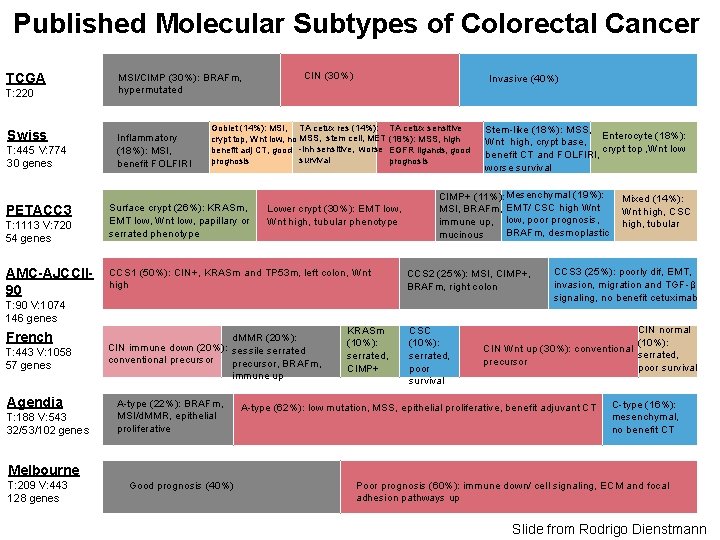
Published Molecular Subtypes of Colorectal Cancer TCGA T: 220 Swiss T: 445 V: 774 30 genes PETACC 3 T: 1113 V: 720 54 genes AMC-AJCCII 90 MSI/CIMP (30%): BRAFm, hypermutated Inflammatory (18%): MSI, benefit FOLFIRI CIN (30%) Invasive (40%) Goblet (14%): MSI, TA cetux res (14%): crypt top, Wnt low, no MSS, stem cell, MET benefit adj CT, good -inh sensitive, worse survival prognosis Surface crypt (26%): KRASm, EMT low, Wnt low, papillary or serrated phenotype TA cetux sensitive (18%): MSS, high EGFR ligands, good prognosis Lower crypt (30%): EMT low, Wnt high, tubular phenotype CCS 1 (50%): CIN+, KRASm and TP 53 m, left colon, Wnt high CIMP+ (11%): Mesenchymal (19%): MSI, BRAFm, EMT/ CSC high Wnt immune up, low, poor prognosis, BRAFm, desmoplastic mucinous CCS 2 (25%): MSI, CIMP+, BRAFm, right colon T: 90 V: 1074 146 genes French T: 443 V: 1058 57 genes Agendia T: 188 V: 543 32/53/102 genes d. MMR (20%): CIN immune down (20%): sessile serrated conventional precursor, BRAFm, immune up A-type (22%): BRAFm, MSI/d. MMR, epithelial proliferative KRASm (10%): serrated, CIMP+ Stem-like (18%): MSS, Enterocyte (18%): Wnt high, crypt base, crypt top , Wnt low benefit CT and FOLFIRI, worse survival CSC (10%): serrated, poor survival Mixed (14%): Wnt high, CSC high, tubular CCS 3 (25%): poorly dif, EMT, invasion, migration and TGF-β signaling, no benefit cetuximab CIN normal (10%): CIN Wnt up (30%): conventional serrated, precursor poor survival A-type (62%): low mutation, MSS, epithelial proliferative, benefit adjuvant CT C-type (16%): mesenchymal, no benefit CT Melbourne T: 209 V: 443 128 genes Good prognosis (40%) Poor prognosis (60%): immune down/ cell signaling, ECM and focal adhesion pathways up Slide from Rodrigo Dienstmann

PIs: Justin Guinney Rodrigo Dienstmann
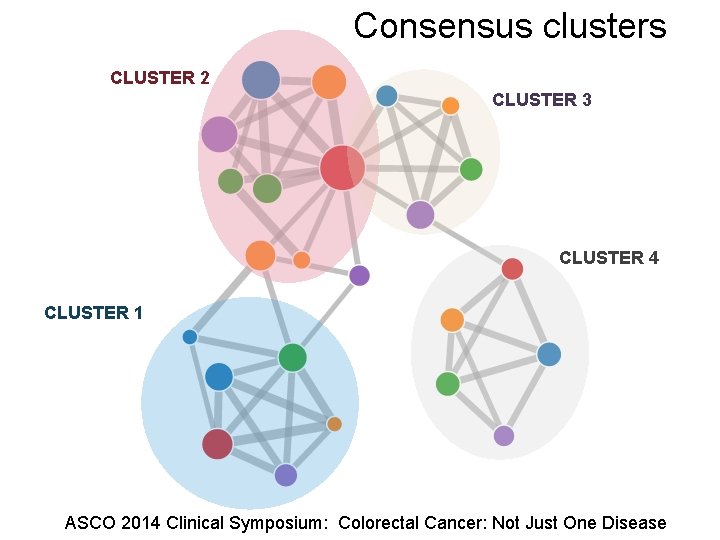
Consensus clusters CLUSTER 2 CLUSTER 3 CLUSTER 4 CLUSTER 1 ASCO 2014 Clinical Symposium: Colorectal Cancer: Not Just One Disease

Why not yet…. ? • We need studies to evaluate the benefit from extended molecular testing (beyond KRAS, NRAS, BRAF) • We need to define the molecular subtypes by gene expression • We need validated assays to move into the clinic 27
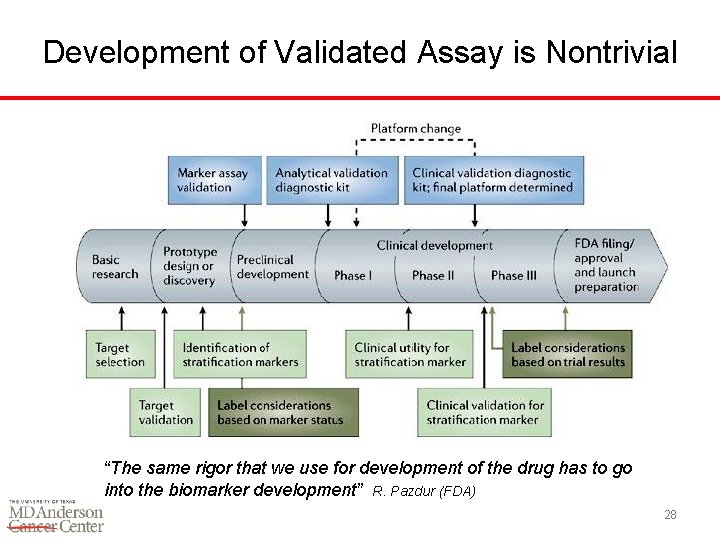
Development of Validated Assay is Nontrivial “The same rigor that we use for development of the drug has to go into the biomarker development” R. Pazdur (FDA) 28

Conclusion • Everyone should be testing for MSI and KRAS, NRAS, BRAF • We need to do the studies to demonstrate benefit of more extended molecular profiling – Low yields for actionable mutations – Need more and better novel therapies • A consensus is building for defining the subsets • The assays need to be built, and moved into clinical labs. 29
- Slides: 29