Molecular phylogenetics Xuhua Xia xxiauottawa ca http dambe

Molecular phylogenetics Xuhua Xia xxia@uottawa. ca http: //dambe. bio. uottawa. ca

Biodiversity Xuhua Xia Slide 2
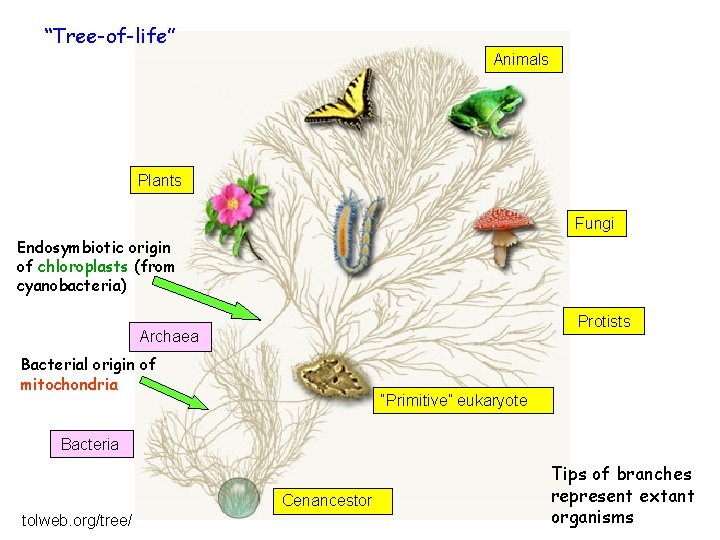
“Tree-of-life” Animals Plants Fungi Endosymbiotic origin of chloroplasts (from cyanobacteria) Protists Archaea Bacterial origin of mitochondria “Primitive” eukaryote Bacteria Cenancestor tolweb. org/tree/ Tips of branches represent extant organisms
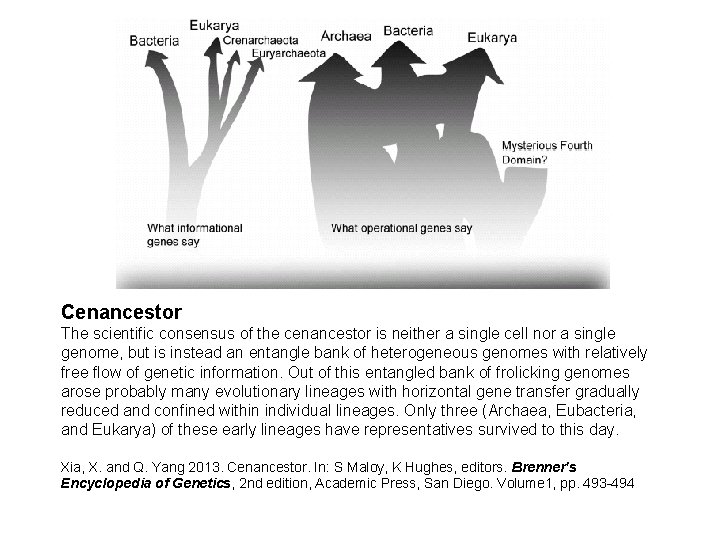
Cenancestor The scientific consensus of the cenancestor is neither a single cell nor a single genome, but is instead an entangle bank of heterogeneous genomes with relatively free flow of genetic information. Out of this entangled bank of frolicking genomes arose probably many evolutionary lineages with horizontal gene transfer gradually reduced and confined within individual lineages. Only three (Archaea, Eubacteria, and Eukarya) of these early lineages have representatives survived to this day. Xia, X. and Q. Yang 2013. Cenancestor. In: S Maloy, K Hughes, editors. Brenner's Encyclopedia of Genetics, 2 nd edition, Academic Press, San Diego. Volume 1, pp. 493 -494
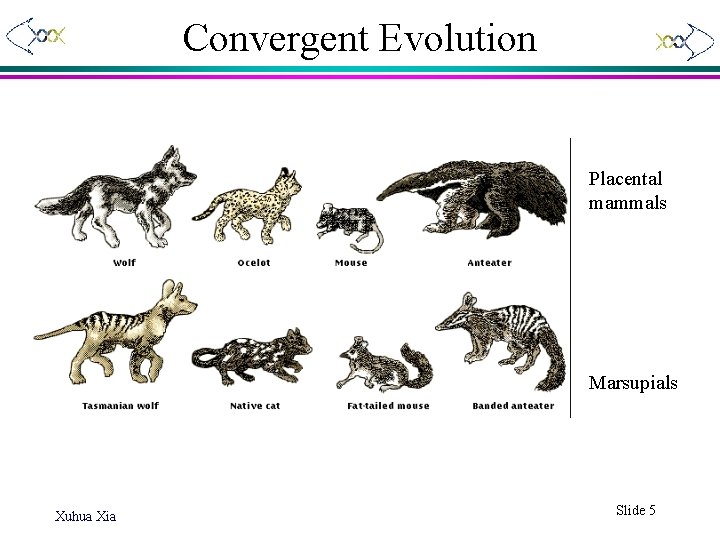
Convergent Evolution Placental mammals Marsupials Xuhua Xia Slide 5
The Story of the German Farmer The elder son of the German Farmer: Strong and Robust Immunological & Electrophoretic Diagnosis German Farmer: Strong and Robust The younger son of the German Farmer: Weak and unmanly Xuhua Xia Slide 6
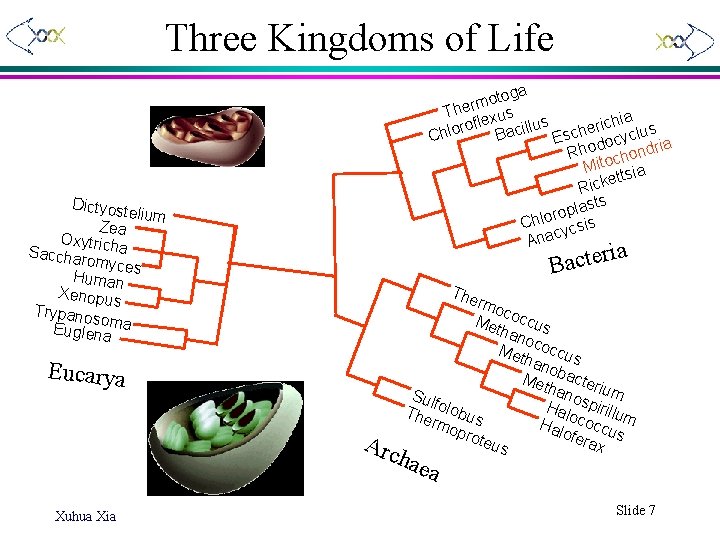
Three Kingdoms of Life toga o m r The xus hia le s f c i u o r l i e o s Bac Chl Esch docyclu ia Rho chondr Mito sia ett Rick sts a l p o r Chlo ycsis Anac Dictyost elium Zea Oxytr Sacchar icha omyces Human Xenopu s Trypano so Euglenama ria e t c a B The Eucarya Arc rmo c Met occus han o Met coccu han s o Met bacter ium han Sul o s f Hal pirillu The olobus oco m rmo H c c a u lofe pro teus rax s hae a Xuhua Xia Slide 7

Where have all the whales gone? • Facts: – North Atlantic minke whales were not taken for commercial purposes under IWC resolutions since 1986 – Fin whales have not been hunted legally since 1986 – Hunting of humpback whales has been prohibited since 1966 – Birth rate was found to be higher than death rate • Why not more whales? • Illegal hunting? Minke whele (North Atlantic) Sample #19 a Sample #9 Sample #15 Sample #19 b Humpback whale Sample #41 Sample #3 Sample #11 Sample WS 4 • Forensics Xuhua Xia Fin whale Slide 8
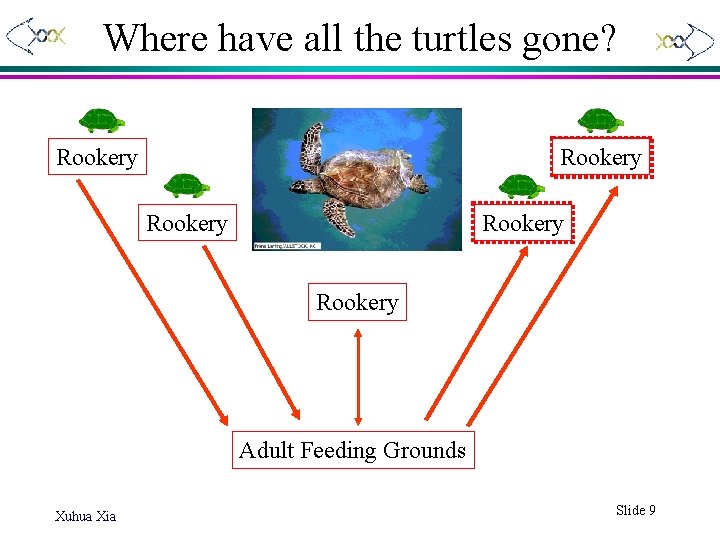
Where have all the turtles gone? Rookery Rookery Adult Feeding Grounds Xuhua Xia Slide 9
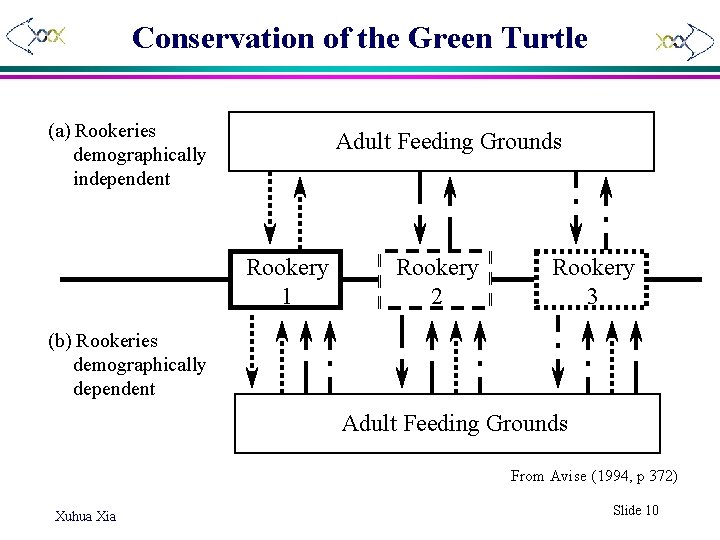
Conservation of the Green Turtle (a) Rookeries demographically independent Adult Feeding Grounds Rookery 1 Rookery 2 Rookery 3 (b) Rookeries demographically dependent Adult Feeding Grounds From Avise (1994, p 372) Xuhua Xia Slide 10
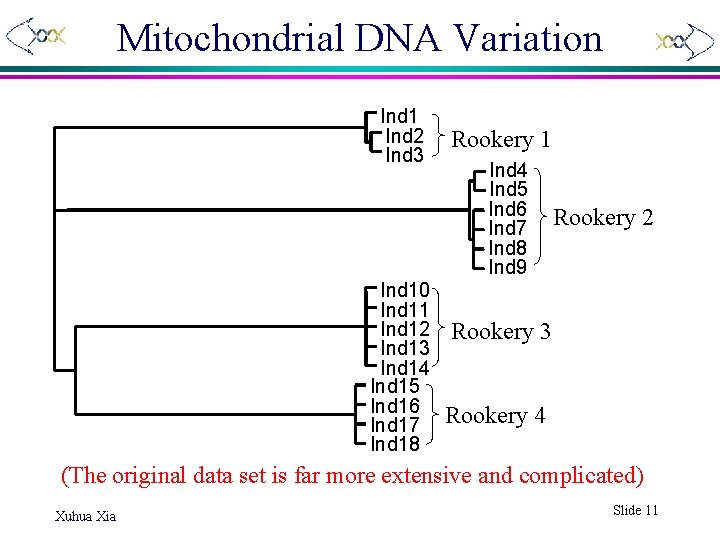
Mitochondrial DNA Variation Ind 1 Ind 2 Ind 3 Ind 10 Ind 11 Ind 12 Ind 13 Ind 14 Ind 15 Ind 16 Ind 17 Ind 18 Rookery 1 Ind 4 Ind 5 Ind 6 Ind 7 Ind 8 Ind 9 Rookery 2 Rookery 3 Rookery 4 (The original data set is far more extensive and complicated) Xuhua Xia Slide 11
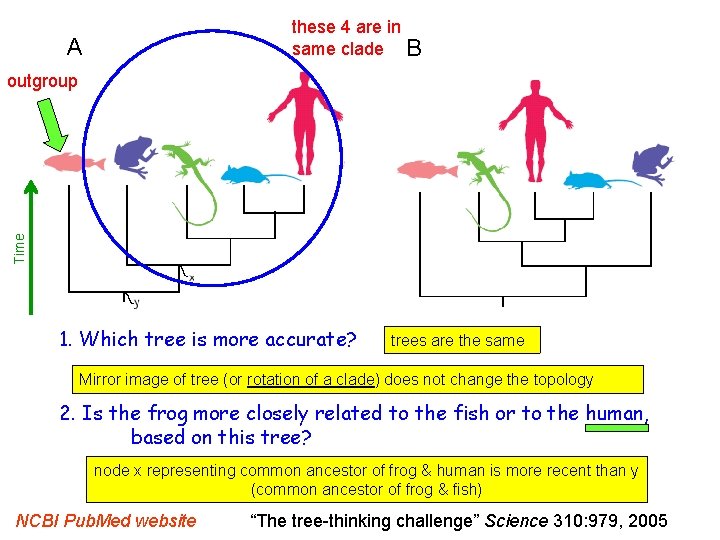
these 4 are in same clade B A Time outgroup 1. Which tree is more accurate? trees are the same Mirror image of tree (or rotation of a clade) does not change the topology 2. Is the frog more closely related to the fish or to the human, based on this tree? node x representing common ancestor of frog & human is more recent than y (common ancestor of frog & fish) NCBI Pub. Med website “The tree-thinking challenge” Science 310: 979, 2005
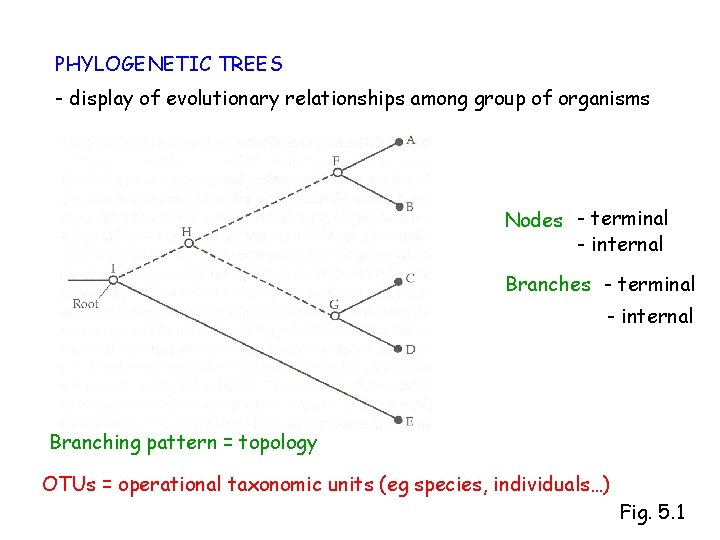
PHYLOGENETIC TREES - display of evolutionary relationships among group of organisms Nodes - terminal - internal Branching pattern = topology OTUs = operational taxonomic units (eg species, individuals…) Fig. 5. 1
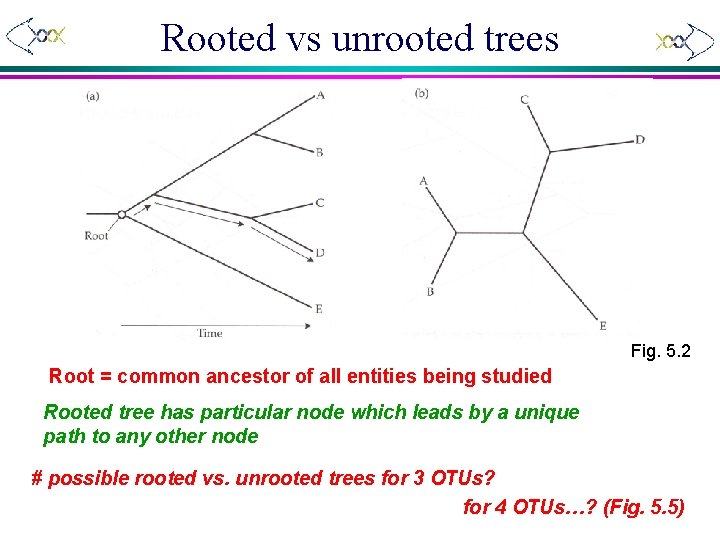
Rooted vs unrooted trees Fig. 5. 2 Root = common ancestor of all entities being studied Rooted tree has particular node which leads by a unique path to any other node # possible rooted vs. unrooted trees for 3 OTUs? for 4 OTUs…? (Fig. 5. 5)

Scaled vs unscaled branches Branch length proportional to number of changes Xuhua Xia Fig. 155. 3 Slide

True vs. inferred trees - only 1 true tree … … but usually must deal with inferred trees (based on certain data set and method of tree reconstruction) Score “similarities” - shared ancestral features - shared derived features - homoplasies (convergences, parallelism, reversals) ie. similarities of traits for reasons other than common ancestry
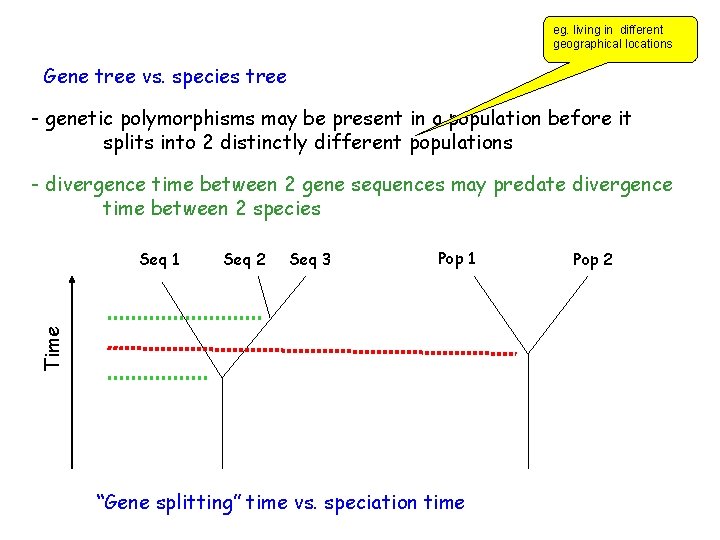
eg. living in different geographical locations Gene tree vs. species tree - genetic polymorphisms may be present in a population before it splits into 2 distinctly different populations - divergence time between 2 gene sequences may predate divergence time between 2 species Seq 2 Seq 3 Pop 1 Time Seq 1 “Gene splitting” time vs. speciation time Pop 2

Gene tree vs species tree -Genetic polymorphisms may be present in a population before it splits into 2 distinctly different populations - divergence time between 2 genes sequences may predate divergence time between 2 species - changes in DNA sequences can occur before or after speciation Gene tree may not always reflect species tree Xuhua Xia Fig. 5. 6 Slide 18

Phylogenetic tree reconstruction • Distance-based methods • Maximum parsimony methods • Maximum likelihood methods • Bayesian inference Xuhua Xia Slide 19

Distance-based methods • Objectives – Grasp the basic concepts distance-based tree-building algorithms – Learn the least-squares criterion and the minimum evolution criterion and how to use them to construct a tree • Distance-based methods – Genetic distance: generally defined as the number of substitutions per site. • • • JC 69 distance K 80 distance TN 84 distance F 84 distance TN 93 distance Log. Det distance – Tree-building algorithms (UPGMA): • • Xuhua Xia UPGMA Neighbor-joining Fitch-Margoliash Fast. ME Slide 20

Calculation of KJC 69 t AACGACGATCG: Species 1 t AACGACGATCG: Species 2 The time is 2 t between Species 1 to Species 2 Sp 1: AAG CCT CGG GGC CCT TAT TTG || | ||| ||| || Sp 2: AAT CTC CGG GGC CTC TAT TTT p = 6/24 = 0. 25 K = 0. 304099 Genetic distances are scaled to be the number of substitutions per site. Xuhua Xia Slide 21

Numerical Illustration Sp 1: AAG CCT CGG GGC CCT TAT TTG || | ||| ||| || Sp 2: AAT CTC CGG GGC CTC TAT TTT What are P and Q? P = 4/24, Q = 2/24 Comparison of distances: P = 0. 25 Poisson P = -ln(1 -p) = 0. 288 KJC 69 = 0. 304099 KK 80 = 0. 3150786 Xuhua Xia Slide 22

A Star Tree (Completely Unresolved Tree) Human Chimpanzee Gorilla Orangutan Gibbon Xuhua Xia Slide 23

Genetic Distance Matrix 10 20 30 40 50 60 ----|----|----|----|----|----|-human CAUGCUACUCCACACACCAAGCUAUCUAGCCUCCCCAAUCCAAAACAUUAAACACUUU. . . chimpanzee CAUACUACUCCACACACCAAACUACCUAGCCUCCCCAAUCCAAAAUAAACAUCAAACACUUU. . . gorilla CAUACUACUCCACACACCAAAUCAUCUAGCCUCCCCAGUCCAGAACACUGAAAAUUUU. . . orangutan CAUACCACUCCACACCCUAUACCAUCCAACUUCCCCUAUCCGAAACAAAUACAAAACACUUC. . . gibbon CAUACUACUCCAUACACCAAAUUAUCCAACUCCCCCAAUCCAGAAUAAACACCGACCAUCUU. . . *** * ****** ** * * * * Matrix of Genetic distances (Dij): Human Chimp Gorilla Orang Gibbon Xuhua Xia Chimp 0. 015 Gorilla 0. 045 0. 030 Orang 0. 143 0. 126 0. 092 0. 179 Gibbon 0. 198 0. 179 Slide 24
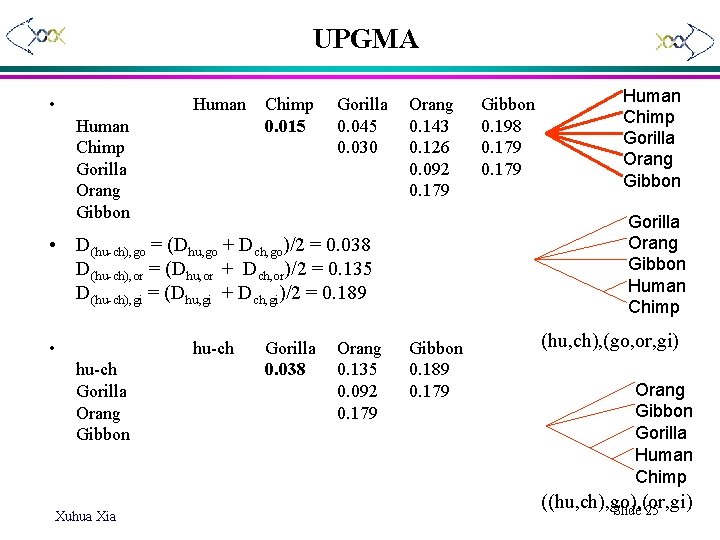
UPGMA • Human Chimp Gorilla Orang Gibbon Chimp 0. 015 Gorilla 0. 045 0. 030 Orang 0. 143 0. 126 0. 092 0. 179 hu-ch Gorilla Orang Gibbon Xuhua Xia Gorilla 0. 038 Orang 0. 135 0. 092 0. 179 Human Chimp Gorilla Orang Gibbon Human Chimp • D(hu-ch), go = (Dhu, go + Dch, go)/2 = 0. 038 D(hu-ch), or = (Dhu, or + Dch, or)/2 = 0. 135 D(hu-ch), gi = (Dhu, gi + Dch, gi)/2 = 0. 189 • Gibbon 0. 198 0. 179 Gibbon 0. 189 0. 179 (hu, ch), (go, or, gi) Orang Gibbon Gorilla Human Chimp ((hu, ch), go), (or, gi) Slide 25

UPGMA • Human Chimp Gorilla Orang Gibbon • Gorilla 0. 045 0. 030 Orang 0. 143 0. 126 0. 092 0. 179 D(hu-ch-go), or = (Dhu, or + Dch, or + Dgo, or)/3 = 0. 120 D(hu-ch-go), gi = (Dhu, gi + Dch, gi +Dgo, gi)/3 = 0. 185 • hu-ch-go Orangutan Gibbon • Chimp 0. 015 Orang 0. 120 Gibbon 0. 185 0. 179 Gibbon 0. 198 0. 179 Orang Gibbon Gorilla Human Chimp Gibbon Orang Gorilla Human Chimp (((hu, ch), go), or), gi) D(hu-ch-go-or), gi = (Dhu, gi + Dch, gi +Dgo, gi + Dor, gi)/4 = 0. 184 Xuhua Xia Slide 26

Phylogenetic Relationship from UPGMA • Human Chimp 0. 015 Gorilla 0. 045 0. 030 Orang 0. 143 0. 126 0. 092 0. 179 hu-ch Gorilla 0. 038 Orang 0. 135 0. 092 0. 179 Gibbon 0. 189 0. 179 Human Chimp Gorilla Orang Gibbon • hu-ch-go Orang Gibbon Xuhua Xia hu-ch-go Orang 0. 120 0. 179 Gibbon 0. 198 0. 179 Gibbon 0. 185 Slide 27
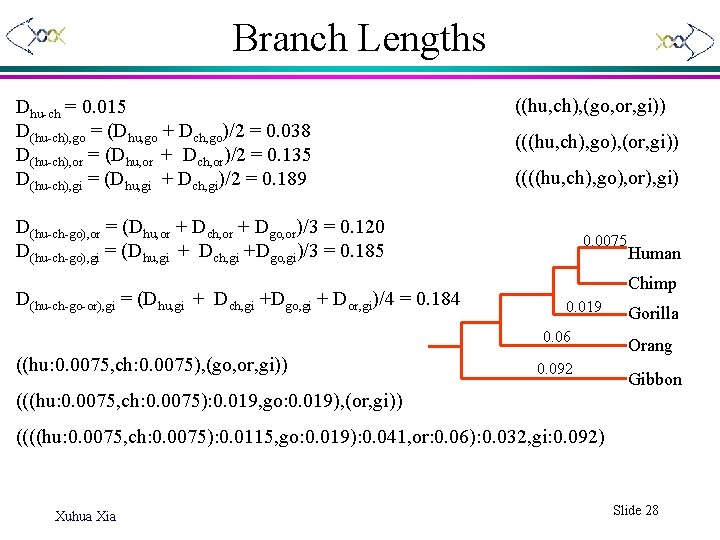
Branch Lengths Dhu-ch = 0. 015 D(hu-ch), go = (Dhu, go + Dch, go)/2 = 0. 038 D(hu-ch), or = (Dhu, or + Dch, or)/2 = 0. 135 D(hu-ch), gi = (Dhu, gi + Dch, gi)/2 = 0. 189 ((hu, ch), (go, or, gi)) (((hu, ch), go), (or, gi)) ((((hu, ch), go), or), gi) D(hu-ch-go), or = (Dhu, or + Dch, or + Dgo, or)/3 = 0. 120 D(hu-ch-go), gi = (Dhu, gi + Dch, gi +Dgo, gi)/3 = 0. 185 D(hu-ch-go-or), gi = (Dhu, gi + Dch, gi +Dgo, gi + Dor, gi)/4 = 0. 184 0. 0075 Chimp 0. 019 0. 06 ((hu: 0. 0075, ch: 0. 0075), (go, or, gi)) Human 0. 092 Gorilla Orang Gibbon (((hu: 0. 0075, ch: 0. 0075): 0. 019, go: 0. 019), (or, gi)) ((((hu: 0. 0075, ch: 0. 0075): 0. 0115, go: 0. 019): 0. 041, or: 0. 06): 0. 032, gi: 0. 092) Xuhua Xia Slide 28
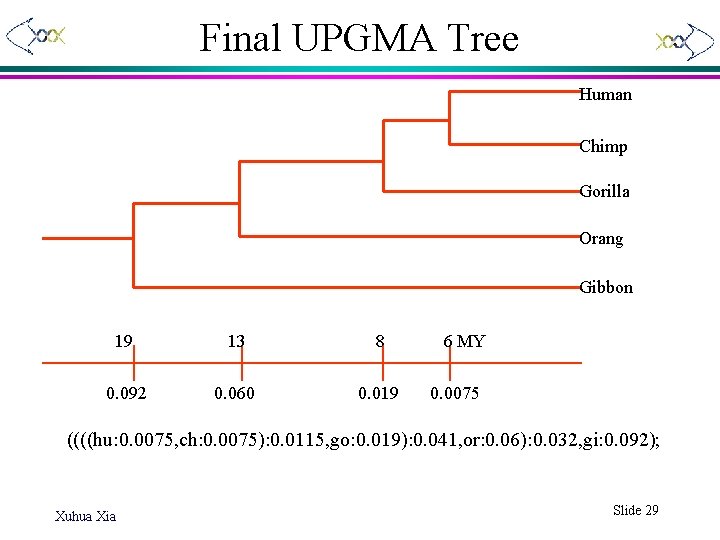
Final UPGMA Tree Human Chimp Gorilla Orang Gibbon 19 13 8 0. 092 0. 060 0. 019 6 MY 0. 0075 ((((hu: 0. 0075, ch: 0. 0075): 0. 0115, go: 0. 019): 0. 041, or: 0. 06): 0. 032, gi: 0. 092); Xuhua Xia Slide 29

Distance-based method • Distance matrix • Tree-building algorithms – UPGMA – Neighbor-joining – Fitch-Margoliash – Fast. ME • Criterion-based methods: the least squares method – Branch-length estimation – Tree-selection criterion Xuhua Xia Slide 30
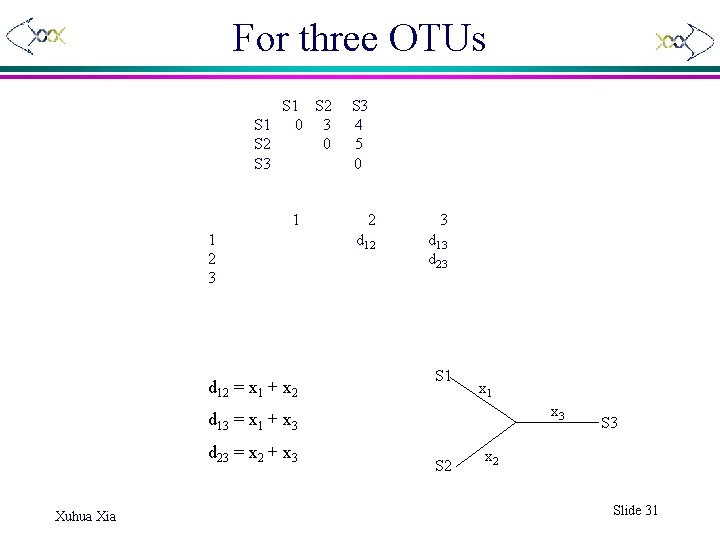
For three OTUs S 1 S 2 S 1 0 3 S 2 0 S 3 1 1 2 3 d 12 = x 1 + x 2 S 3 4 5 0 2 3 d 12 d 13 d 23 S 1 x 3 d 13 = x 1 + x 3 d 23 = x 2 + x 3 Xuhua Xia S 2 S 3 x 2 Slide 31

Least-square method 4 Sp 1 Sp 2 Sp 3 Sp 4 d’ 12 = x 1 + x 2 0. 3 0. 4 0. 5 0. 4 0. 6 d’ 13 = x 1 + x 5+ x 3 d’ 14 = x 1 + x 5 + x 4 4 d’ 23 = x 2 + x 5 + x 3 Sp 1 d’ 24 = x 2 + x 5 + x 4 Sp 2 d 12 Sp 3 d 13 d 23 Sp 4 d 14 d 24 1 d’ 34 = x 3 + x 4 d 34 x 3 x 1 3 x 5 2 Xuhua Xia x 2 x 4 4 Slide 32

The LS method in linear regression X Y R(Residual) 3 11. 5 a+b*3 – 11. 5 2 7. 5 a+b*2 – 7. 5 1 5 a+b*1 – 5 4 14 a+b*4 – 14 RSS = 0 means a perfect fit of the linear model to the data. A large RSS means a poor fit. Y=a+bx Slide 33

Least-square method 1 x 3 x 1 3 x 5 2 x 4 4 d’ 12 = x 1 + x 2 (d 12 - d’ 12)2= [d 12 – (x 1 + x 2)]2 d’ 13 = x 1 + x 5+ x 3 (d 13 - d’ 13)2 = [d 13 – (x 1 + x 5+ x 3)]2 d’ 14 = x 1 + x 5 + x 4 (d 14 - d’ 14)2 = [d 14 – (x 1 + x 5 + x 4)]2 d’ 23 = x 2 + x 5 + x 3 (d 23 - d’ 23)2 = [d 23 – (x 2 + x 5 + x 3)]2 d’ 24 = x 2 + x 5 + x 4 (d 24 - d’ 24)2 = [d 24 – (x 2 + x 5 + x 4)]2 d’ 34 = x 3 + x 4 (d 34 - d’ 34)2 = [d 34 – (x 3 + x 4)]2 Least-squares method: Find xi values that minimize SS Xuhua Xia Slide 34
![Least-squares method SS = [d 12 – (x 1 + x 2)]2 + [d Least-squares method SS = [d 12 – (x 1 + x 2)]2 + [d](http://slidetodoc.com/presentation_image_h2/95ab863ca9a99fb141eade885725b0b3/image-35.jpg)
Least-squares method SS = [d 12 – (x 1 + x 2)]2 + [d 13 – (x 1 + x 5+ x 3)]2 + [d 14 – (x 1 + x 5 + x 4)]2 + [d 23 – (x 2 + x 5 + x 3)]2+ [d 24 – (x 2 + x 5 + x 4)]2+ [d 34 – (x 3 + x 4)]2 Take the partial derivative of SS with respective to xi, we have SS/ x 1 : = -2 d 12 + 6 x 1 + 2 x 2 - 2 d 13 + 4 x 5 + 2 x 3 - 2 d 14 + 2 x 4 SS/ x 2 : = -2 d 12 + 2 x 1 + 6 x 2 - 2 d 23 + 4 x 5 + 2 x 3 - 2 d 24 + 2 x 4 SS/ x 3 : = -2 d 13 + 2 x 1 + 4 x 5 + 6 x 3 - 2 d 23 + 2 x 2 - 2 d 34 + 2 x 4 SS/ x 4 : = -2 d 14 + 2 x 1 + 4 x 5 + 6 x 4 - 2 d 24 + 2 x 2 - 2 d 34 + 2 x 3 SS/ x 5 : = -2 d 13 + 4 x 1 + 8 x 5 + 4 x 3 - 2 d 14 + 4 x 4 - 2 d 23 + 4 x 2 - 2 d 24 Setting these partial derivatives to 0 and solve for xi, we have x 1 = d 13/4 + d 12/2 - d 23/4 + d 14/4 - d 24/4 x 2 = d 12/2 - d 13/4 + d 23/4 - d 14/4 + d 24/4, x 3 = d 13/4 + d 23/4 + d 34/2 - d 14/4 - d 24/4, x 4 = d 14/4 - d 13/4 - d 23/4 + d 34/2 + d 24/4, x 5 = - d 12/2 + d 23/4 - d 34/2 + d 14/4 + d 24/4 + d 13/4 Xuhua Xia Slide 35

Least-squares method x 1 = d 13/4 + d 12/2 - d 23/4 + d 14/4 - d 24/4 x 2 = d 12/2 - d 13/4 + d 23/4 - d 14/4 + d 24/4, x 3 = d 13/4 + d 23/4 + d 34/2 - d 14/4 - d 24/4, x 4 = d 14/4 - d 13/4 - d 23/4 + d 34/2 + d 24/4, x 5 = - d 12/2 + d 23/4 - d 34/2 + d 14/4 + d 24/4 + d 13/4 4 Sp 1 Sp 2 Sp 3 Sp 4 0. 3 0. 4 0. 5 0. 4 0. 6 x 1 = 0. 075 x 2 = 0. 225 x 3 = 0. 275 x 4 = 0. 325 x 5 = 0. 025 Xuhua Xia 1 x 3 x 1 3 x 5 2 x 4 4 Slide 36
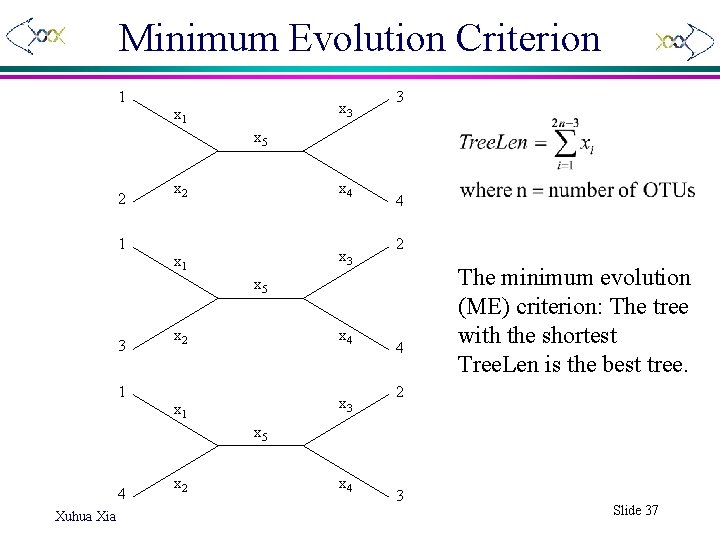
Minimum Evolution Criterion 1 x 3 x 1 3 x 5 2 1 x 2 x 4 x 1 x 3 4 2 x 5 3 1 x 2 x 4 x 1 x 3 4 The minimum evolution (ME) criterion: The tree with the shortest Tree. Len is the best tree. 2 x 5 4 Xuhua Xia x 2 x 4 3 Slide 37

Maximum Parsimony (MP) Method • Mapping character state changes to alternative topologies • Apply the maximum parsimony criterion to choose the best tree. • Efficient dynamic programming algorithm developed by Walter Fitch and David Sankoff • The only method with branch-and-bound search • Use only informative sites to discriminate among alternative topologies • Problems – Long-branch attraction – Failure to account to multiple substitutions Xuhua Xia Slide 38

Informative sites • A site with at least two different characters each being represented by at least two OTUs. • Meaningful only in Fitch Parsimony where all nucleotides or amino acids are equally likely to replace each other. • Sankoff parsimony introduces the step matrix and can use information in a "non-informative" site for discriminate among alternative topologies, e. g. , when transitions and transversions are associated with different costs. Xuhua Xia Slide 39
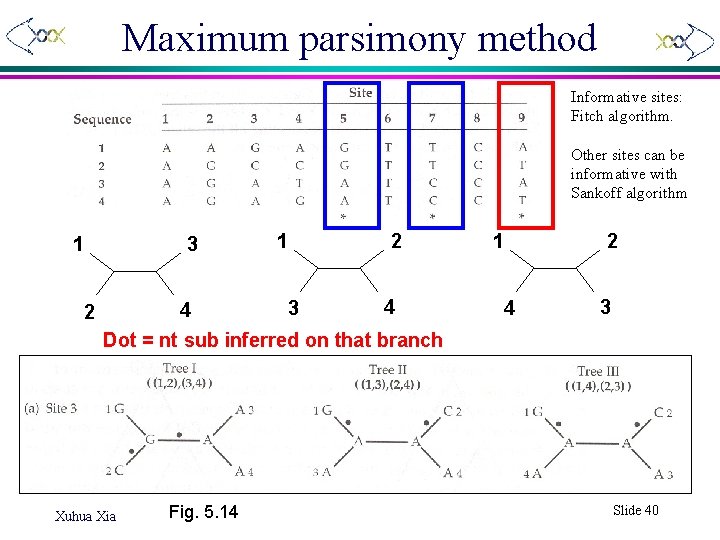
Maximum parsimony method Informative sites: Fitch algorithm. Other sites can be informative with Sankoff algorithm 1 2 3 1 2 4 3 4 Dot = nt sub inferred on that branch Xuhua Xia Fig. 5. 14 1 4 2 3 Slide 40
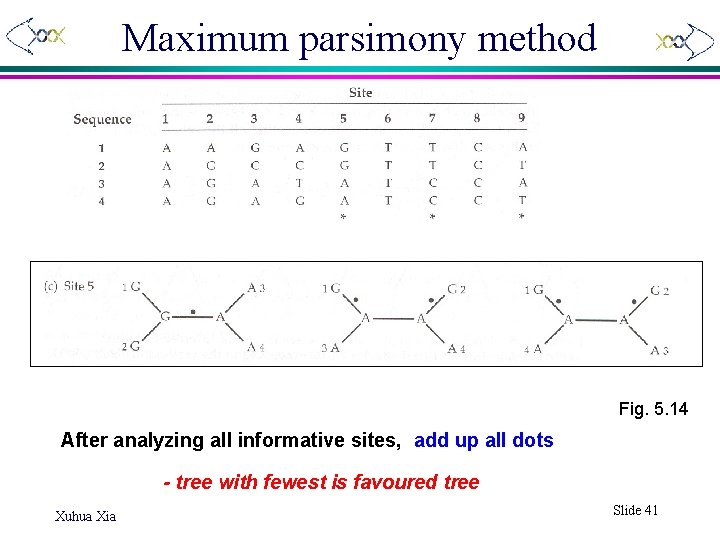
Maximum parsimony method Fig. 5. 14 After analyzing all informative sites, add up all dots - tree with fewest is favoured tree Xuhua Xia Slide 41

Computing N 1 • • Each node is represented by a set of characters, with the terminal nodes (leaves) each represented by a set containing a single character. The MP method traverses through each internal node, starting from the node closest to the leaves. – If two sets of the two daughter nodes have an empty intersection, then the node will be represented by the union of the two daughter sets, otherwise the node will be represented by the intersection. – Once the operation reaches the root, then the number of union operations is the minimum number of changes needed to map the site to the tree. Xuhua Xia Slide 42

Tree Length • Site 1 requires four union operations • Sites 3, 5, and 8 each require only one union operation • Sites 6 and 7, which are polymorphic with two nucleotide states but not informative, will require one change for any topology. • The tree length for the topology above: 4+(1+1+1)+(1+1) = 9 Xuhua Xia Slide 43

Criteria for a good estimator • Unbiased • Efficient • Consistent Xuhua Xia Slide 44

Inconsistency (Felsenstein, 1978) A B Model tree p p q C q q D A Rates or Branch lengths p >> q B MP tree C Wrong D • With more data the certainty that parsimony will give the wrong tree increases - so that parsimony is statistically inconsistent • It is now recognised that long-branch attraction is one of the most serious problems in phylogenetic inference Xuhua Xia Slide 45

Maximum likelihood Method • Likelihood L of a tree is the probability of observing the data given the tree L = P(data|tree) • Find the tree with the highest L value • Results depends on model of nucleotide substitution Xuhua Xia Slide 46
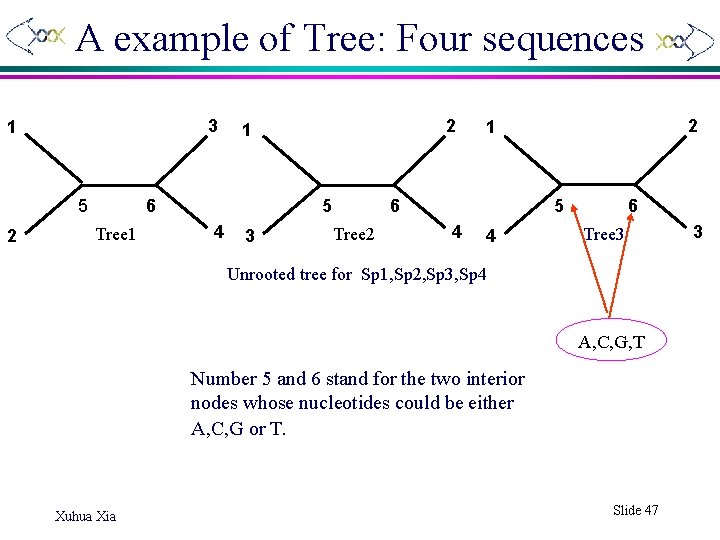
A example of Tree: Four sequences 3 1 5 2 6 Tree 1 2 1 5 4 3 6 Tree 2 2 1 5 4 4 6 Tree 3 Unrooted tree for Sp 1, Sp 2, Sp 3, Sp 4 A, C, G, T Number 5 and 6 stand for the two interior nodes whose nucleotides could be either A, C, G or T. Xuhua Xia Slide 47 3
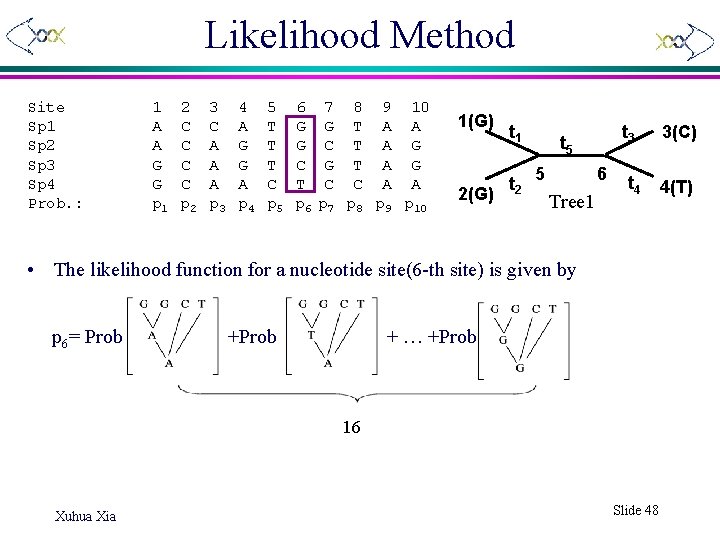
Likelihood Method Site Sp 1 Sp 2 Sp 3 Sp 4 Prob. : 1 A A G G p 1 2 C C p 2 3 C A A A p 3 4 A G G A p 4 5 T T T C p 5 6 G G C T p 6 7 8 9 10 G T A A C T A G G T A G C C A A p 7 p 8 p 9 p 10 1(G) 2(G) t 1 t 2 t 3 t 5 5 6 Tree 1 t 4 • The likelihood function for a nucleotide site(6 -th site) is given by p 6= Prob + … +Prob 16 Xuhua Xia Slide 48 3(C) 4(T)

Calculation of Likelihood where P(A), P(T), P(C), P(G) are empirical nucleotide frequencies satisfying P(A)+P(T)+P(C)+P(G)=1, Pii(t) and Pij(t) are given by JC 69 when • ln. LTree 1= ln(p 1)+ln(p 2)+…+ln(p 10) • Calculate ln. LTree 2, ln. LTree 3 similarly. • We choose the tree which has the highest ln. L i. e. Max(ln. LTree 1, ln. LTree 2 , ln. LTree 3 ). 1(G) 2(G) Xuhua Xia t 1 t 2 t 3 t 5 5 6 Tree 1 t 4 3(C) 4(T) Slide 49

Problems with ML method The ML method is strictly data-based. If we sampled 6 fish all being males, then our estimation of p is 6/6 = 1. Xuhua Xia Slide 50
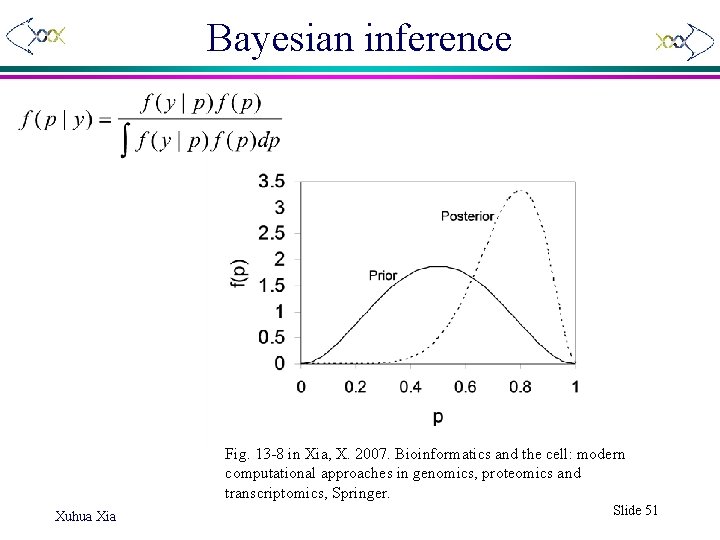
Bayesian inference Fig. 13 -8 in Xia, X. 2007. Bioinformatics and the cell: modern computational approaches in genomics, proteomics and transcriptomics, Springer. Xuhua Xia Slide 51

Tree quality assessment • Re-sampling methods for subtree support – Bootstrap – Jackknife (delete-half jackknife) – Monte-Carlo method • Significant tests for alternative trees – Distance-based method – Maximum parsimony method – Maximum likelihood method Xuhua Xia Slide 52
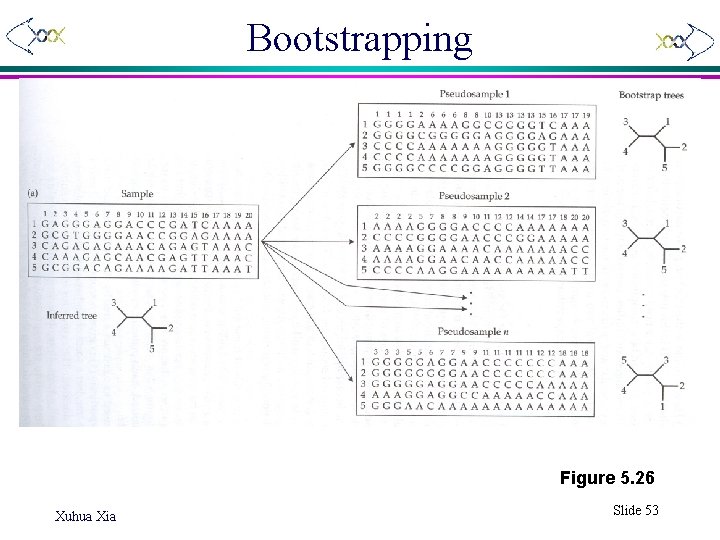
Bootstrapping Figure 5. 26 Xuhua Xia Slide 53
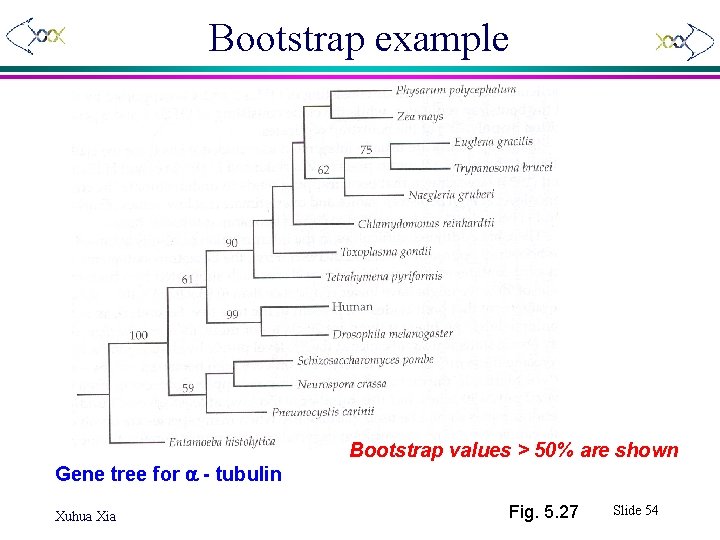
Bootstrap example Gene tree for a - tubulin Xuhua Xia Bootstrap values > 50% are shown Fig. 5. 27 Slide 54
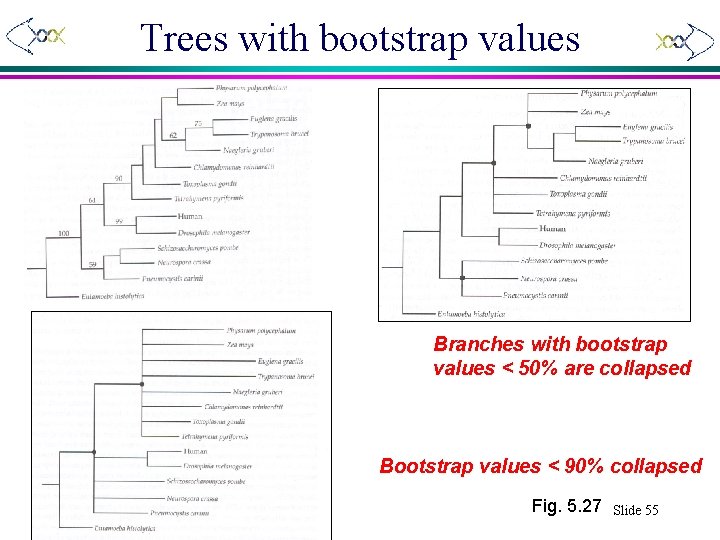
Trees with bootstrap values Branches with bootstrap values < 50% are collapsed Bootstrap values < 90% collapsed Xuhua Xia Fig. 5. 27 Slide 55

Phylogenetic Hypothesis Testing: MP Tree 1 Tree 2 AAGGT GTGGT GAGGT Frog AAGGT Pigeon GTGGC Eagle GTGGT Elephant GAAAC GAAAT Lion GAAAT Frog Eagle Pigeon Lion Elephant 1 2 34 5 ACCCAAAGGCCTT GCCCTAAGGCCTC GCCCAAAAACCTT GCCCAAAAACCTC Tree 1 Tree 2 1 1 Xuhua Xia 1 2 11 22 AAGGT GAGGT GAGGC Frog AAGGT Lion GAAAT Eagle GTGGT Elephant GAAAC Pigeon 2 1 Slide 56 GTGGC

Test Phylogenetic Hypotheses: ML • Maximum likelihood-based method – Kishino-Hasegawa’s RELL test • • ln. L 1 from Tree 1 ln. L 2 from Tree 2 D = ln. L 1 – ln. L 2 Var(D) obtained by resampling – DNLML test Xuhua Xia Slide 57

Things to consider • Data: – – – Mixture of paralogous and orthologous genes Gene conversion Convergent evolution Horizontal gene transfer Why mt. DNA is popular in molecular phylogenetics? • Too little or too much variation (Substitution saturation): choose rapidly evolving sequences for recently diverged taxa and highly conserved genes for resolving deep phylogenies. • Sequence alignment (circularity between alignment and phylogenetics) • Substitution models • Tree selection criteria • Tree search algorithms A phylogenetic tree represents a hypothesized phylogenetic relationship among ingroup species, and published trees often contain errors. (P. 217) Xuhua Xia Slide 58

What gene to use? If comparing very divergent organisms: - need ones “universally” present and highly conserved otherwise possible errors in alignment, multiple subs… eg. ribosomal RNAs: the universal yardstick translation elongation factors, ribosomal proteins glycolytic pathway enzymes

Testing phylogenetic hypotheses Out. Group “Reptilian” Mammal Bird Out. Group Mammal Bird “Reptilian” Xuhua Xia “Using molecular sequence data to determine the phylogenetic relationships of the major groups of organisms has yielded some spectacular successes but has also thrown up some conundrums. One such is the relationship of birds to the rest of the tetrapods. Morphological data and most molecular studies have placed the birds closer to the crocodiles than to any other tetrapod group, but analysis of sequence data from 18 S ribosomal RNA (r. RNA) has persistently allied the birds more closely to the mammals. There have been several attempts to account for this niggling doubt, and Xia et al. (2003, Syst. Biol. 52: 283) now show that the discrepancy arose because of methodological flaws in the analysis of 18 S r. RNA data, which caused, among other things, misalignment of sequences from the different taxa. When structure-based alignment is carried out, the resulting phylogeny matches those obtained by other means, with the birds allied to the crocodiles via a common reptilian ancestor. ” Science 301: 279 (Editors’ Choice) Slide 60
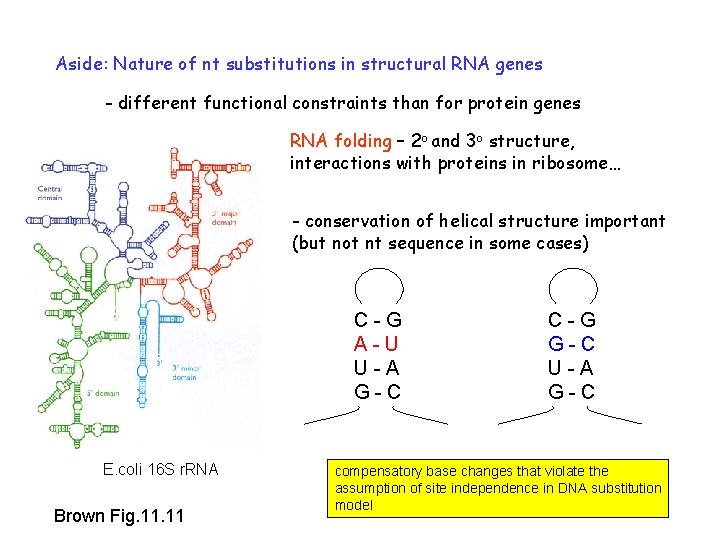
Aside: Nature of nt substitutions in structural RNA genes - different functional constraints than for protein genes RNA folding – 2 o and 3 o structure, interactions with proteins in ribosome… - conservation of helical structure important (but not nt sequence in some cases) C-G A-U U-A G-C E. coli 16 S r. RNA Brown Fig. 11 C-G G-C U-A G-C compensatory base changes that violate the assumption of site independence in DNA substitution model
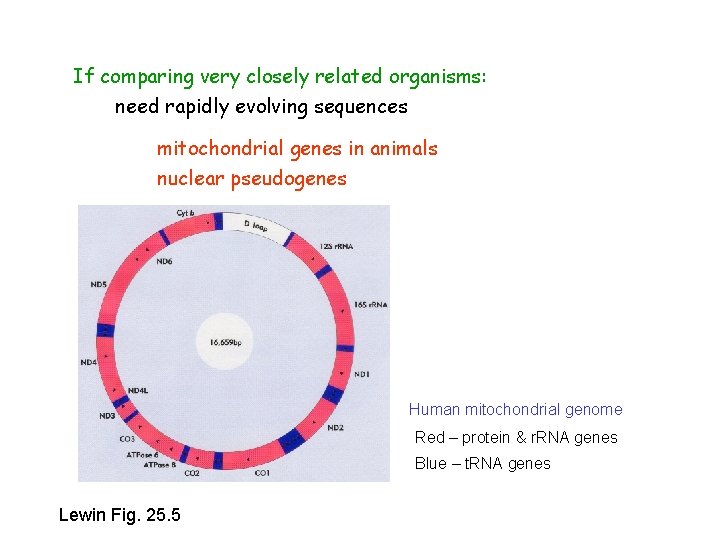
If comparing very closely related organisms: need rapidly evolving sequences mitochondrial genes in animals nuclear pseudogenes Human mitochondrial genome Red – protein & r. RNA genes Blue – t. RNA genes Lewin Fig. 25. 5
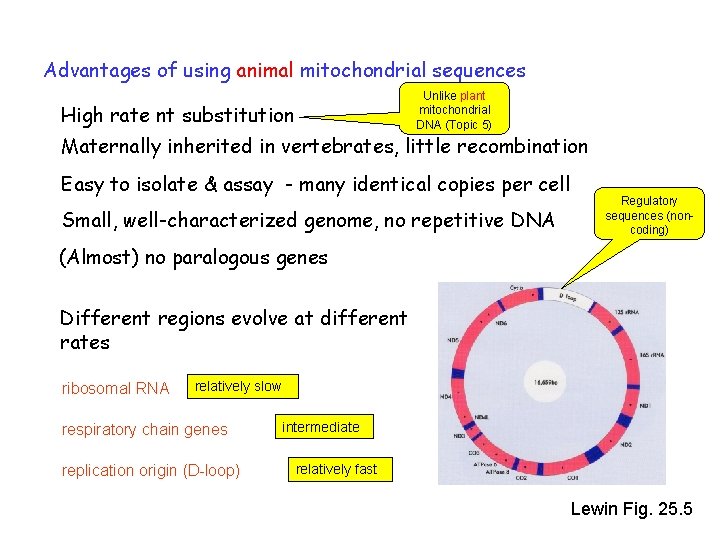
Advantages of using animal mitochondrial sequences Unlike plant mitochondrial DNA (Topic 5) High rate nt substitution Maternally inherited in vertebrates, little recombination Easy to isolate & assay - many identical copies per cell Small, well-characterized genome, no repetitive DNA Regulatory sequences (noncoding) (Almost) no paralogous genes Different regions evolve at different rates ribosomal RNA relatively slow respiratory chain genes replication origin (D-loop) intermediate relatively fast Lewin Fig. 25. 5

“Barcode of life” Quick, inexpensive way to identify species using short DNA sequences that are universally present among organisms of interest For animal kingdom, part of mitochondrial COI gene is PCR-amplified with primers that map to conserved coding region & then sequenced “[The authors] imagine a day when a handheld scanner (similar to a GPS device) will link to a database of the barcodes of all species. Then, by inserting a snippet of tissue into the scanner, anyone can get an instant identification of a creature or plant. ” Paul Hebert U. Guelph Stoekle & Hebert Scient Amer Oct. 2008
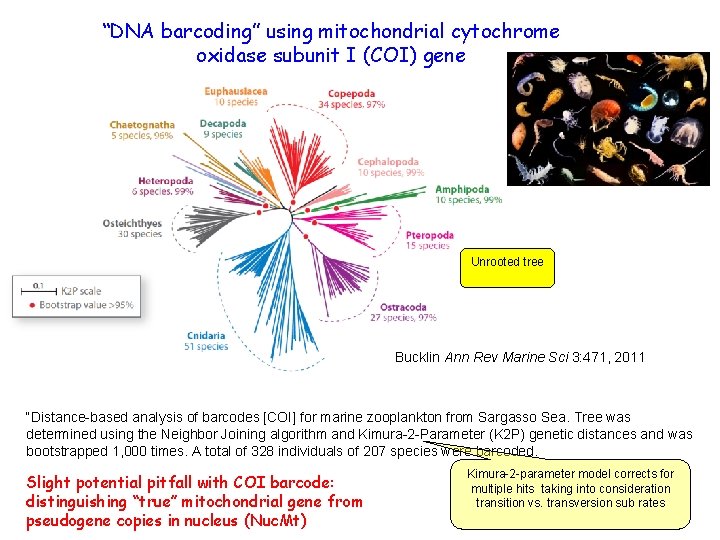
“DNA barcoding” using mitochondrial cytochrome oxidase subunit I (COI) gene Unrooted tree Bucklin Ann Rev Marine Sci 3: 471, 2011 “Distance-based analysis of barcodes [COI] for marine zooplankton from Sargasso Sea. Tree was determined using the Neighbor Joining algorithm and Kimura-2 -Parameter (K 2 P) genetic distances and was bootstrapped 1, 000 times. A total of 328 individuals of 207 species were barcoded. Slight potential pitfall with COI barcode: distinguishing “true” mitochondrial gene from pseudogene copies in nucleus (Nuc. Mt) Kimura-2 -parameter model corrects for multiple hits taking into consideration transition vs. transversion sub rates
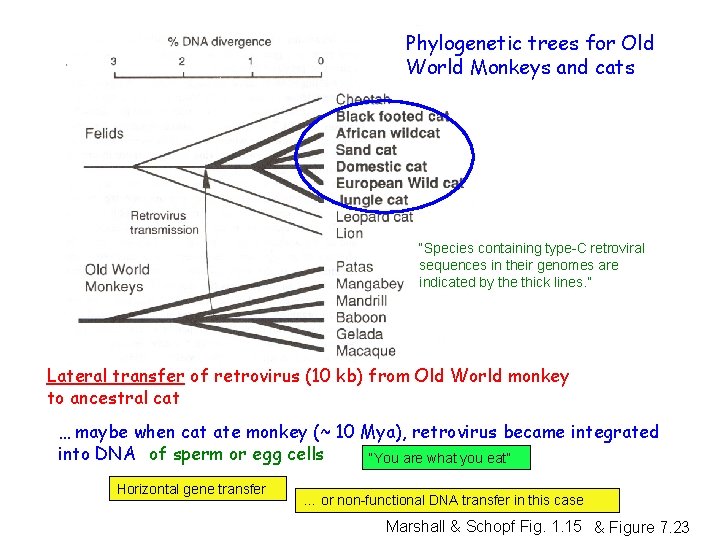
Phylogenetic trees for Old World Monkeys and cats “Species containing type-C retroviral sequences in their genomes are indicated by the thick lines. ” Lateral transfer of retrovirus (10 kb) from Old World monkey to ancestral cat … maybe when cat ate monkey (~ 10 Mya), retrovirus became integrated into DNA of sperm or egg cells “You are what you eat” Horizontal gene transfer … or non-functional DNA transfer in this case Marshall & Schopf Fig. 1. 15 & Figure 7. 23
- Slides: 66