Molecular Modeling The Computer is the Lab Niels

Molecular Modeling: The Computer is the Lab Niels Johan Christensen IGM/Bioinorganic Chemistry/NP 3 centre

Overview • Brief intro to molecular modeling • Molecular modeling at the NP 3 centre: Application to novel insulin complexes • Clustering • Acknowledgements • Questions Slide 2

What is Molecular Modeling? Wikipedia´(http: //en. wikipedia. org/wiki/Molecular_modelling): Molecular modelling encompasses all theoretical methods and computational techniques used to model or mimic the behaviour of molecules. The techniques are used in the fields of computational chemistry, computational biology and materials science for studying molecular systems ranging from small chemical systems to large biological molecules and material assemblies…. inevitably computers are required to perform molecular modelling of any reasonably sized system…. Andrew R. Leach, ”Molecular modelling, principles and applications”, second edition: …we shall not concern ourselves with semantics but rather shall consider any theoretical or computational technique that provides insight into the behaviour of molecular systems to be an example of molecular modelling. Slide 3

The Molecular Modeling Toolbox Molecular Mechanics Methods Quantum Mechanical Methods Molecules modeled as spheres (atoms) connected by springs (bonds) Molecules represented using electron structure (Schrödinger equation) • Fast, >106 atoms • Computationally expensive , <10 -100 atoms, depending on method • Limited flexibility due to lack of electron treatment • Highly flexible – any property can in principle be calculated Typical applications § Simulating biomolecules in explicit solvent/membrane § Geometry optimization § Conformational search Slide 4 § Chemical reactions § Spectra § Accurate (gas phase) structures, energies

The insulin project at the NP 3 centre* • Synthesis: Engineered insulin with a novel metalion bindingsite • Experimental data: CD, UV-vis • Goal: Elucidate the structure of a the novel insulin-complex in solution • Molecular modeling methodologies employed: Slide 5 • Molecular mechanics • Molecular dynamics • Quantum mechanics (Density functional theory) *http: //www. np 3. life. ku. dk/
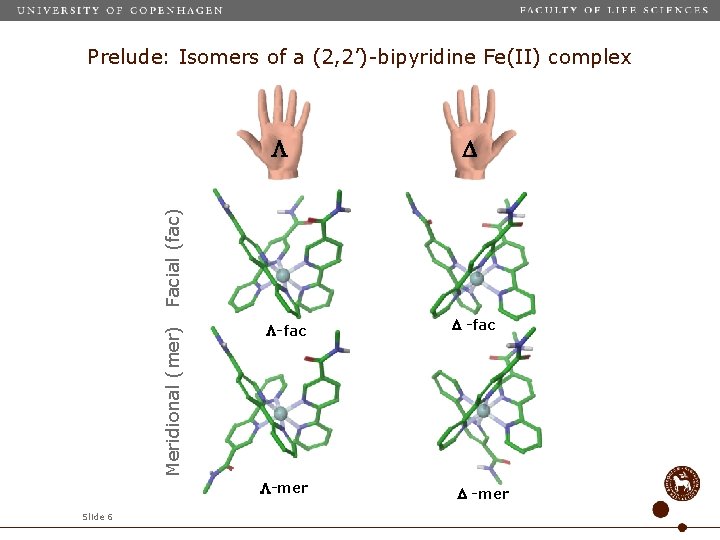
Prelude: Isomers of a (2, 2’)-bipyridine Fe(II) complex Meridional (mer) Facial (fac) -fac -mer Slide 6 -fac -mer
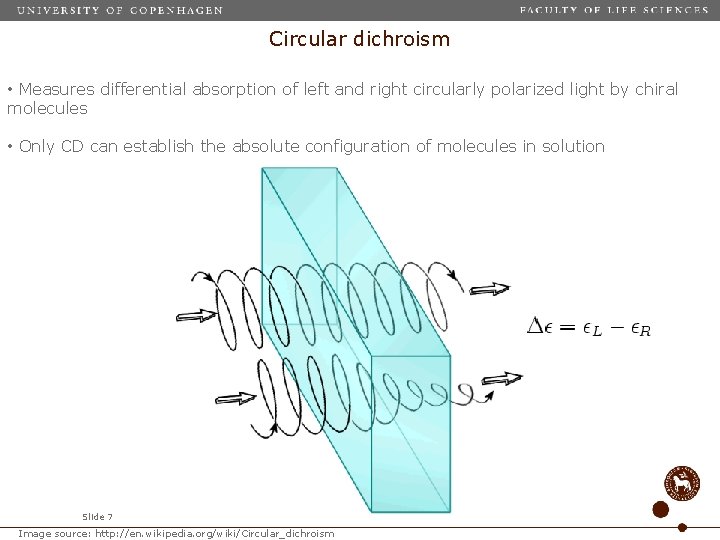
Circular dichroism • Measures differential absorption of left and right circularly polarized light by chiral molecules • Only CD can establish the absolute configuration of molecules in solution Slide 7 Image source: http: //en. wikipedia. org/wiki/Circular_dichroism
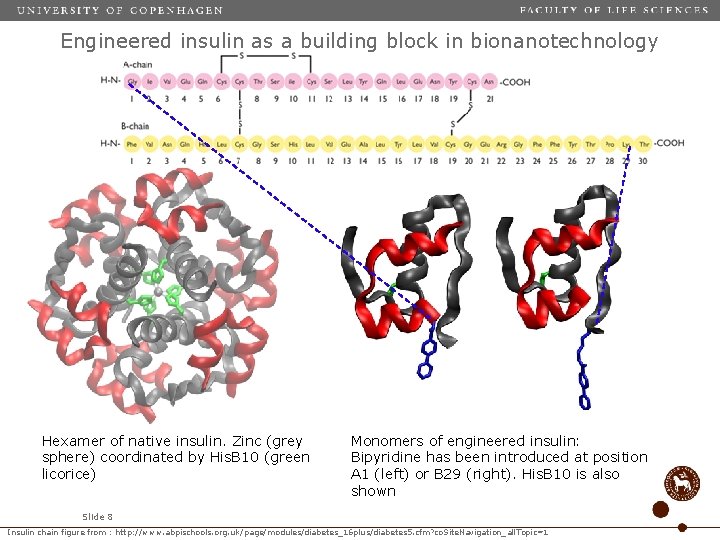
Engineered insulin as a building block in bionanotechnology Hexamer of native insulin. Zinc (grey sphere) coordinated by His. B 10 (green licorice) Monomers of engineered insulin: Bipyridine has been introduced at position A 1 (left) or B 29 (right). His. B 10 is also shown Slide 8 Insulin chain figure from : http: //www. abpischools. org. uk/page/modules/diabetes_16 plus/diabetes 5. cfm? co. Site. Navigation_all. Topic=1
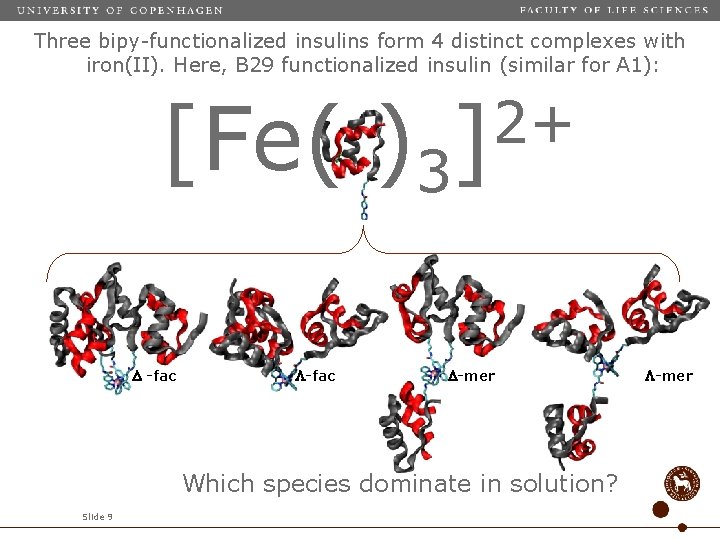
Three bipy-functionalized insulins form 4 distinct complexes with iron(II). Here, B 29 functionalized insulin (similar for A 1): [Fe( )3 -fac 2+ ] -mer Which species dominate in solution? Slide 9 -mer
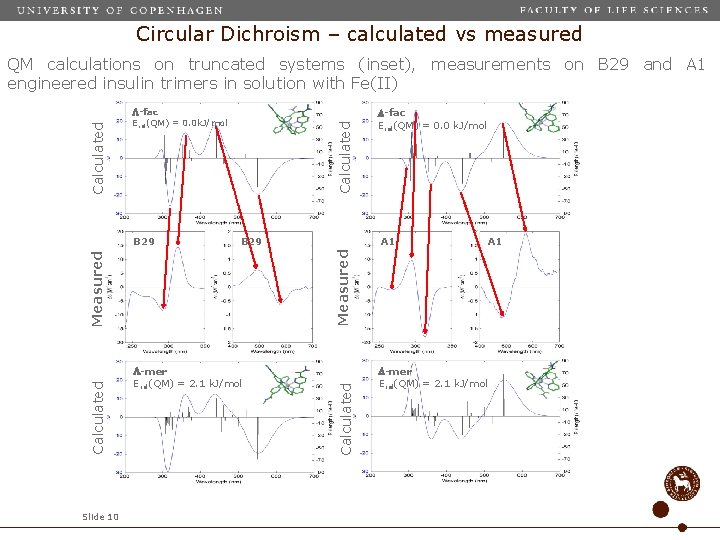
Circular Dichroism – calculated vs measured B 29 Measured Calculated Slide 10 -fac Erel(QM) = 0. 0 k. J/mol A 1 Measured B 29 Calculated -fac Erel(QM) = 0. 0 k. J/mol -mer Erel(QM) = 2. 1 k. J/mol Calculated QM calculations on truncated systems (inset), measurements on B 29 and A 1 engineered insulin trimers in solution with Fe(II) -mer Erel(QM) = 2. 1 k. J/mol A 1

Circular Dichroism – calculated vs measured • Comparison of measured/calculated CD sign changes allows determination of enantiomer dominating in solution: A 1 ( ), B 29 ( ) • Meridional (mer) and facial (fac) configuation cannot be firmly established from CD alone. • Energies from a conformational search on (truncated) systems may help in determining fac/mer preferences Slide 11
![Conformational search on a truncated B 29 trimer Conformational search: [Fe(bipy)3]2+ core fixed, rotate Conformational search on a truncated B 29 trimer Conformational search: [Fe(bipy)3]2+ core fixed, rotate](http://slidetodoc.com/presentation_image/1ccd00d48921059fc1c48f99b57a9a61/image-12.jpg)
Conformational search on a truncated B 29 trimer Conformational search: [Fe(bipy)3]2+ core fixed, rotate remaining groups systematically to find lowest energy: -fac 0. 0 k. J/mol Slide 12 -fac 14. 3 k. J/mol -mer 25. 4 k. J/mol -mer 30. 0 k. J/mol
Molecular dynamics simulations can be used to elucidate the dynamics of biomolecules • Example: Rearrangement of an engineered insulin monomer Slide 13

Clustering: Building a larger calculator Slide 14

Acknowledgements Henrik K. Munchb, Søren Thiis Heidea, Thomas Hoeg-Jensenc, Peter Waaben Thulstrupa and Knud J. Jensenb a Bioinorganic Chemistry, Department of Basic Sciences and Environment, Faculty of Life Sciences, University of Copenhagen, Denmark b Bioorganic Chemistry, Department of Basic Sciences and Environment, Faculty of Life Sciences, University of Copenhagen, Denmark c Novo Nordisk , Maaloev, Denmark Det Strategiske Forskningsråds Programkomite for Nanovidenskab og -teknologi, Bioteknologi og IT (NABIIT) Slide 15
- Slides: 15