Molecular modeling group Bioengineering department Biology Faculty M

Molecular modeling group Bioengineering department Biology Faculty M. V. Lomonosov Moscow State University Calculation of interaction energy between voltage-gated potassium channel Kv 1. 2 and blocker agitoxin Valery N. Novoseletsky Maria A. Bolshakova Konstantin V. Shaitan Moscow 2013
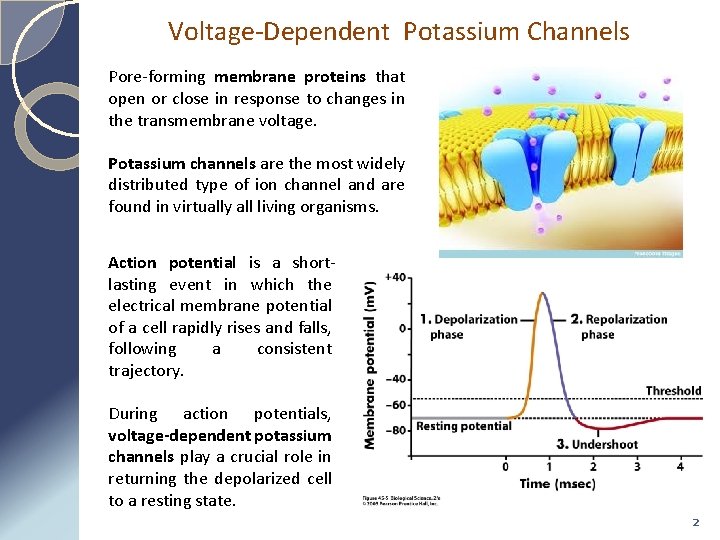
Voltage-Dependent Potassium Channels Pore-forming membrane proteins that open or close in response to changes in the transmembrane voltage. Potassium channels are the most widely distributed type of ion channel and are found in virtually all living organisms. Action potential is a shortlasting event in which the electrical membrane potential of a cell rapidly rises and falls, following a consistent trajectory. During action potentials, voltage-dependent potassium channels play a crucial role in returning the depolarized cell to a resting state. 2

Crystal Structure of a Mammalian Voltage-Dependent K+ Channel (Long, Campbell, Mac. Kinnon, 2005) 3
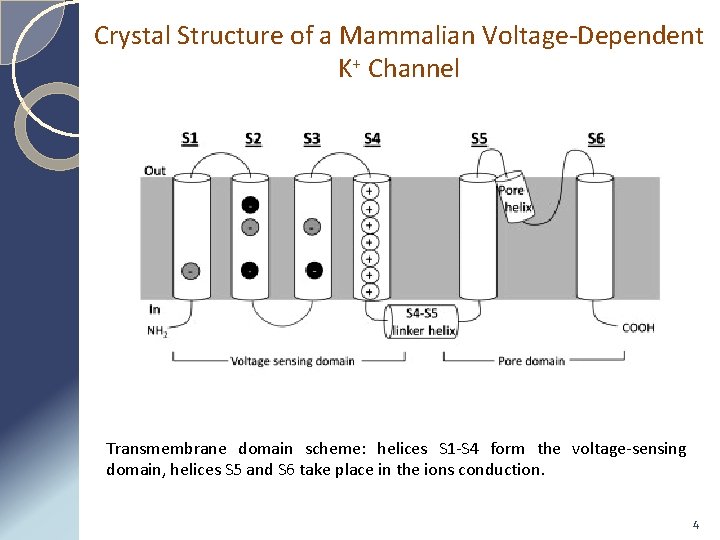
Crystal Structure of a Mammalian Voltage-Dependent K+ Channel Transmembrane domain scheme: helices S 1 -S 4 form the voltage-sensing domain, helices S 5 and S 6 take place in the ions conduction. 4

Crystal Structure of a Mammalian Voltage-Dependent K+ Channel Side view: voltage-sensing domain , pore domain and selective filter. 5

Scorpion venom toxin – agitoxin Agitoxin structure was determined by NMR (Kresel, Kasibhatha, 1995). Structure has highly conserved motif consisting of - 3 antiparallel β-sheets - α-helix - 3 disulfide bridges Agitoxin specifically blocks Kv 1. 2 channel with high affinity (Kd < 1 nmol/L) 6
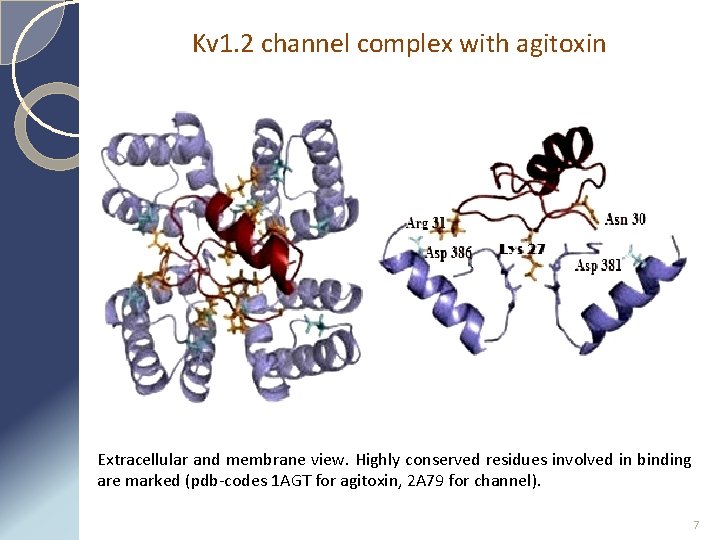
Kv 1. 2 channel complex with agitoxin Extracellular and membrane view. Highly conserved residues involved in binding are marked (pdb-codes 1 AGT for agitoxin, 2 A 79 for channel). 7
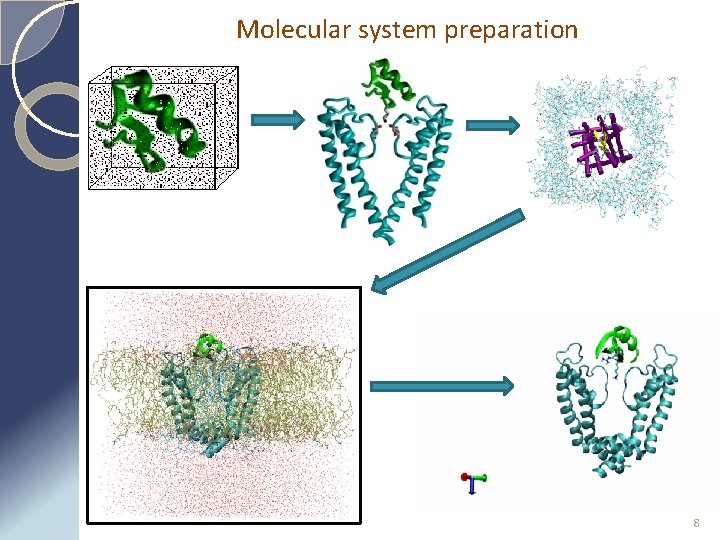
Molecular system preparation 8
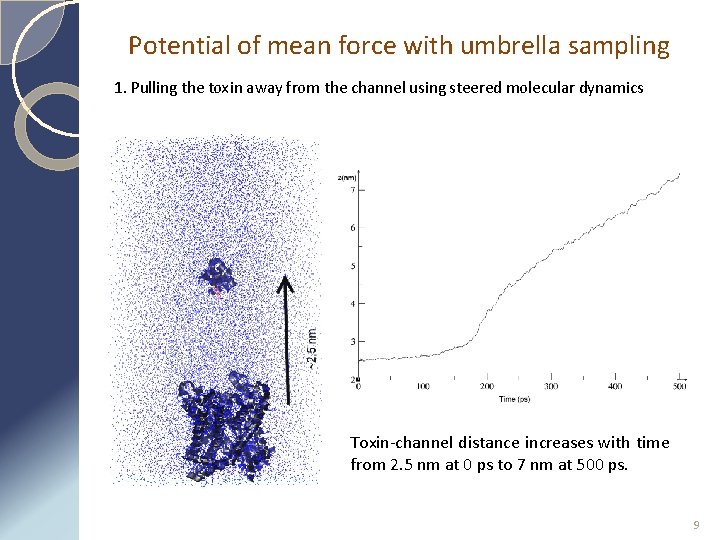
Potential of mean force with umbrella sampling 1. Pulling the toxin away from the channel using steered molecular dynamics Toxin-channel distance increases with time from 2. 5 nm at 0 ps to 7 nm at 500 ps. 9
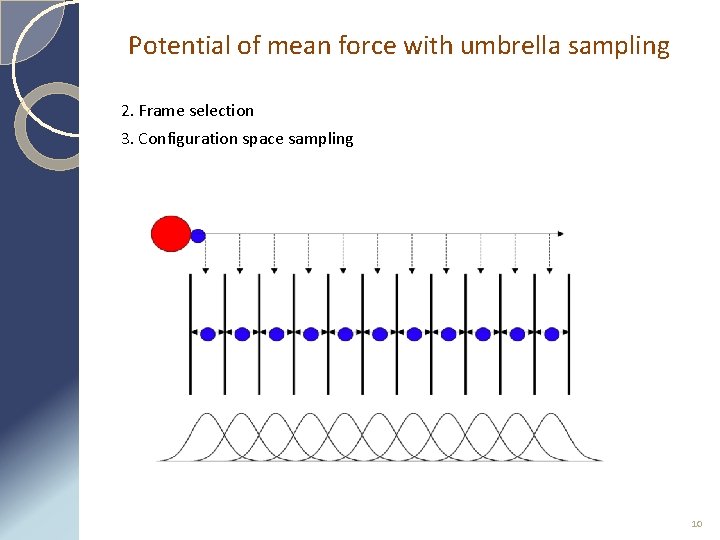
Potential of mean force with umbrella sampling 2. Frame selection 3. Configuration space sampling 10
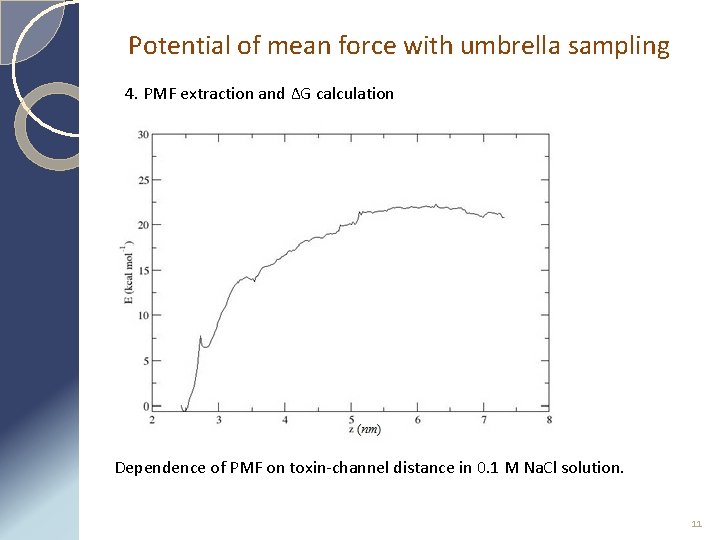
Potential of mean force with umbrella sampling 4. PMF extraction and ΔG calculation Dependence of PMF on toxin-channel distance in 0. 1 M Na. Cl solution. 11
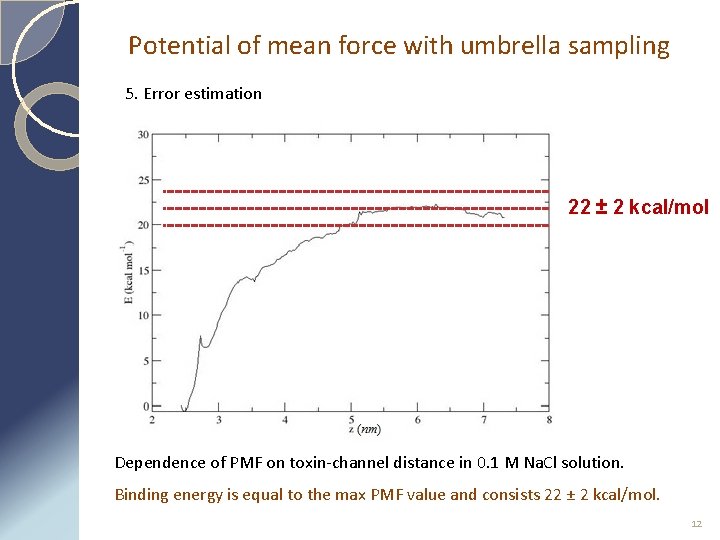
Potential of mean force with umbrella sampling 5. Error estimation 22 ± 2 kcal/mol Dependence of PMF on toxin-channel distance in 0. 1 M Na. Cl solution. Binding energy is equal to the max PMF value and consists 22 ± 2 kcal/mol. 12
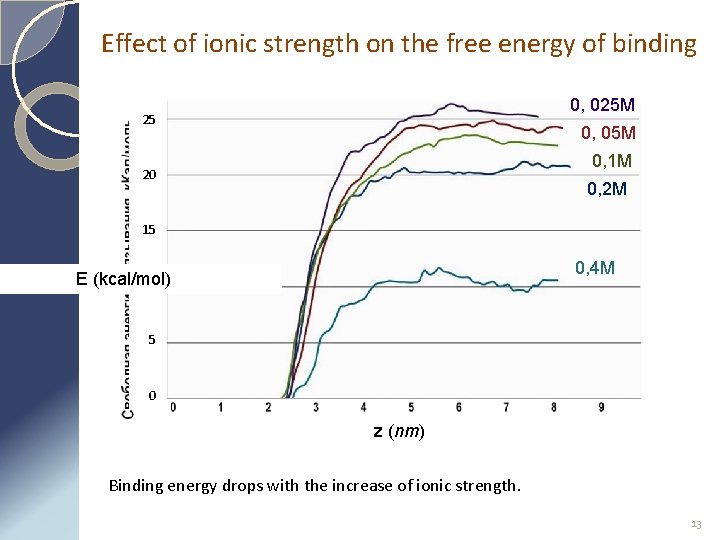
Effect of ionic strength on the free energy of binding 0, 025 M 25 0, 05 M 0, 1 M 20 0, 2 M 15 0, 4 M E (kcal/mol) 10 5 0 z (nm) Binding energy drops with the increase of ionic strength. 13
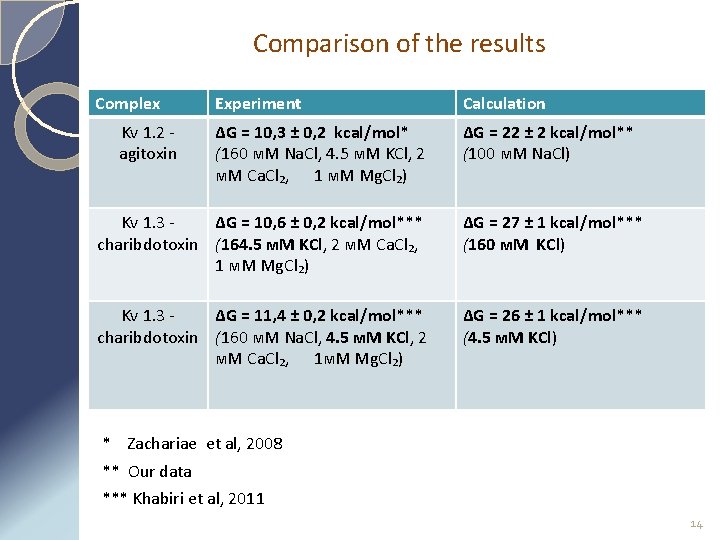
Comparison of the results Complex Kv 1. 2 agitoxin Experiment Calculation ∆G = 10, 3 ± 0, 2 kcal/mol* (160 м. М Na. Cl, 4. 5 м. М KCl, 2 м. М Ca. Cl₂, 1 м. М Mg. Cl₂) ∆G = 22 ± 2 kcal/mol** (100 м. М Na. Cl) Kv 1. 3 ∆G = 10, 6 ± 0, 2 kcal/mol*** charibdotoxin (164. 5 м. М KCl, 2 м. М Ca. Cl₂, 1 м. М Mg. Cl₂) ∆G = 27 ± 1 kcal/mol*** (160 м. М KCl) Kv 1. 3 ∆G = 11, 4 ± 0, 2 kcal/mol*** charibdotoxin (160 м. М Na. Cl, 4. 5 м. М KCl, 2 м. М Ca. Cl₂, 1 м. М Mg. Cl₂) ∆G = 26 ± 1 kcal/mol*** (4. 5 м. М KCl) * Zachariae et al, 2008 ** Our data *** Khabiri et al, 2011 14

Conclusion 1. Structural model of agitoxin-channel complex was constructed 2. Free energy binding was calculated using potential of mean force and umbrella sampling 3. Obtained results differ from experimental data. Probable reason is in the mistake of system preparation 15

Thank you for attention! 16
- Slides: 16