MOLECULAR GENETICS How is a gene different from

























- Slides: 41

MOLECULAR GENETICS How is a gene different from a chromosome? 2. How is a gene expressed? 3. What are some characteristics that must be exhibited by a gene 4. What is the chemical composition of a gene? Characteristics of DNA 1. Must be variable 2. Must store information 3. Must be able to replicate 4. Must control protein production 1.

THE DOUBLE HELIX n Historical Survey n 1869 Frederich Meischer – discovered Nucleic Acids “white, sugary, slightly acidic and phosphorous chemicals found in the cell’s nucleus” •

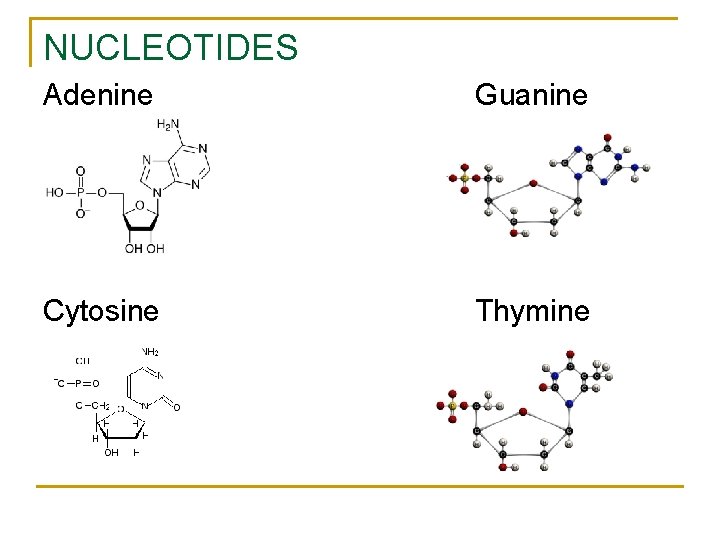
NUCLEOTIDE STRUCTURE Nucleotides – the building blocks of nucleic acids 1. Phosphate → H 3 PO 4 2. 5 – Carbon Sugar → C 5 H 10 O 5 (DNA = C 5 H 10 O 4) 3. Nitrogen base compound Adenine purines → double ring structures Guanine Cytosine pyrimidines → single ring structures Thymine Tetranucleotide Theory – grouped in clusters of 4

NUCLEOTIDES Adenine Guanine Cytosine Thymine

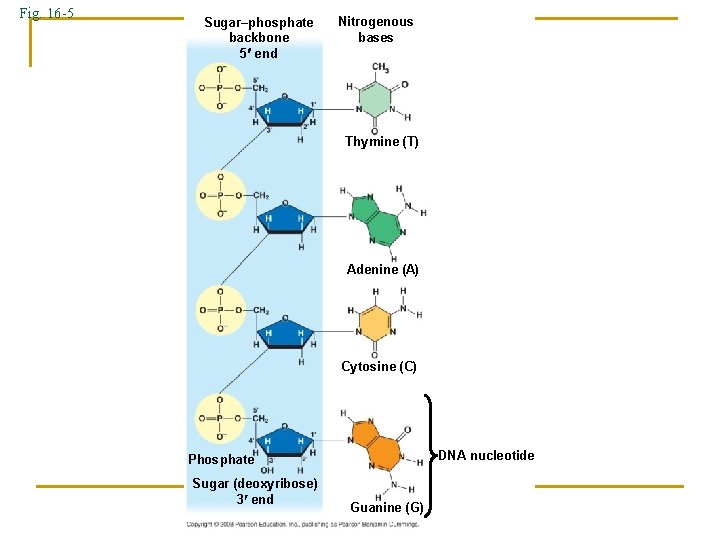
Fig. 16 -5 Sugar–phosphate backbone 5 end Nitrogenous bases Thymine (T) Adenine (A) Cytosine (C) DNA nucleotide Phosphate Sugar (deoxyribose) 3 end Guanine (G)

ANSWER THE FOLLOWING: 1. What is the name of the bacterium that Frederick Griffith studied? 2. What happened in this experiment that caused the mice to develop pneumonia? 3. What is the principle of transformation? 4. What did Frederick Griffith have to do with DNA?

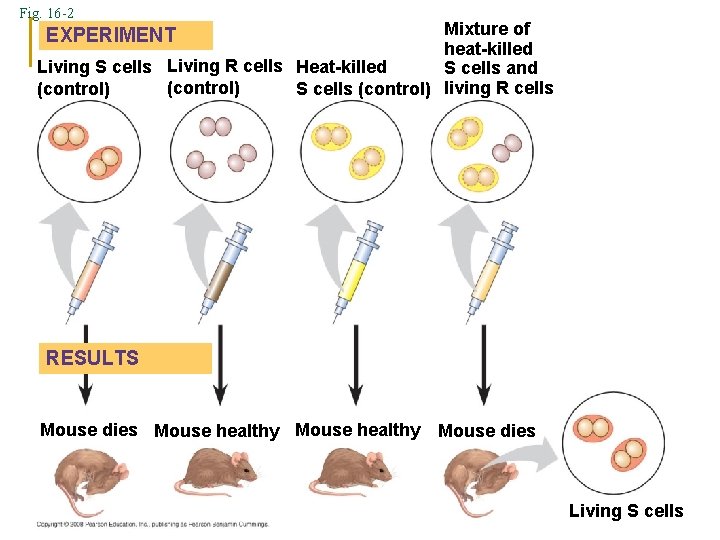
BACTERIAL TRANSFORMATION 1928 Frederick Griffith 2 Types of Bacteria S = Smooth + Virulent (encapsulated) R = Rough + Non-Virulent (without capsule) 1. S injected → mice die 2. R injected → mice live 3. Heat killed S injected → mice live 4. Dead S + R → ? mice die Why?

Fig. 16 -2 Mixture of heat-killed Living S cells Living R cells Heat-killed S cells and (control) S cells (control) living R cells EXPERIMENT RESULTS Mouse dies Mouse healthy Mouse dies Living S cells

Transformation – a genetic change caused by the incorporation into a cell of a “transforming factor” from the external medium R bacteria received genetic material from S R took on the characteristics of S 1943 Avery, Macloud and Mc. Carty → transforming factor = DNA

Answer the following: 1. 2. 3. 4. 5. 6. What is a bacteriophage? What is the structure of the bacteriophage? What does the virus do after is attaches to a bacterium? Describe the activities of the viral DNA after it enters the cell. How does the answer to question 4 relate to the functions of DNA? Complete the template.

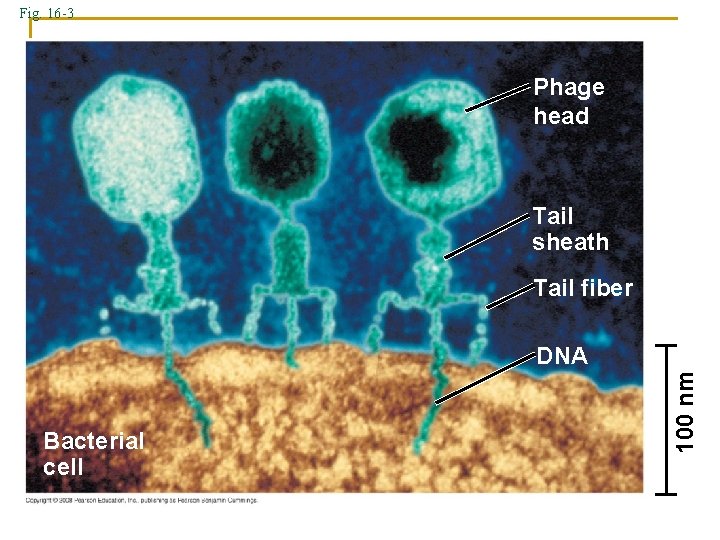
Fig. 16 -3 Phage head Tail sheath Tail fiber Bacterial cell 100 nm DNA

Bacteriophage Experiments 1940 Delbruck and Luria → viral studies Results 1. Viral Structure 2. Viruses inject DNA into host 3. Viral DNA replicates 4. Viral DNA directs production of protein coats DNA controls reproduction, heredity, and all life processes by directing protein (enzyme) production How? DNA structure must be determined

Fig. 16 -1

DNA Structure – Watson and Crick n 1. 2. 3. 4. 5. 6. Known Data DNA → large, long, thin, made of nucleotides Assembled in repeating units of 4? ? Chargaff’s Rule (% A = % T, % G = % C) X-Ray diffraction measurements by Maurice Wilkins and Rosaline Franklin Must show variety. Must indicate a mechanism for replication.

Fig. 16 -6 (a) Rosalind Franklin (b) Franklin’s X-ray diffraction photograph of DNA

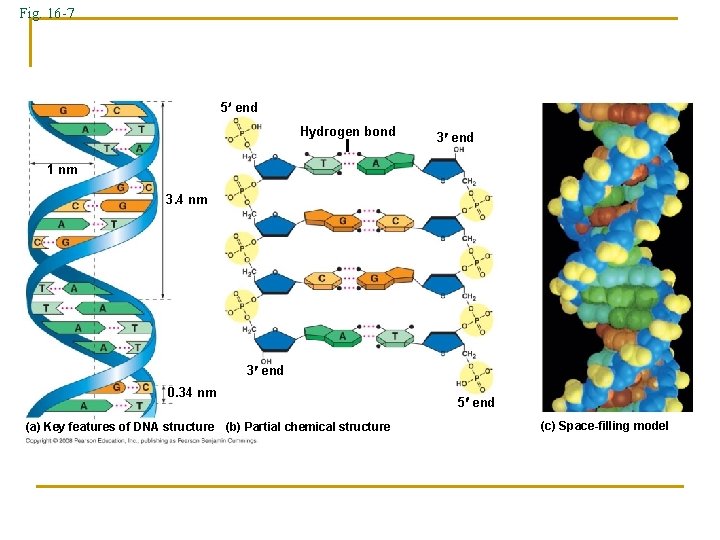
Fig. 16 -7 5 end Hydrogen bond 3 end 1 nm 3. 4 nm 3 end 0. 34 nm (a) Key features of DNA structure (b) Partial chemical structure 5 end (c) Space-filling model

DNA LAB 1. 2. 3. 4. Construct a chain (strand) of nucleotides with the following sequence: What is the complementary sequence for your DNA molecule? Attach a complementary nucleotide to each nucleotide in your strand a) be sure to attach nucleotides in order b) be sure to attach nucleotides 5’ to 3’ Draw a diagram of your DNA model.

ANSWER THE FOLLOWING: If DNA is compared to a ladder…. 1. What are the “rungs” of the ladder? 2. How are they held together? 3. What are the “uprights” of the ladder? 4. How are they held together? 5. How is your model different from others? 6. Propose a possible explanation for DNA replication.

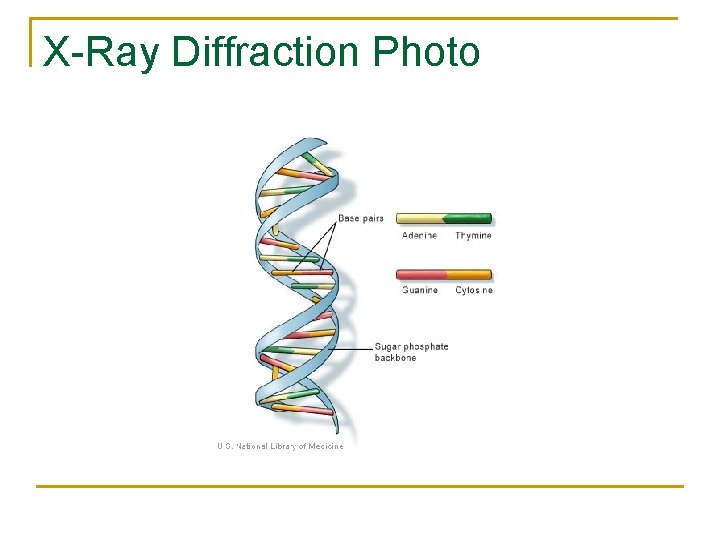
X-Ray Diffraction Photo

1953 Watson and Crick’s Model DNA = Double Helix (twisted ladder) 2. N-base pairs = rungs 3. must be a purine and a pyrimidine Why? 2 purines > 2 nm 2 pyrimidines < 2 nm A – T bond with 2 H bonds G – C bond with 3 H bonds Paired bases are COMPLEMENTARY 1.

DNA Structure 4. Each strand has direction 5' end → phosphate to sugar 3' end → sugar (open end) Strands are ANTIPARALLEL 5 ' TTCAG 3 ' 3‘ 5. Uprights = alternating phosphates and sugars held by strong covalent bonds 6. Nucleotide sequence is always written 5 '→ 3 '

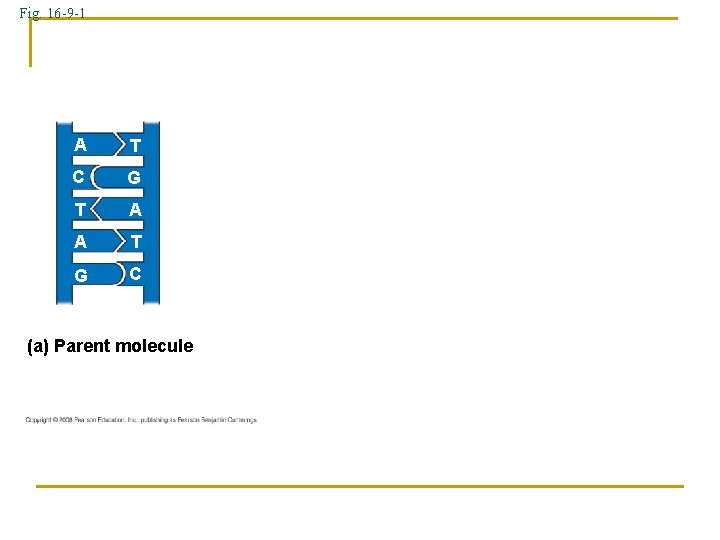
Fig. 16 -9 -1 A T C G T A A T G C (a) Parent molecule

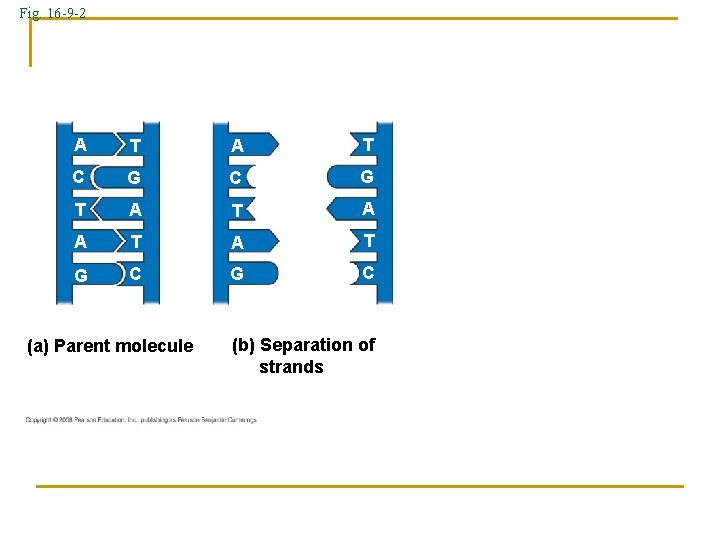
Fig. 16 -9 -2 A T C G T A A T G C (a) Parent molecule (b) Separation of strands

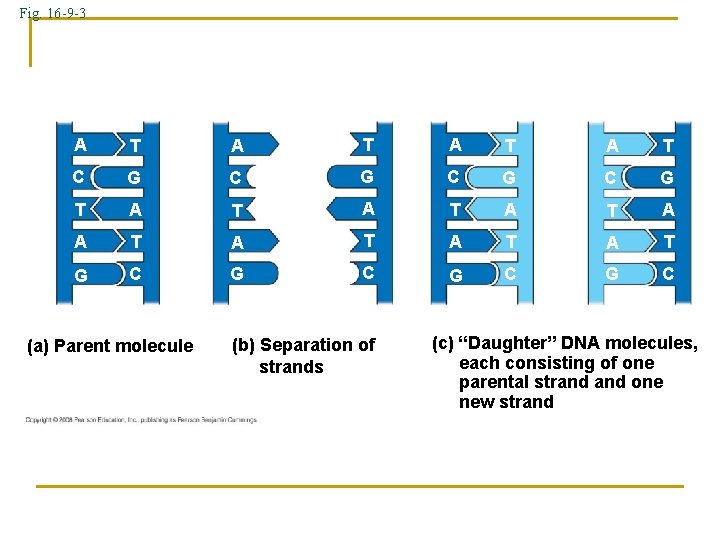
Fig. 16 -9 -3 A T A T C G C G T A T A T G C G C (a) Parent molecule (b) Separation of strands (c) “Daughter” DNA molecules, each consisting of one parental strand one new strand

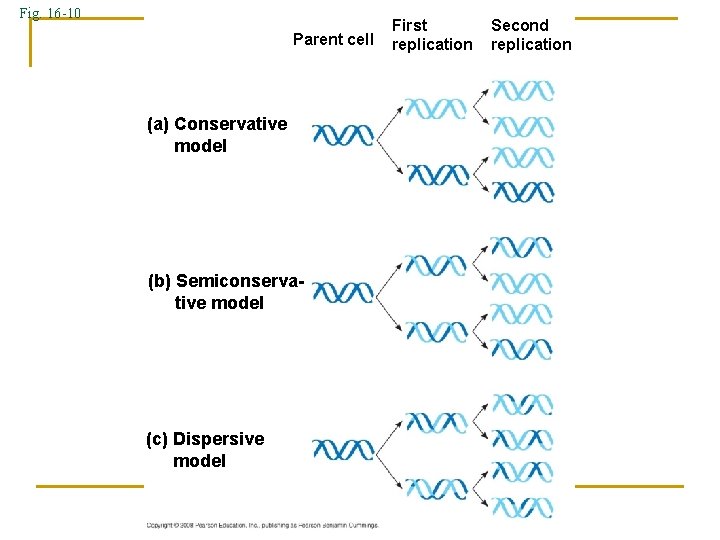
DNA REPLICATION Meselson and Stahl 1. Conservative Replication 2. *Semi-Conservative Replication 3. Dispersive Replication

Fig. 16 -10 Parent cell (a) Conservative model (b) Semiconservative model (c) Dispersive model First replication Second replication

DNA REPLICATION n n n 1. 2. 3. Occurs during S phase Leads to mitosis or meiosis 50 nucleotides/sec (500/sec in prok) DNA uncoils ands unzips Each strand picks up complementary nucleotides in a 5’ to 3’ direction Fragments are joined

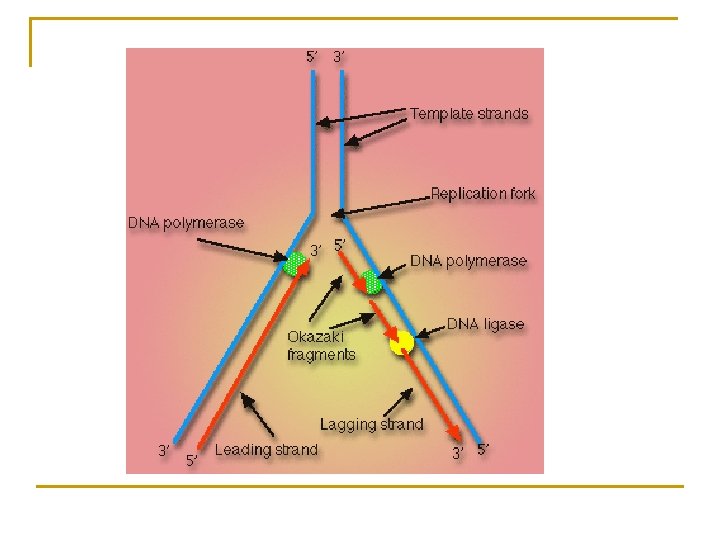
TERMINOLOGY FOR REPLICATION 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. Origin of Replication bubble Replication Fork Topoisomerase Helicase DNA Polymerase RNA Primer Leading Strand Lagging Strand Ligase

Origin of Replication n n n n Position on DNA where “unzipping” begins 1. helicase → to break H bonds 2. topoisomerase → to prevent supercoiling 3. single strand binding protein → to prevent kinking Under an electron microscope replication bubble replication fork Replication is bi-directional


REPLICATION TERMINOLOGY Leading Strand – produced in the direction of unzipping Lagging Strand – produced in the opposite direction Okasaki Fragments – fragments on lagging strand RNA primer – initial nucleotides attached to lagging strand RNA Primase – enzyme which regulates RNA primer DNA Polymerase III– regulates synthesis of new DNA strand

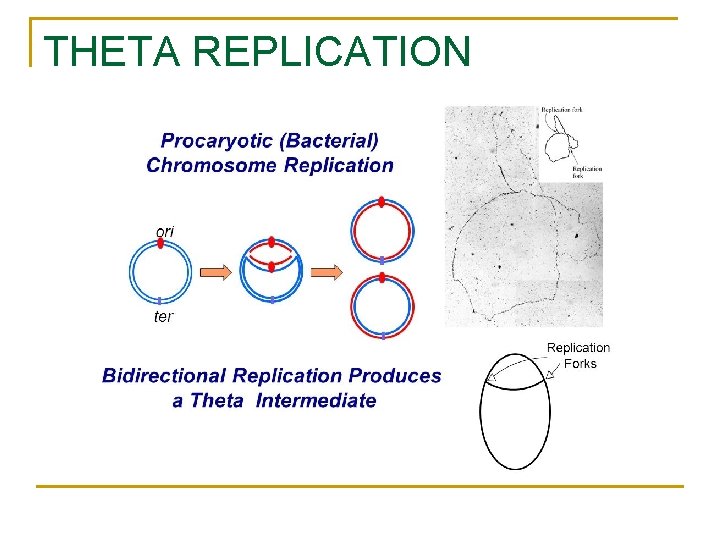
Eukaryotic vs Prokaryotic DNA Eukaryotes Shape Structure Origins of Replication Method Junk DNA Telomeres Linear DNA + Protein (Histones) Many Regular Much Present Prokaryotes Circular DNA One Theta Little or None Absent

THETA REPLICATION

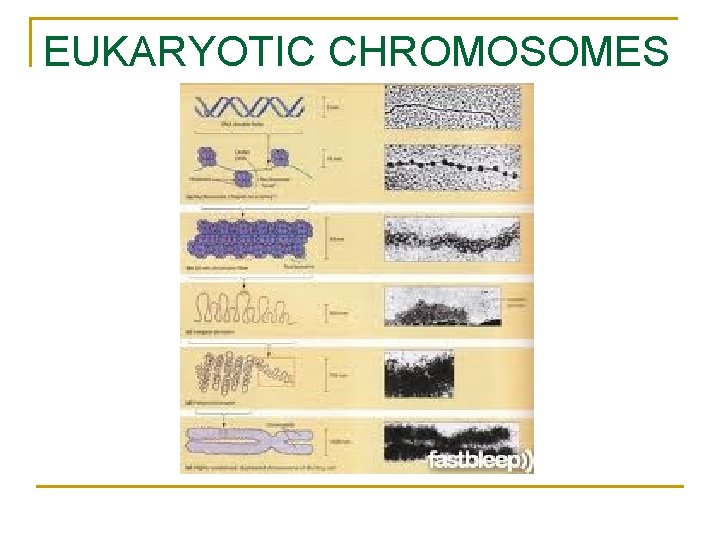
EUKARYOTIC CHROMOSOMES

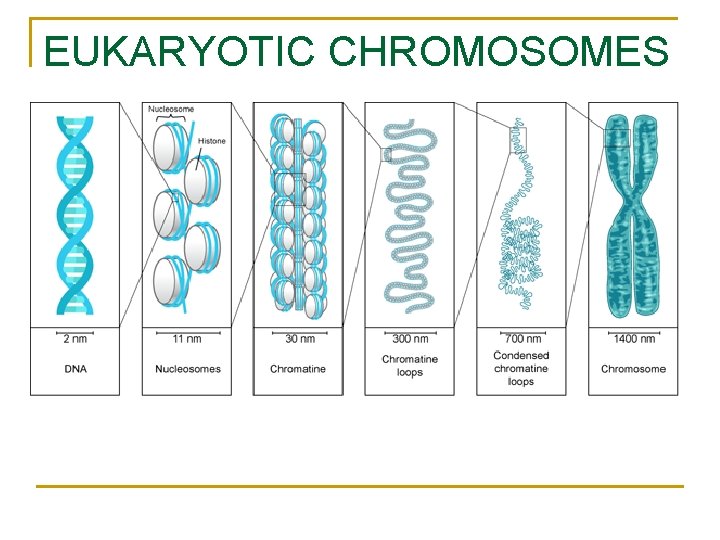
EUKARYOTIC CHROMOSOMES

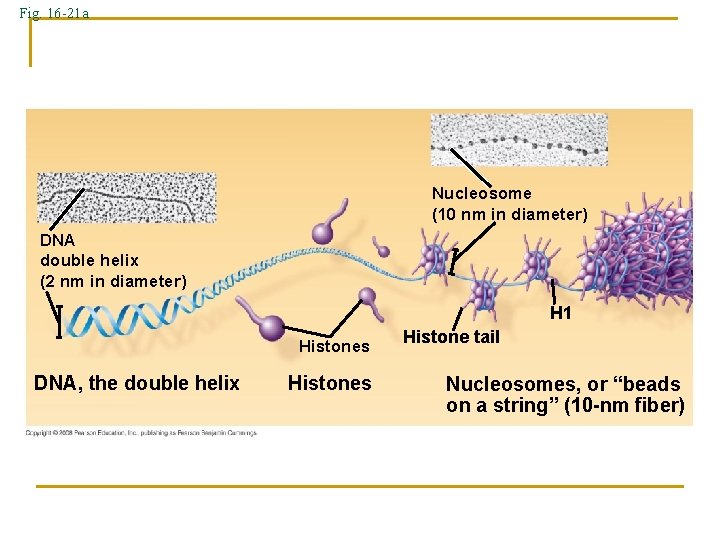
n Organization of Eukaryotic Chromosomes n 10 -nm fiber q q n DNA winds around histones to form nucleosome “beads” Nucleosomes are strung together like beads on a string by linker DNA 30 -nm fiber q Interactions between nucleosomes cause thin fiber to coil or fold into this thicker fiber Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

300 -nm fiber q The 30 -nm fiber forms looped domains that attach to proteins Metaphase chromosome q q q The looped domains coil further The width of a chromatid is 700 nm The width of a replicated chromosome is 1400 nm Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

CLASSES OF EUKARYTIC DNA 1. Simple-sequence DNA – repetitive sequences - makes up most DNA ex. ATAT, (Telomeres) 2. Intermediate-repeat DNA – code for histones 3. Single-copy DNA – code for proteins Discovered by Hybridization techniques

TELOMERES 1. 2. 3. 4. 5. Tips of chromosomes Made of Simple Sequence DNA (TTAGGG) Maintains chromosome structure Degenerates with each mitotic division May be related to the aging process No wonder Dolly died of old age at a young age.

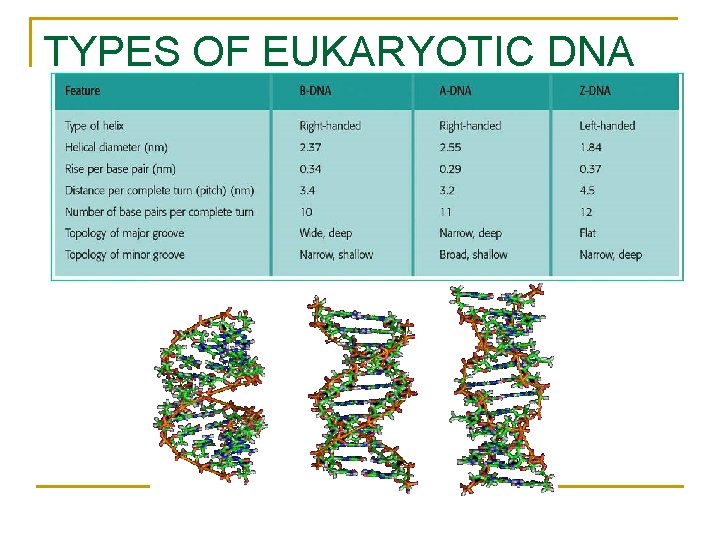
TYPES OF EUKARYOTIC DNA

Fig. 16 -21 a Nucleosome (10 nm in diameter) DNA double helix (2 nm in diameter) H 1 Histones DNA, the double helix Histones Histone tail Nucleosomes, or “beads on a string” (10 -nm fiber)