Molecular Flow of Genetic Information The processes by
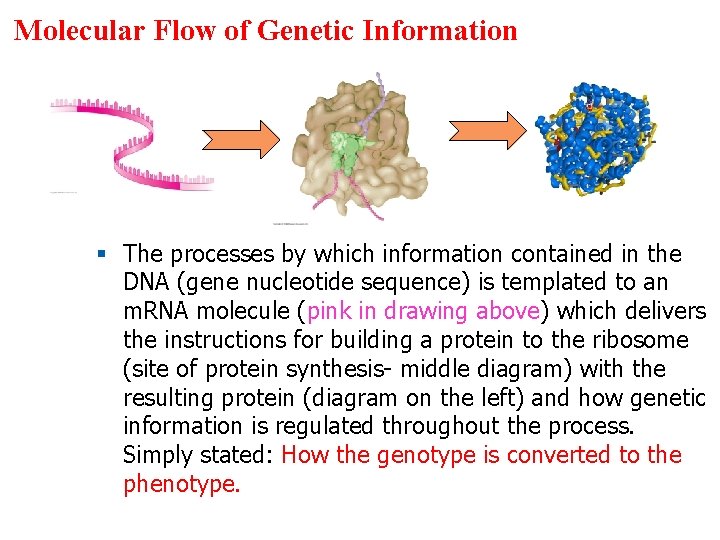
Molecular Flow of Genetic Information § The processes by which information contained in the DNA (gene nucleotide sequence) is templated to an m. RNA molecule (pink in drawing above) which delivers the instructions for building a protein to the ribosome (site of protein synthesis- middle diagram) with the resulting protein (diagram on the left) and how genetic information is regulated throughout the process. Simply stated: How the genotype is converted to the phenotype.
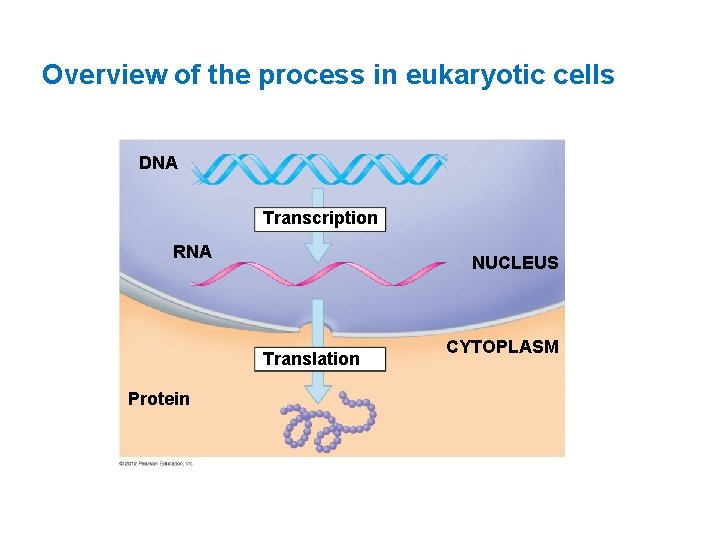
Overview of the process in eukaryotic cells DNA Transcription RNA NUCLEUS Translation Protein CYTOPLASM
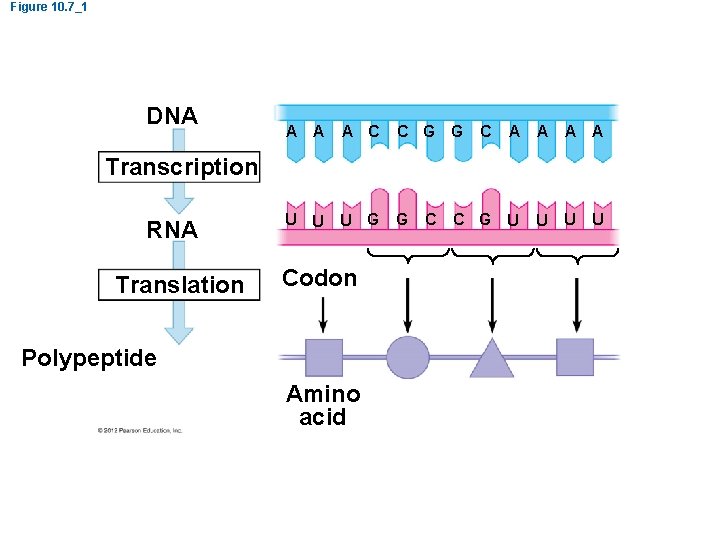
Figure 10. 7_1 DNA A C C G G U U U G G C A A C G U U U Transcription RNA Translation Codon Polypeptide Amino acid C U
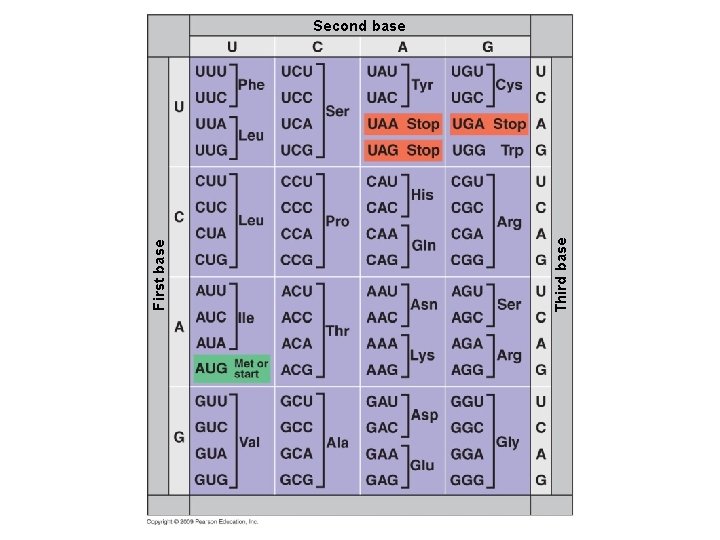
Third base First base Second base

Characteristics of the genetic code § Genetic code – Triplet: Three nucleotides specify one amino acid – 61 codons correspond to amino acids – AUG codes for methionine and signals the start of transcription – 3 “stop” codons signal the end of translation – Redundant: More than one codon for some amino acids – Unambiguous: Any codon for one amino acid does not code for any other amino acid – Does not contain spacers or punctuation: Codons are adjacent to each other with no gaps in between – Nearly universal Copyright © 2009 Pearson Education, Inc.

Transfer RNA molecules serve as interpreters during translation § Transfer RNA (t. RNA) molecules match an amino acid to its corresponding m. RNA codon – t. RNA structure allows it to convert one language to the other – An amino acid attachment site allows each t. RNA to carry a specific amino acid – An anticodon allows the t. RNA to bind to a specific m. RNA codon, complementary in sequence – A pairs with U, G pairs with C Copyright © 2009 Pearson Education, Inc.
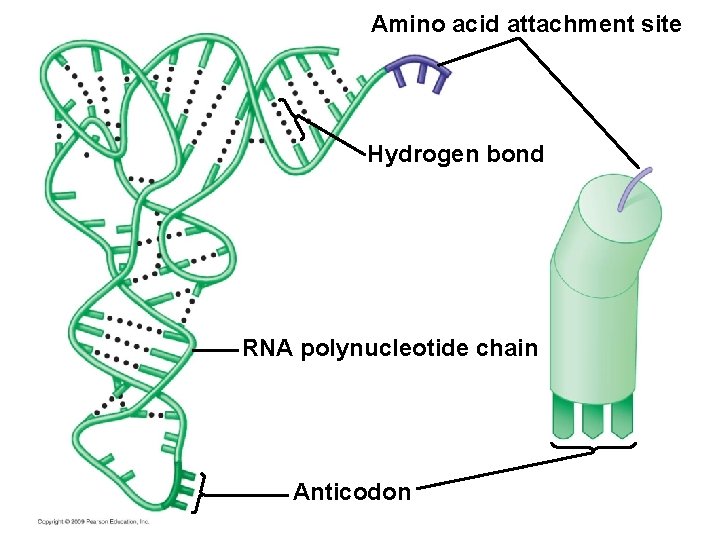
Amino acid attachment site Hydrogen bond RNA polynucleotide chain Anticodon

10. 12 Ribosomes build polypeptides § Translation occurs on the surface of the ribosome – Ribosomes have two subunits: small and large – Each subunit is composed of ribosomal RNAs and proteins – Ribosomal subunits come together during translation – Ribosomes have binding sites for m. RNA and t. RNAs Copyright © 2009 Pearson Education, Inc.
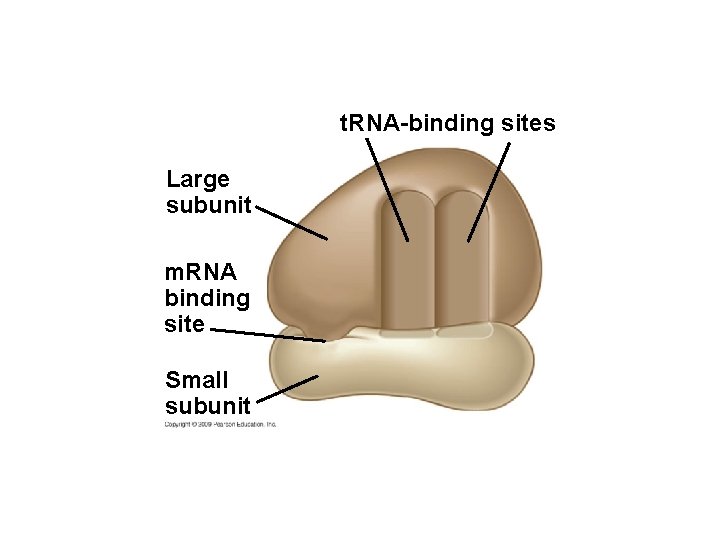
t. RNA-binding sites Large subunit m. RNA binding site Small subunit
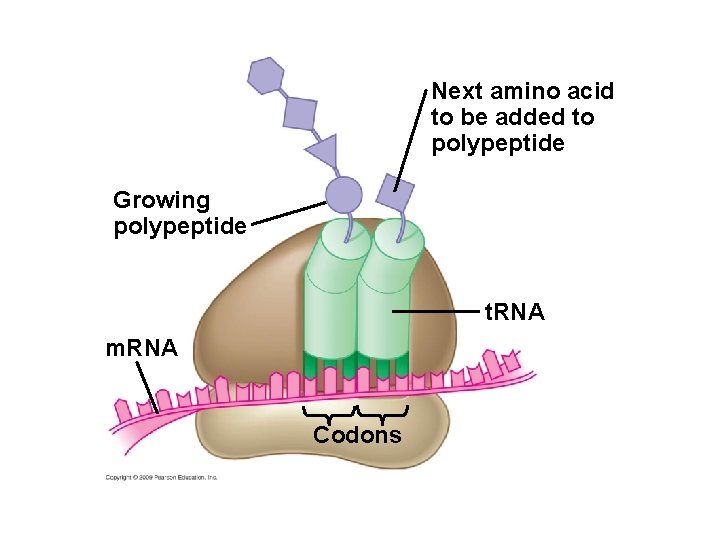
Next amino acid to be added to polypeptide Growing polypeptide t. RNA m. RNA Codons
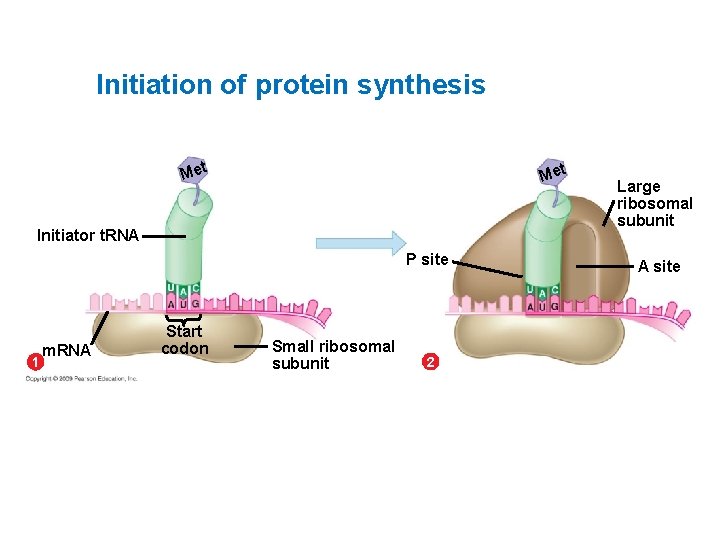
Initiation of protein synthesis Met Initiator t. RNA P site 1 m. RNA Start codon Small ribosomal subunit 2 Large ribosomal subunit A site

Elongation adds amino acids to the polypeptide chain until a stop codon terminates translation § Elongation is the addition of amino acids to the polypeptide chain § Each cycle of elongation has three steps 1. Codon recognition: next t. RNA binds to the m. RNA at the A site 2. Peptide bond formation: joining of the new amino acid to the chain – Amino acids on the t. RNA at the P site are attached by a covalent bond to the amino acid on the t. RNA at the A site 3. Translocation: t. RNA is released from the P site and the ribosome moves t. RNA from the A site into the P site Copyright © 2009 Pearson Education, Inc.
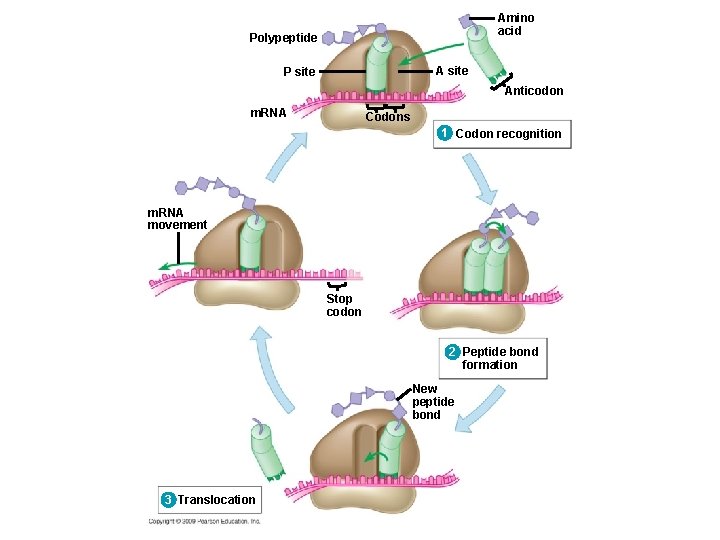
Amino acid Polypeptide A site P site Anticodon m. RNA Codons 1 Codon recognition m. RNA movement Stop codon 2 Peptide bond formation New peptide bond 3 Translocation

Peptide Bond Structure
Elongation adds amino acids to the polypeptide chain until a stop codon terminates translation § Elongation continues until the ribosome reaches a stop codon § Termination – The ribosomal subunits separate – m. RNA is released and can be translated again – The completed polypeptide is released § An m. RNA is usually translated simultaneously by multiple ribosomes. Animation: Translation Copyright © 2009 Pearson Education, Inc.
Animation: Translation Local copy
DNAi - Protein Synthesis Animation Go to the DNAi website select “Code. . Reading the Code…Putting It Together…Translation”. Watch Andrew Berry’s wonderful depiction of protein synthesis in real time. local copy

Protein Folding Gone Wrong: Prions Video: Mad Cow Disease
Video: Mad Cow Disease video backup link
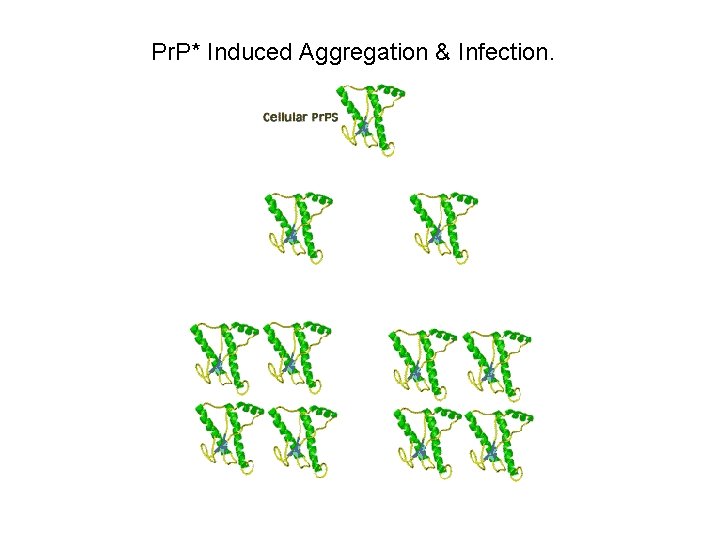
Pr. P* Induced Aggregation & Infection.
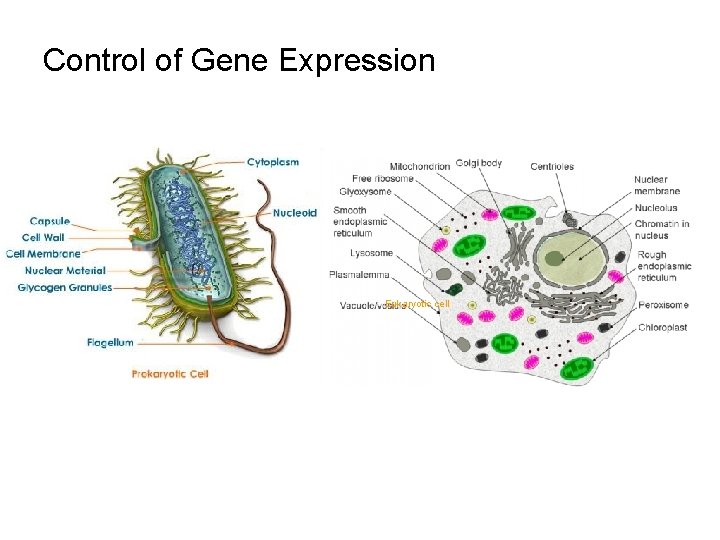
Control of Gene Expression Eukaryotic cell
Gene Expression: The process by which information flows from DNA to RNA to protein or genotype to phenotype. E. coli as model organism to understand prokaryotic gene expression. Features of prokaryotic transcriptional regulation • Ability to change metabolic activities in response to the environment. • Capability to induce/repress entire enzyme pathways- genome organization. • Examples of regulated gene networks: lac operon, trp operon
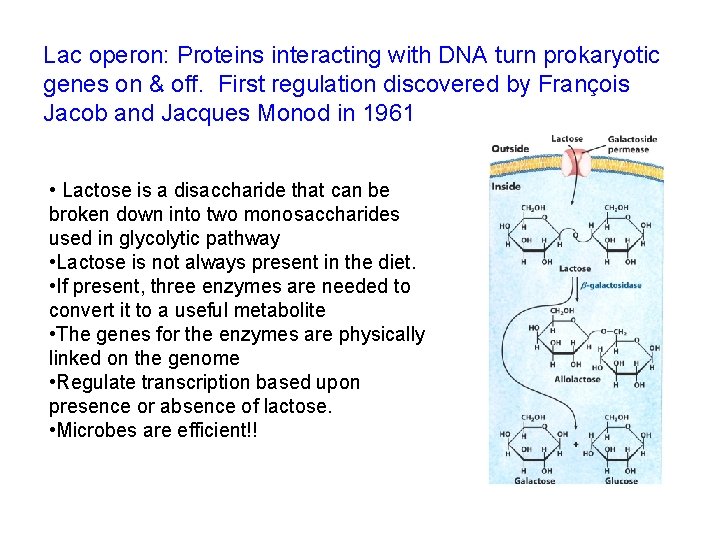
Lac operon: Proteins interacting with DNA turn prokaryotic genes on & off. First regulation discovered by François Jacob and Jacques Monod in 1961 • Lactose is a disaccharide that can be broken down into two monosaccharides used in glycolytic pathway • Lactose is not always present in the diet. • If present, three enzymes are needed to convert it to a useful metabolite • The genes for the enzymes are physically linked on the genome • Regulate transcription based upon presence or absence of lactose. • Microbes are efficient!!
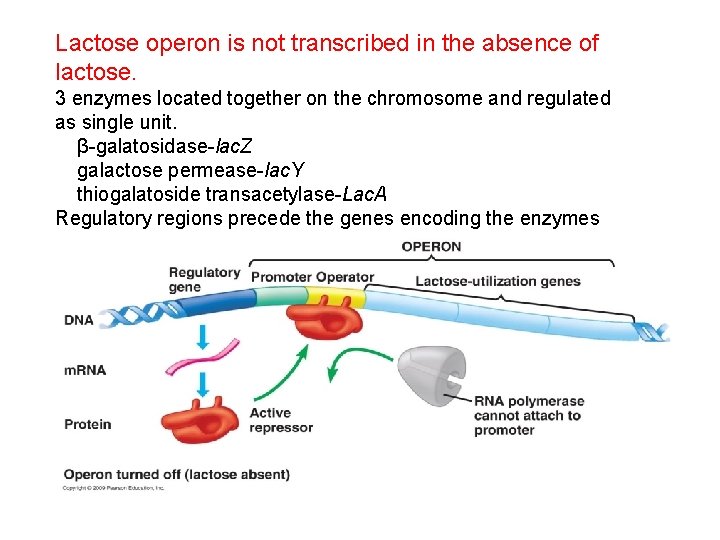
Lactose operon is not transcribed in the absence of lactose. 3 enzymes located together on the chromosome and regulated as single unit. β-galatosidase-lac. Z galactose permease-lac. Y thiogalatoside transacetylase-Lac. A Regulatory regions precede the genes encoding the enzymes
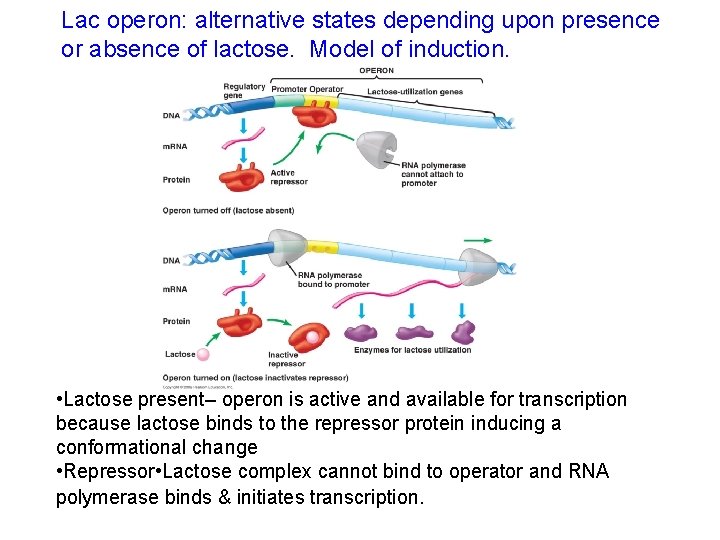
Lac operon: alternative states depending upon presence or absence of lactose. Model of induction. • Lactose present– operon is active and available for transcription because lactose binds to the repressor protein inducing a conformational change • Repressor • Lactose complex cannot bind to operator and RNA polymerase binds & initiates transcription.
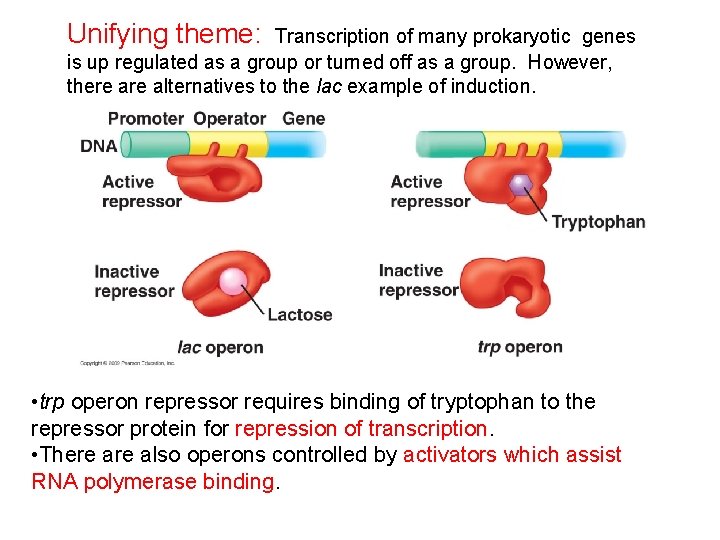
Unifying theme: Transcription of many prokaryotic genes is up regulated as a group or turned off as a group. However, there alternatives to the lac example of induction. • trp operon repressor requires binding of tryptophan to the repressor protein for repression of transcription. • There also operons controlled by activators which assist RNA polymerase binding.

Eukaryotic mechanisms of gene regulation 1. All cells contain the same genotype. Differential gene expression leads to different cell types. 2. Multiple mechanisms to regulate gene expression. • Eukaryotic transcription has multiple enhancers, silencers, and variety of proteins that are assembled into a complex. • DNA packing. Prevents access of the DNA for transcription. • Alternative RNA splicing leads to multiple m. RNAs for translation. • micro. RNAs and RNA interference. • Translation and later events also regulated.
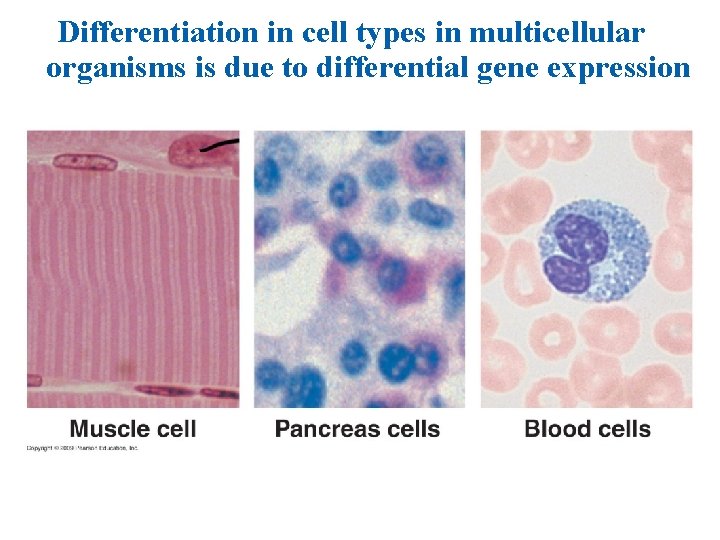
Differentiation in cell types in multicellular organisms is due to differential gene expression
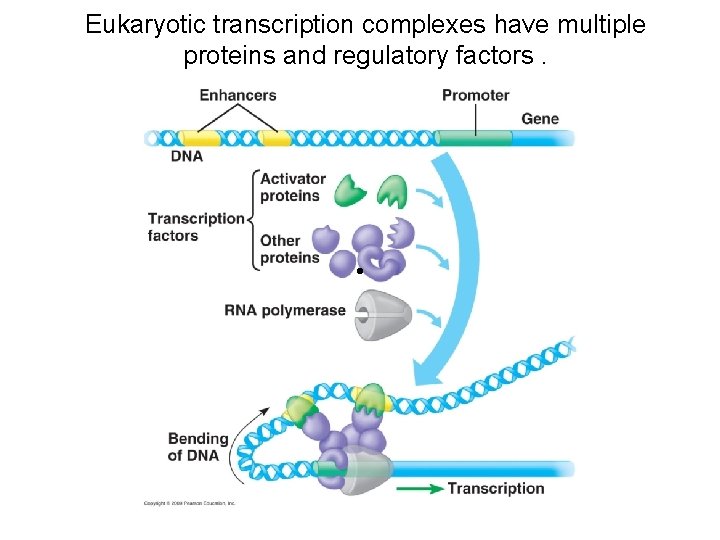
Eukaryotic transcription complexes have multiple proteins and regulatory factors. •
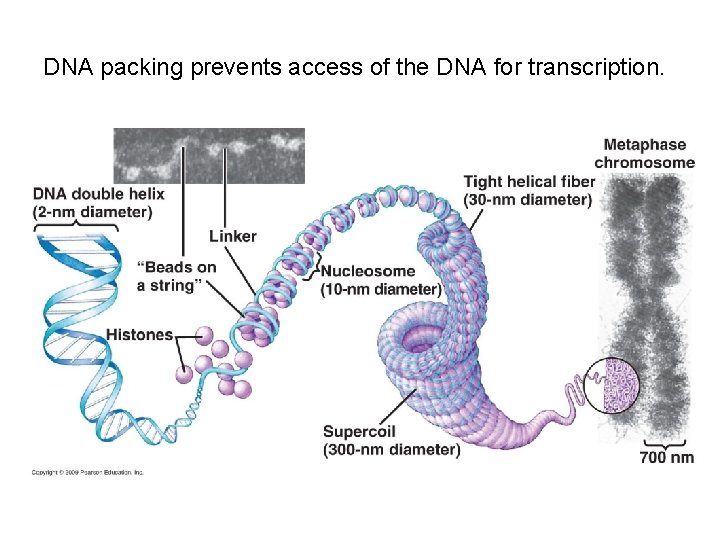
DNA packing prevents access of the DNA for transcription.
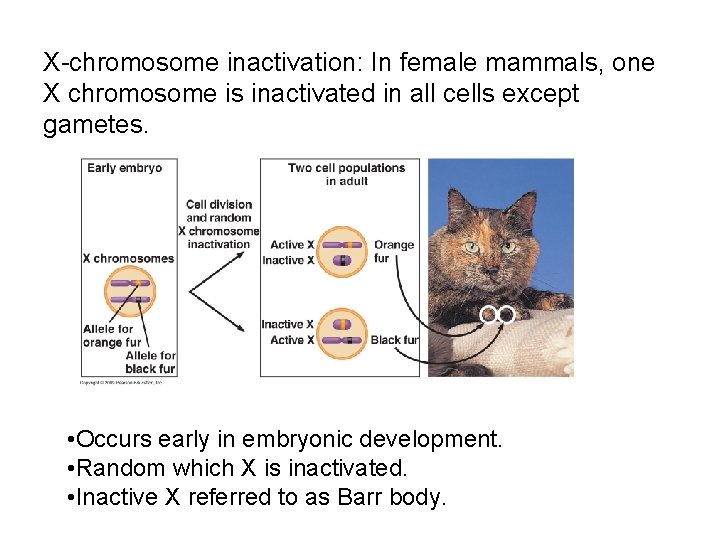
X-chromosome inactivation: In female mammals, one X chromosome is inactivated in all cells except gametes. • Occurs early in embryonic development. • Random which X is inactivated. • Inactive X referred to as Barr body.
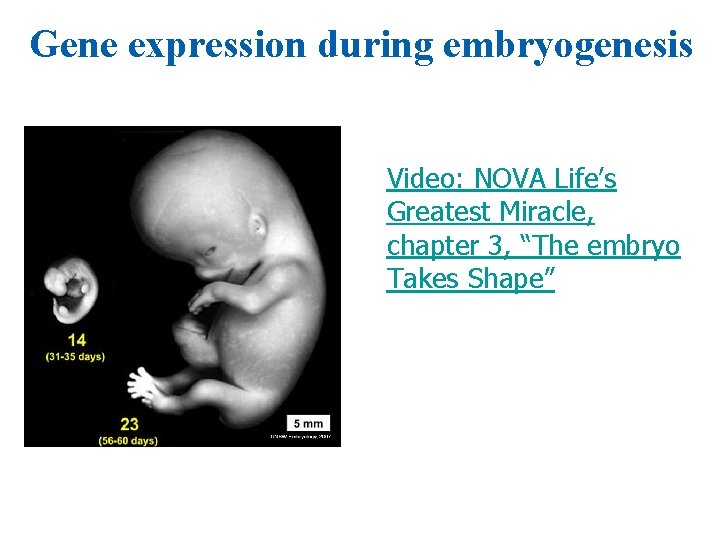
Gene expression during embryogenesis Video: NOVA Life’s Greatest Miracle, chapter 3, “The embryo Takes Shape”
Video: The embryo Takes Shape Original PBS site link (move the slider to 26: 48 time point) backup link
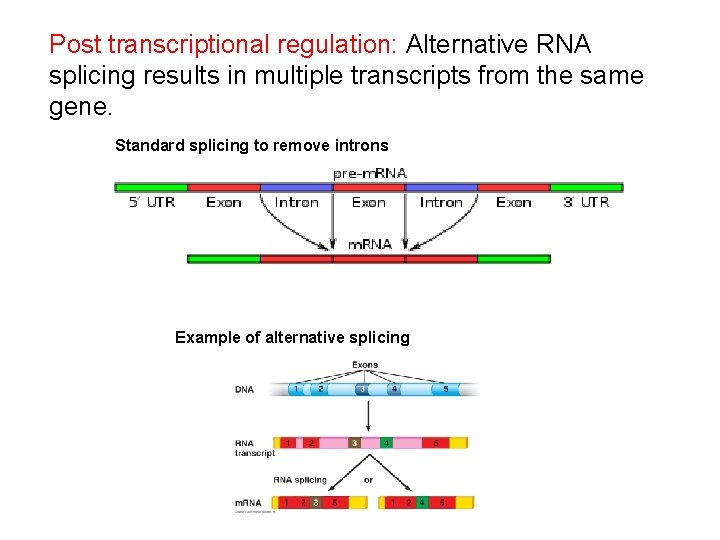
Post transcriptional regulation: Alternative RNA splicing results in multiple transcripts from the same gene. Standard splicing to remove introns Example of alternative splicing
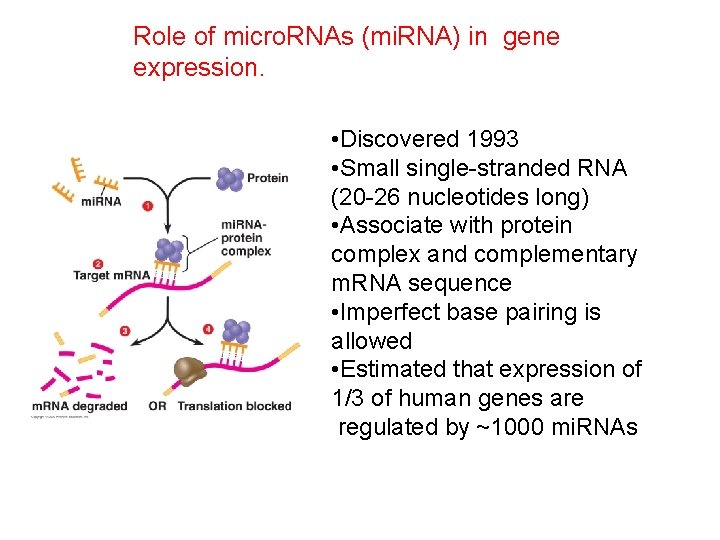
Role of micro. RNAs (mi. RNA) in gene expression. • Discovered 1993 • Small single-stranded RNA (20 -26 nucleotides long) • Associate with protein complex and complementary m. RNA sequence • Imperfect base pairing is allowed • Estimated that expression of 1/3 of human genes are regulated by ~1000 mi. RNAs

Gene expression regulated at later stages. • m. RNA degradation: t 1/2 of transcripts variable. prokaryotic: very short, minutes, because m. RNA lasting for too long could have negative impact on environmental change. eukaryotic: variable, hours to months and can be regulated particularly by hormones • translation highly regulated inhibitory protein blocks transcription of hemoglobin m. RNA if heme supply is low (iron containing compund that binds oxygen)
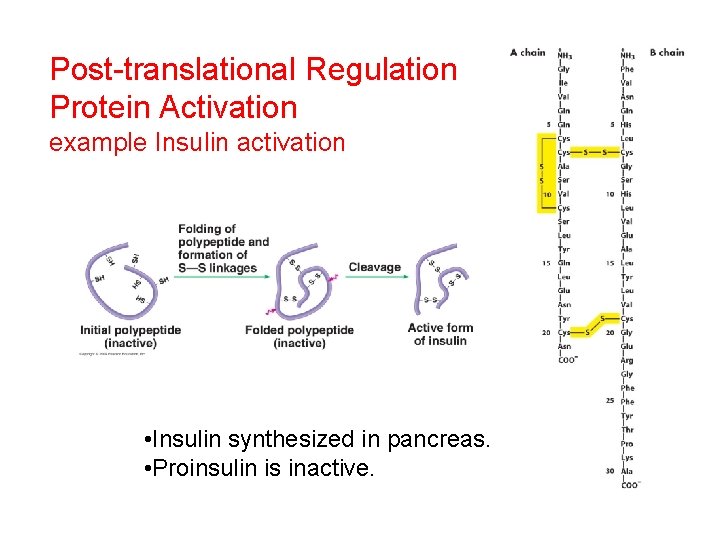
Post-translational Regulation Protein Activation example Insulin activation • Insulin synthesized in pancreas. • Proinsulin is inactive.
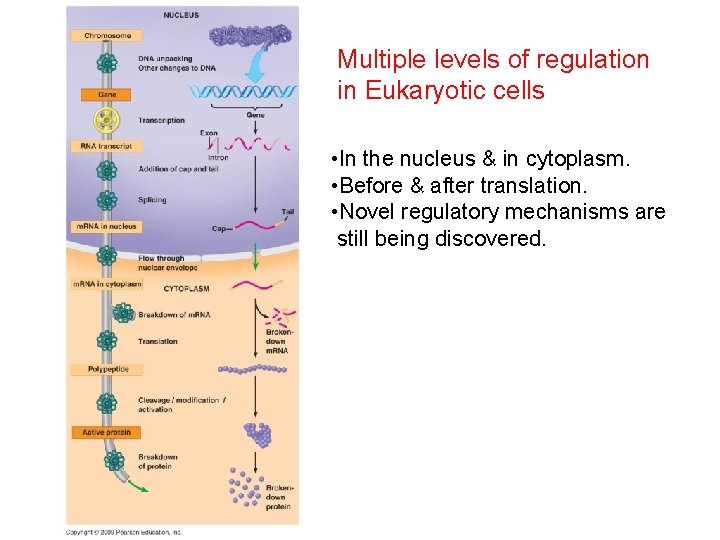
Multiple levels of regulation in Eukaryotic cells • In the nucleus & in cytoplasm. • Before & after translation. • Novel regulatory mechanisms are still being discovered.
- Slides: 38