Molecular Dynamics simulations Bert de Groot Max Planck
Molecular Dynamics simulations Bert de Groot Max Planck institute for biophysical chemistry Göttingen, Germany

Molecular Dynamics Simulations Schrödinger equation Born-Oppenheimer approximation Nucleic motion described classically Empirical force field
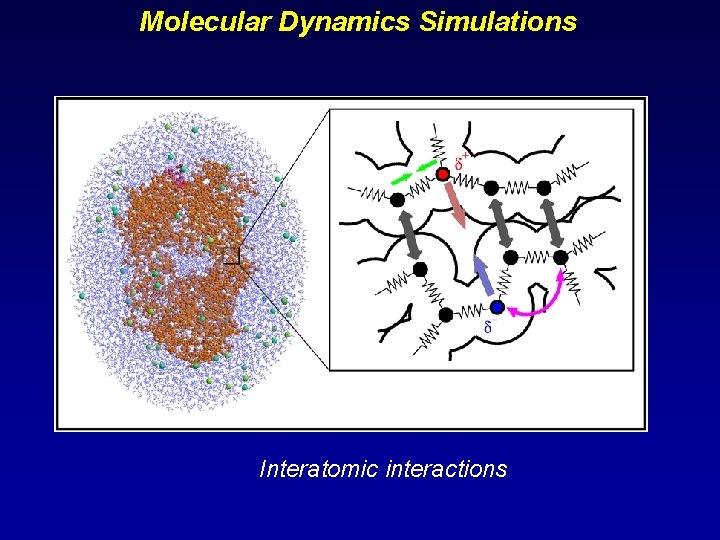
Molecular Dynamics Simulations Interatomic interactions
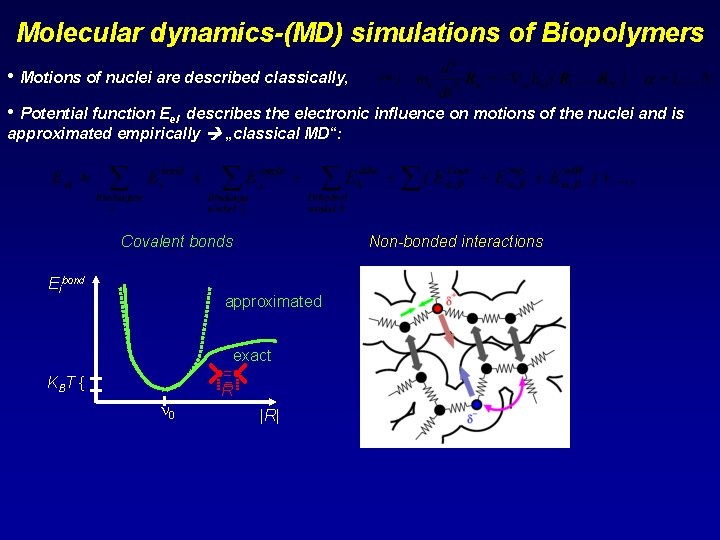
Molecular dynamics-(MD) simulations of Biopolymers • Motions of nuclei are described classically, • Potential function Eel describes the electronic influence on motions of the nuclei and is approximated empirically „classical MD“: Covalent bonds Eibond Non-bonded interactions approximated exact = = R KBT { 0 |R|
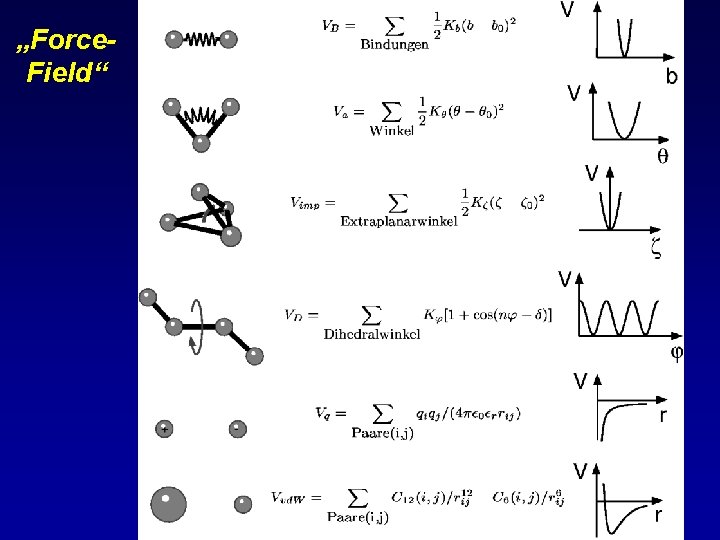
„Force. Field“

Molecular Dynamics Simulation Molecule: (classical) N-particle system Newtonian equations of motion: with Integrate numerically via the „leapfrog“ scheme: with Δt 1 fs! (equivalent to the Verlet algorithm)
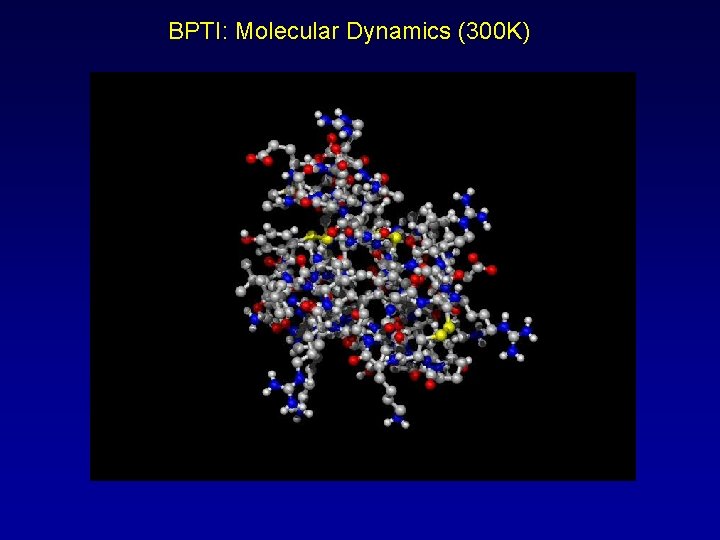
BPTI: Molecular Dynamics (300 K)

Computational task: Solve the Newtonian equations of motion:
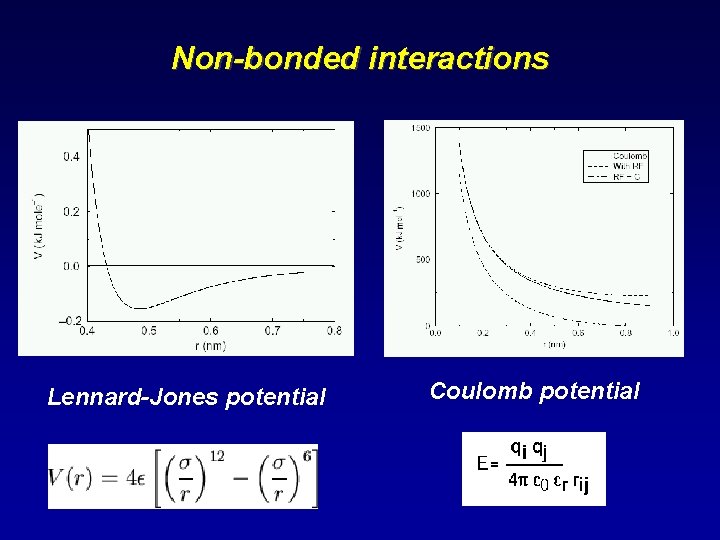
Non-bonded interactions Lennard-Jones potential Coulomb potential
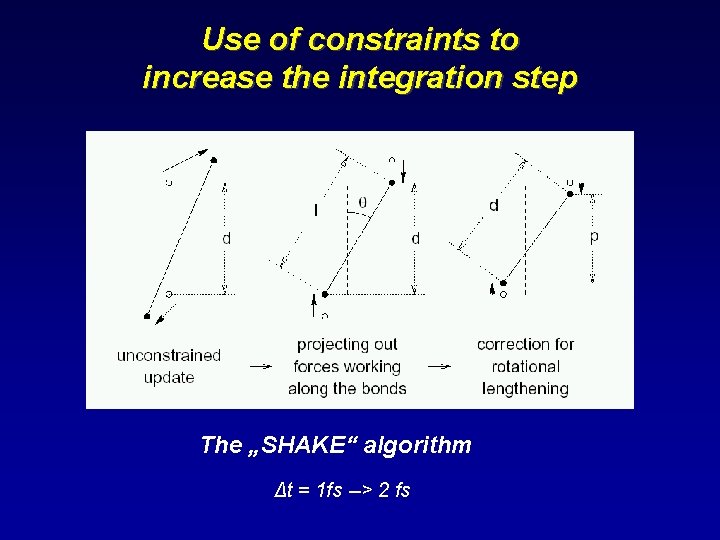
Use of constraints to increase the integration step The „SHAKE“ algorithm Δt = 1 fs --> 2 fs

Molecular dynamics is very expensive. . . Example: F 1 -ATPase in water (183 674 atoms), 1 nanosecond: 106 integration steps 8. 4 * 1011 flop per step [n(n-1)/2 interactions] total: 8. 4 * 1017 flop on a 100 Mflop/s workstation: ca 250 years . . . but performance has been improved by use of: multiple time stepping ca. 25 years + structure adapted multipole methods ca. 6 years + FAMUSAMM ca. 2 years + parallel computers ca. 55 days

Limits of MD-Simulations • classical description: chemical reactions not described poor description of H-atoms (proton-transfer) poor description of low-T (quantum) effects simplified electrostatic model simplified force field • only small systems accessible (104. . . 106 atoms) • only short time spans accessible (ps. . . μs)

MD-Experiments with Argon Gas

Role of environment - solvent explicit or implicit? box or droplet?

Surface (tension) effects? periodic boundary conditions and the minimum image convention
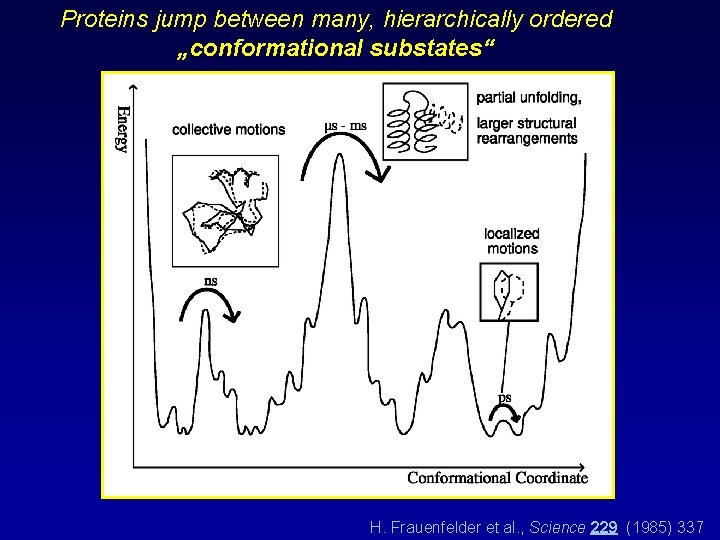
Proteins jump between many, hierarchically ordered „conformational substates“ H. Frauenfelder et al. , Science 229 (1985) 337

Reversible Folding Dynamics of a β-Peptide X. Daura, B. Jaun, D. Seebach, W. F. van Gunsteren, A. E. Mark, J. Mol. Biol. 280 (1998) 925

MD Simulations • external coupling: temperature (potential truncation, integration errors) pressure (density equilibration) system translation/rotation • analysis energies (individual terms, pressure, temperature) coordinates (numerical analysis, visual inspection!) mechanisms

- Slides: 20