Molecular Dynamics of Naproxen LiquidLiquid Phase Separation Lubna
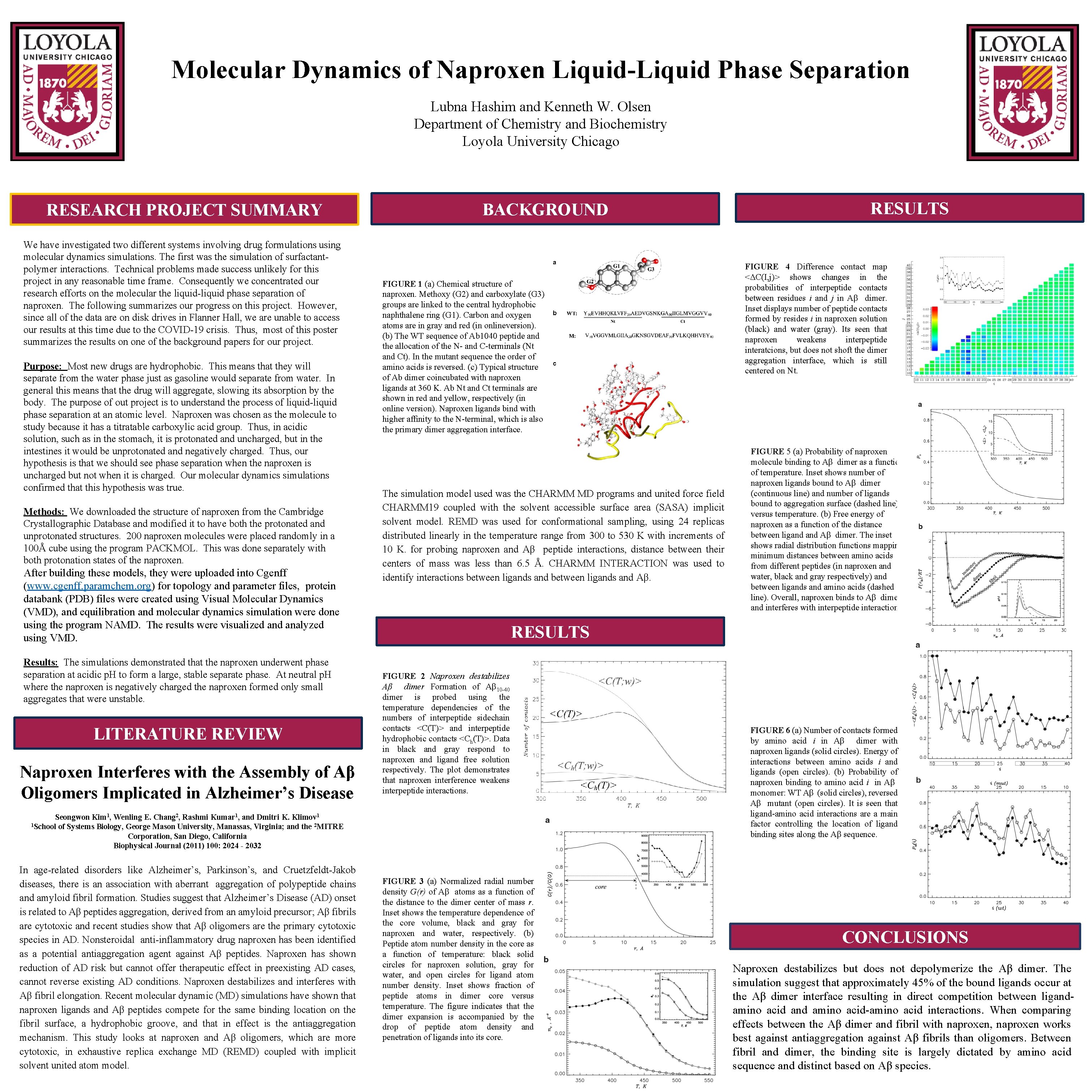
Molecular Dynamics of Naproxen Liquid-Liquid Phase Separation Lubna Hashim and Kenneth W. Olsen Department of Chemistry and Biochemistry Loyola University Chicago RESEARCH PROJECT SUMMARY We have investigated two different systems involving drug formulations using molecular dynamics simulations. The first was the simulation of surfactantpolymer interactions. Technical problems made success unlikely for this project in any reasonable time frame. Consequently we concentrated our research efforts on the molecular the liquid-liquid phase separation of naproxen. The following summarizes our progress on this project. However, since all of the data are on disk drives in Flanner Hall, we are unable to access our results at this time due to the COVID-19 crisis. Thus, most of this poster summarizes the results on one of the background papers for our project. Purpose: Most new drugs are hydrophobic. This means that they will separate from the water phase just as gasoline would separate from water. In general this means that the drug will aggregate, slowing its absorption by the body. The purpose of out project is to understand the process of liquid-liquid phase separation at an atomic level. Naproxen was chosen as the molecule to study because it has a titratable carboxylic acid group. Thus, in acidic solution, such as in the stomach, it is protonated and uncharged, but in the intestines it would be unprotonated and negatively charged. Thus, our hypothesis is that we should see phase separation when the naproxen is uncharged but not when it is charged. Our molecular dynamics simulations confirmed that this hypothesis was true. Methods: We downloaded the structure of naproxen from the Cambridge Crystallographic Database and modified it to have both the protonated and unprotonated structures. 200 naproxen molecules were placed randomly in a 100Å cube using the program PACKMOL. This was done separately with both protonation states of the naproxen. After building these models, they were uploaded into Cgenff (www. cgenff. paramchem. org) for topology and parameter files, protein databank (PDB) files were created using Visual Molecular Dynamics (VMD), and equilibration and molecular dynamics simulation were done using the program NAMD. The results were visualized analyzed using VMD. Results: The simulations demonstrated that the naproxen underwent phase separation at acidic p. H to form a large, stable separate phase. At neutral p. H where the naproxen is negatively charged the naproxen formed only small aggregates that were unstable. LITERATURE REVIEW Naproxen Interferes with the Assembly of Aβ Oligomers Implicated in Alzheimer’s Disease BACKGROUND FIGURE 1 (a) Chemical structure of naproxen. Methoxy (G 2) and carboxylate (G 3) groups are linked to the central hydrophobic naphthalene ring (G 1). Carbon and oxygen atoms are in gray and red (in onlineversion). (b) The WT sequence of Ab 1040 peptide and the allocation of the N- and C-terminals (Nt and Ct). In the mutant sequence the order of amino acids is reversed. (c) Typical structure of Ab dimer coincubated with naproxen ligands at 360 K. Ab Nt and Ct terminals are shown in red and yellow, respectively (in online version). Naproxen ligands bind with higher affinity to the N-terminal, which is also the primary dimer aggregation interface. The simulation model used was the CHARMM MD programs and united force field CHARMM 19 coupled with the solvent accessible surface area (SASA) implicit solvent model. REMD was used for conformational sampling, using 24 replicas distributed linearly in the temperature range from 300 to 530 K with increments of 10 K. for probing naproxen and Aβ peptide interactions, distance between their centers of mass was less than 6. 5 Å. CHARMM INTERACTION was used to identify interactions between ligands and Aβ. FIGURE 4 Difference contact map <ΔC(I, j)> shows changes in the probabilities of interpeptide contacts between residues i and j in Aβ dimer. Inset displays number of peptide contacts formed by resides i in naproxen solution (black) and water (gray). Its seen that naproxen weakens interpeptide interatcions, but does not shoft the dimer aggregation interface, which is still centered on Nt. FIGURE 5 (a) Probability of naproxen molecule binding to Aβ dimer as a function of temperature. Inset shows number of naproxen ligands bound to Aβ dimer (continuous line) and number of ligands bound to aggregation surface (dashed line) versus temperature. (b) Free energy of naproxen as a function of the distance between ligand Aβ dimer. The inset shows radial distribution functions mapping minimum distances between amino acids from different peptides (in naproxen and water, black and gray respectively) and between ligands and amino acids (dashed line). Overall, naproxen binds to Aβ dimer and interferes with interpeptide interactions. RESULTS FIGURE 2 Naproxen destabilizes Aβ dimer Formation of Aβ 10 -40 dimer is probed using the temperature dependencies of the numbers of interpeptide sidechain contacts <C(T)> and interpeptide hydrophobic contacts <Ch(T)>. Data in black and gray respond to naproxen and ligand free solution respectively. The plot demonstrates that naproxen interference weakens interpeptide interactions. Seongwon Kim 1, Wenling E. Chang 2, Rashmi Kumar 1, and Dmitri K. Klimov 1 1 School of Systems Biology, George Mason University, Manassas, Virginia; and the 2 MITRE Corporation, San Diego, California Biophysical Journal (2011) 100: 2024 - 2032 In age-related disorders like Alzheimer’s, Parkinson’s, and Cruetzfeldt-Jakob diseases, there is an association with aberrant aggregation of polypeptide chains and amyloid fibril formation. Studies suggest that Alzheimer’s Disease (AD) onset is related to Aβ peptides aggregation, derived from an amyloid precursor; Aβ fibrils are cytotoxic and recent studies show that Aβ oligomers are the primary cytotoxic species in AD. Nonsteroidal anti-inflammatory drug naproxen has been identified as a potential antiaggregation agent against Aβ peptides. Naproxen has shown reduction of AD risk but cannot offer therapeutic effect in preexisting AD cases, cannot reverse existing AD conditions. Naproxen destabilizes and interferes with Aβ fibril elongation. Recent molecular dynamic (MD) simulations have shown that naproxen ligands and Aβ peptides compete for the same binding location on the fibril surface, a hydrophobic groove, and that in effect is the antiaggregation mechanism. This study looks at naproxen and Aβ oligomers, which are more cytotoxic, in exhaustive replica exchange MD (REMD) coupled with implicit solvent united atom model. RESULTS FIGURE 3 (a) Normalized radial number density G(r) of Aβ atoms as a function of the distance to the dimer center of mass r. Inset shows the temperature dependence of the core volume, black and gray for naproxen and water, respectively. (b) Peptide atom number density in the core as a function of temperature: black solid circles for naproxen solution, gray for water, and open circles for ligand atom number density. Inset shows fraction of peptide atoms in dimer core versus temperature. The figure indicates that the dimer expansion is accompanied by the drop of peptide atom density and penetration of ligands into its core. FIGURE 6 (a) Number of contacts formed by amino acid i in Aβ dimer with naproxen ligands (solid circles). Energy of interactions between amino acids i and ligands (open circles). (b) Probability of naproxen binding to amino acid i in Aβ monomer: WT Aβ (solid circles), reversed Aβ mutant (open circles). It is seen that ligand-amino acid interactions are a main factor controlling the location of ligand binding sites along the Aβ sequence. CONCLUSIONS Naproxen destabilizes but does not depolymerize the Aβ dimer. The simulation suggest that approximately 45% of the bound ligands occur at the Aβ dimer interface resulting in direct competition between ligandamino acid and amino acid-amino acid interactions. When comparing effects between the Aβ dimer and fibril with naproxen, naproxen works best against antiaggregation against Aβ fibrils than oligomers. Between fibril and dimer, the binding site is largely dictated by amino acid sequence and distinct based on Aβ species.
- Slides: 1