Molecular Cloning Gene Cloning The basics Introduction to

Molecular Cloning Gene Cloning The basics

Introduction to DNA Cloning • 1970 s: Gene cloning became a reality – Clone – a molecule, cell, or organism that was produced from another single entity • Made possible by the discovery of: – Restriction Enzymes – DNA cutting enzymes (molecular scissors) – Plasmid DNA Vectors – circular form of self-replicating DNA • Can be manipulated to carry and clone other pieces of DNA
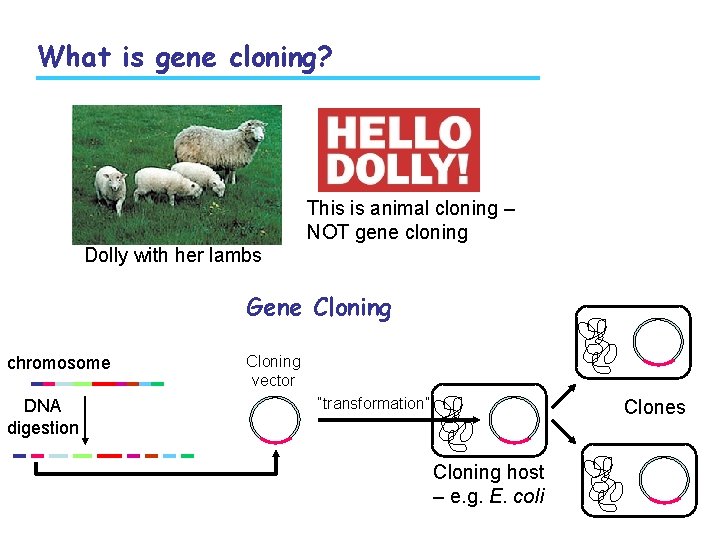
What is gene cloning? This is animal cloning – NOT gene cloning Dolly with her lambs Gene Cloning chromosome DNA digestion Cloning vector “transformation” Clones Cloning host – e. g. E. coli

Gene Cloning • Techniques for gene cloning enable scientists to prepare multiple identical copies of gene-sized pieces of DNA. • Most methods for cloning pieces of DNA share certain general features. – For example, a foreign gene is inserted into a bacterial plasmid and this recombinant DNA molecule is returned to a bacterial cell. – Every time this cell reproduces, the recombinant plasmid is replicated as well and passed on to its descendents. – Under suitable conditions, the bacterial clone will make the protein encoded by the foreign gene.
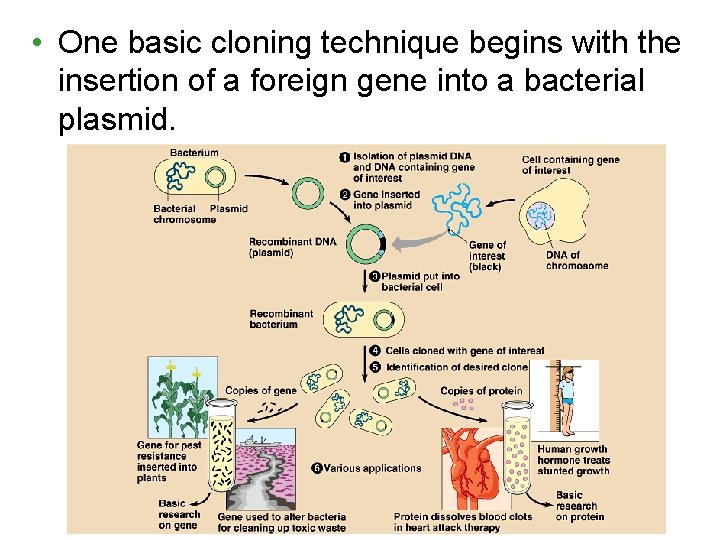
• One basic cloning technique begins with the insertion of a foreign gene into a bacterial plasmid.
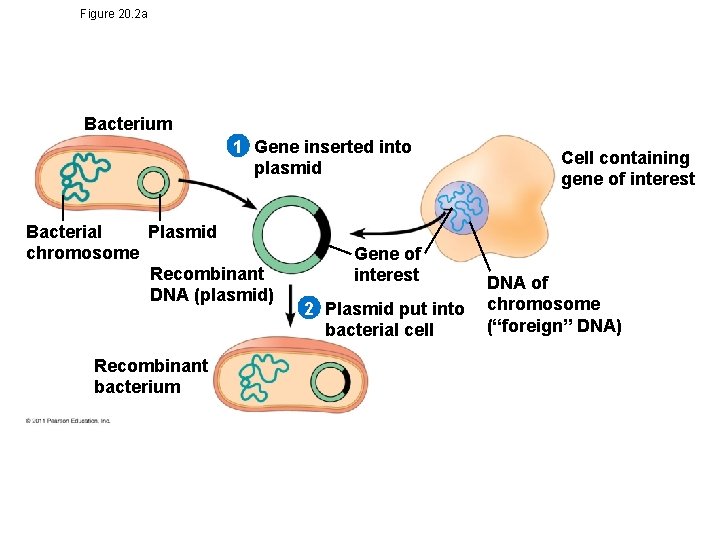
Figure 20. 2 a Bacterium 1 Gene inserted into plasmid Bacterial Plasmid chromosome Recombinant DNA (plasmid) Recombinant bacterium Gene of interest 2 Plasmid put into bacterial cell Cell containing gene of interest DNA of chromosome (“foreign” DNA)

• One goal may be to produce a protein product for use. • A second goal may be to prepare many copies of the gene itself. – This may enable scientists to determine the gene’s nucleotide sequence or provide an organism with a new metabolic capability by transferring a gene from another organism.

Restriction Enzymes • In nature, bacteria use restriction enzymes to cut foreign DNA, such as from phages or other bacteria. • Most restrictions enzymes are very specific, recognizing short DNA nucleotide sequences and cutting at specific point in these sequences.

• Each restriction site reads same forward and backward on opposite strands of DNA – These are often a symmetrical series of four to eight bases on both strands running in opposite directions. – If the restriction site on one strand is 3’-CTTAAG-5’, the complementary strand is 5’-GAATTC-3 • Restriction enzymes cut covalent phosphodiester bonds of both strands, creating single-stranded ends, sticky ends. – These extensions will form hydrogen-bonded base pairs with complementary single-stranded stretches on other DNA molecules cut with the same restriction enzyme

Using Restriction Enzymes to Make Recombinant DNA • Bacterial restriction enzymes cut DNA molecules at specific DNA sequences called restriction sites • A restriction enzyme usually makes many cuts, yielding restriction fragments • The most useful restriction enzymes cut DNA in a staggered way, producing fragments with “sticky ends. ” • Sticky ends can bond with complementary sticky ends of other fragments • DNA ligase is an enzyme that seals the bonds between restriction fragments • Why restriction enzymes don't digest bacteria DNA?
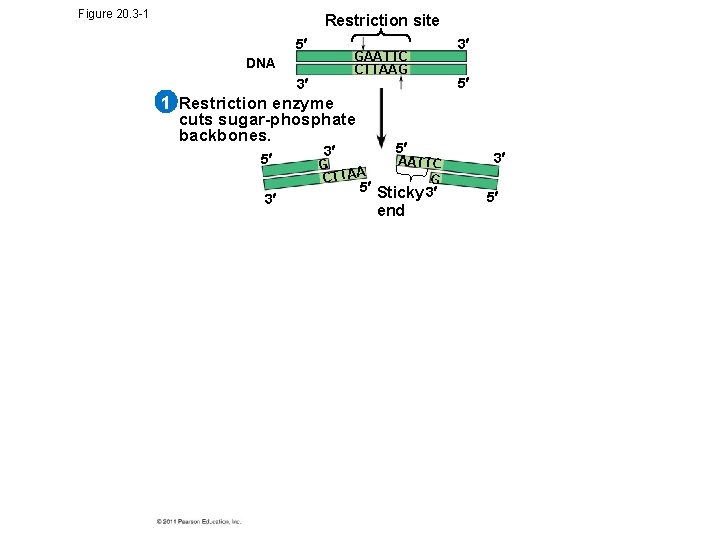
Figure 20. 3 -1 Restriction site 5 GAATTC CTTAAG DNA 3 3 5 1 Restriction enzyme cuts sugar-phosphate backbones. 5 3 3 5 G CTTAA AATTC G 5 Sticky 3 end 3 5
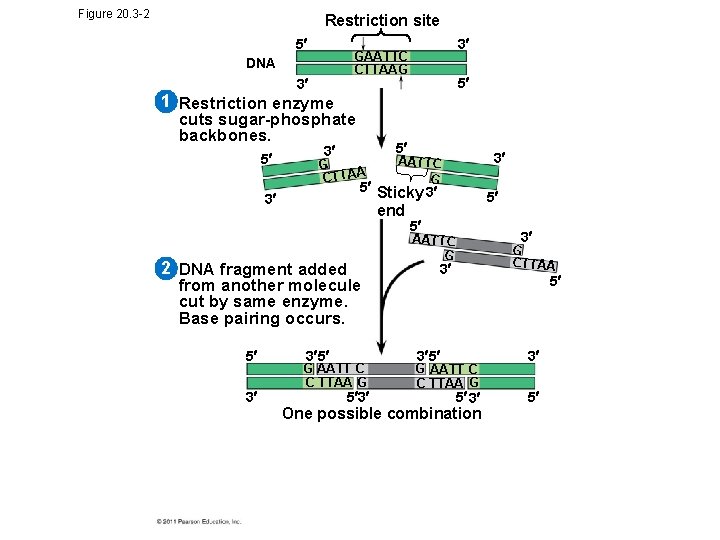
Figure 20. 3 -2 Restriction site 5 3 GAATTC CTTAAG DNA 3 5 1 Restriction enzyme cuts sugar-phosphate backbones. 5 5 3 G CTTAA 5 Sticky 3 3 end 2 DNA fragment added from another molecule cut by same enzyme. Base pairing occurs. 5 3 3 AATTC G 3 5 G AATT C C TTAA G 5 3 5 5 3 AATTC G G CTTAA 3 5 3 5 G AATT C C TTAA G 5 3 One possible combination 3 5
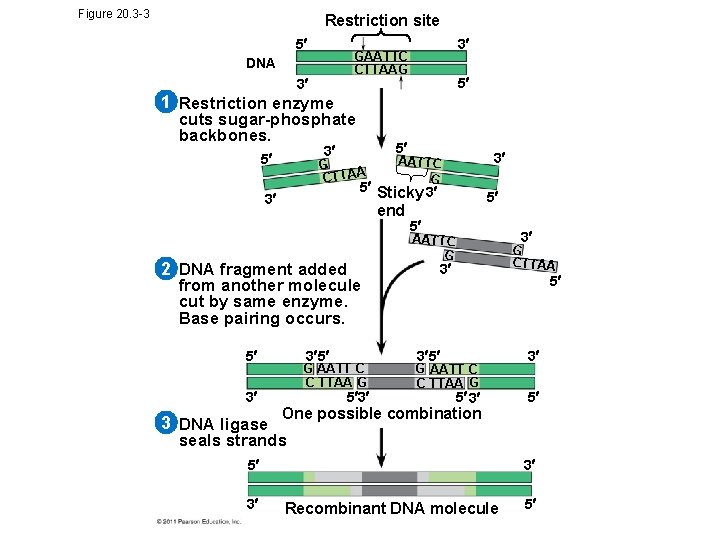
Figure 20. 3 -3 Restriction site 5 3 GAATTC CTTAAG DNA 3 5 1 Restriction enzyme cuts sugar-phosphate backbones. 5 3 5 G CTTAA 5 Sticky 3 3 end 2 DNA fragment added from another molecule cut by same enzyme. Base pairing occurs. 5 3 5 G AATT C C TTAA G 3 3 DNA ligase 3 AATTC G 5 3 5 5 3 AATTC G G CTTAA 3 5 3 5 G AATT C C TTAA G 5 3 3 5 One possible combination seals strands 5 3 3 Recombinant DNA molecule 5

Cloning a Gene in a Bacterial Plasmid • In gene cloning, the original plasmid is called a cloning vector • A cloning vector is a DNA molecule that can carry foreign DNA into a host cell and replicate there • Recombinant plasmids are produced by splicing restriction fragments from foreign DNA into plasmids. – A plasmid is a circular piece of DNA found in bacteria and contain genes. – Plasmids can be used to insert DNA from another organism into a bacterial cell. • Then, as a bacterium carrying a recombinant plasmid reproduces, the plasmid replicates within it.

• Bacteria are most commonly used as host cells for gene cloning because DNA can be easily isolated and reintroduced into their cells. • Bacteria cultures also grow quickly, rapidly replicating the foreign genes. • Bacteria will also produce large amounts of the protein of interest
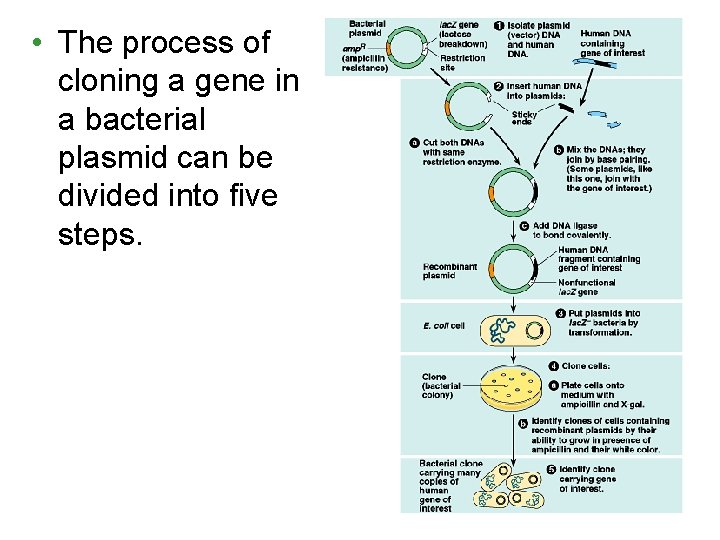
• The process of cloning a gene in a bacterial plasmid can be divided into five steps.

1. Isolation of vector and gene-source DNA. • The source of DNA may come from human tissue cells. • The source of the plasmid is typically E. coli. – This plasmid carries useful genes, such as amp. R, conferring resistance to the antibiotic ampicillin

2. Insertion of DNA into the vector. • By digesting both the plasmid and human DNA with the same restriction enzyme we can create thousands of human DNA fragments, one fragment with the gene that we want, and with compatible sticky ends on bacterial plasmids. • After mixing, the human fragments and cut plasmids form complementary pairs that are then joined by DNA ligase. • This creates a mixture of recombinant DNA molecules.

3. Introduction of the cloning vector into cells. • Bacterial cells take up the recombinant plasmids by transformation. • This creates a diverse pool of bacteria, some bacteria that have taken up the desired recombinant plasmid DNA, other bacteria that have taken up other DNA, both recombinant and nonrecombinant.

4. Cloning of cells (and foreign genes). • We can plate out the transformed bacteria on a solid nutrient medium containing ampicillin. – Only bacteria that have the ampicillinresistance plasmid will grow. 5. Identifying cell clones with the right gene. • In the final step, we will sort through the thousands of bacterial colonies with foreign DNA to find those containing our gene of interest.

• One technique to test the clones of interest is to use DNA or Nucleic acid hybridization. – This technique depends on base pairing between our gene and a short piece of DNA or RNA with a complementary sequence to the gene called a Probe, – The sequence of the RNA or DNA probe depends on knowledge of at least part of the sequence of our gene. – A radioactive or fluorescent tag labels the probe so that if it bind with our gene we can detect it’s presence.

The polymerase chain reaction (PCR) clones DNA entirely in vitro • When the source of DNA is scanty or impure, the polymerase chain reaction (PCR) is quicker and more selective. Its limitation is that PCR only produces short DNA segments within a gene and not entire genes. • This technique can quickly amplify any piece of DNA without using cells. • Devised in 1985, PCR has had a major impact on biological research and technology.

• Polymerase Chain Reaction – Developed in the mid-1980 s by Kary Mullis – Technique for making copies, or amplifying, a specific sequence of DNA in a short period of time – Process • Target DNA to be amplified is added to a tube, mixed with nucleotides (d. ATP, d. CTP, d. GTP, d. TTP), buffer, and DNA polymerase. • Paired set of Forward and Reverse Primers are added – short single-stranded DNA oligonucleotides (20– 30 bp long) – Primers are complementary to nucleotides flanking opposite ends of target DNA • Reaction tube is placed in an instrument called a thermocycler

• PCR Process continued – Thermocycler will take DNA through a series of reactions called a PCR cycle – Each cycle consists of three stages 1. Denaturation – heat to 94 °C to 96 °C 2. Annealing (hybridization) – in which primers H bond with complementary bases at the opposite ends of target sequence at 55 °C to 65 °C 3. Extension (elongation) – DNA Pol copies target DNA at 70 to 75 °C – At the end of one cycle, the amount of DNA has doubled – Cycles are repeated 20– 30 times
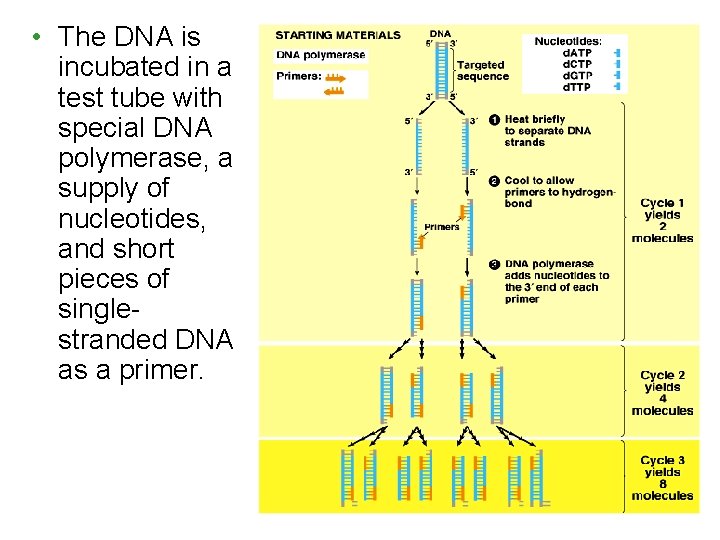
• The DNA is incubated in a test tube with special DNA polymerase, a supply of nucleotides, and short pieces of singlestranded DNA as a primer.

• PCR has amplified DNA from a variety of sources: – fragments of ancient DNA from a 40, 000 -yearold frozen wooly mammoth, – DNA from tiny amount of blood or semen found at the scenes of violent crimes, – DNA from single embryonic cells for rapid prenatal diagnosis of genetic disorders, – DNA of viral genes from cells infected with difficult-to-detect viruses such as HIV.

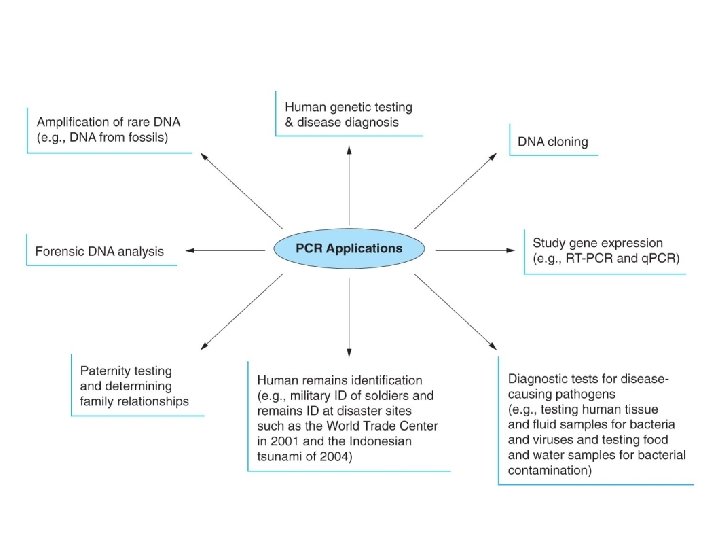
• To calculate the number of copies of target DNA starting with 1 molecule of DNA use this equation 2 N in which N represents number of PCR cycles • Applications – – – Making DNA probes Studying gene expression Detection of viral and bacterial infections Diagnosis of genetic conditions Detection of trace amounts of DNA from tissue found at crime scene – Detection of DNA from fossilized dinosaur tissue
- Slides: 28