Molecular Cloning Definitions q Cloning q Obtaining a
Molecular Cloning

Definitions q Cloning : q Obtaining a piece of DNA from its original source (Genome) and introducing it in a DNA vector q Sub-cloning: q Transfer of a cloned DNA insert or a part thereof from one vector to another vector q Recombinant Plasmid : q Vector into which foreign DNA was introduced q Recombinant organism q Organism with a recombinant vector

Why clone? q Separate, identify, manipulate or express a specific DNA fragment 3

Step 1 - Separate q Two approaches: q Fragment/digest genomic DNA q Generates a vast number of fragments q May be difficult to find fragment of interest q PCR Amplification q Much less fragments q Much easier to find sequence of interest

DNA Replication & Amplification The Polymerase Chain Reaction
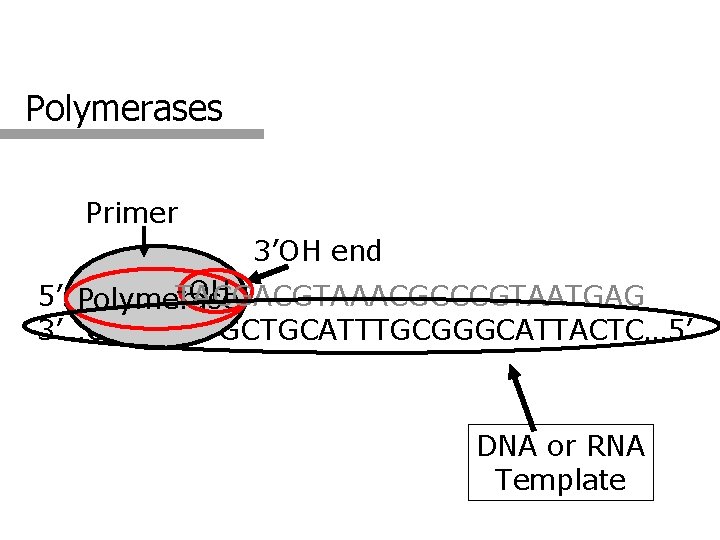
Polymerases Primer 3’OH end -OH 5’…GTACT TACGACGTAAACGCCCGTAATGAG Polymerase 3’…CATGAATGCTGCATTTGCGGGCATTACTC… 5’ DNA or RNA Template

The Polymerase Chain Reaction-PCR q Repetitive replication of a given region of DNA q Allows the exponential amplification of a given region of DNA q Increases the relative representation of the region of interest q Allows the isolation of a given region of DNA 7
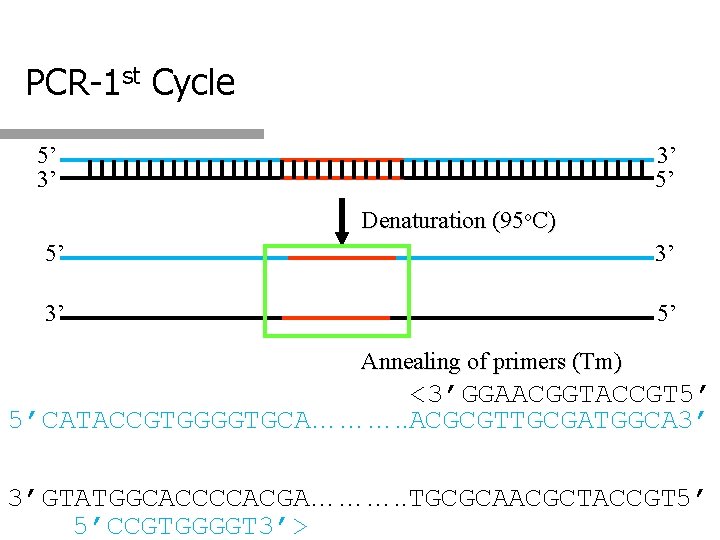
PCR-1 st Cycle 5’ 3’ 3’ 5’ Denaturation (95 o. C) 5’ 3’ 3’ 5’ Annealing of primers (Tm) <3’GGAACGGTACCGT 5’ 5’CATACCGTGGGGTGCA………. . ACGCGTTGCGATGGCA 3’ 3’GTATGGCACCCCACGA………. . TGCGCAACGCTACCGT 5’ 5’CCGTGGGGT 3’>
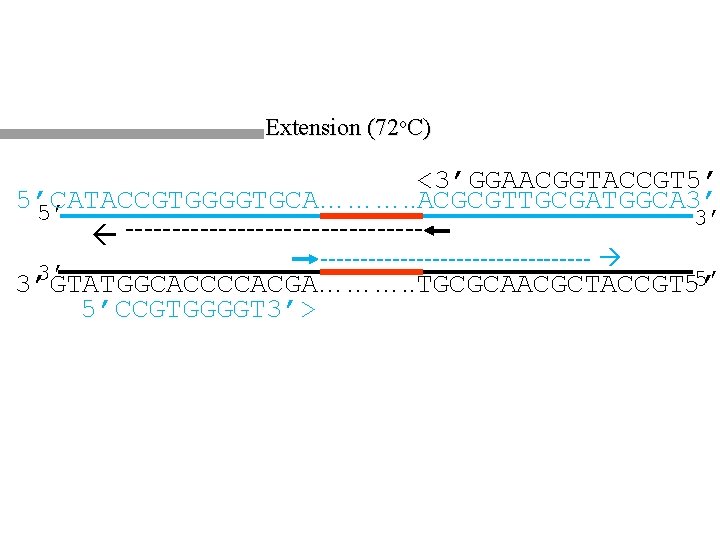
Extension (72 o. C) <3’GGAACGGTACCGT 5’ 5’CATACCGTGGGGTGCA………. . ACGCGTTGCGATGGCA 3’ 5’ 3’ ----------------- 3’ 5’ 3’GTATGGCACCCCACGA………. . TGCGCAACGCTACCGT 5’ 5’CCGTGGGGT 3’> 9

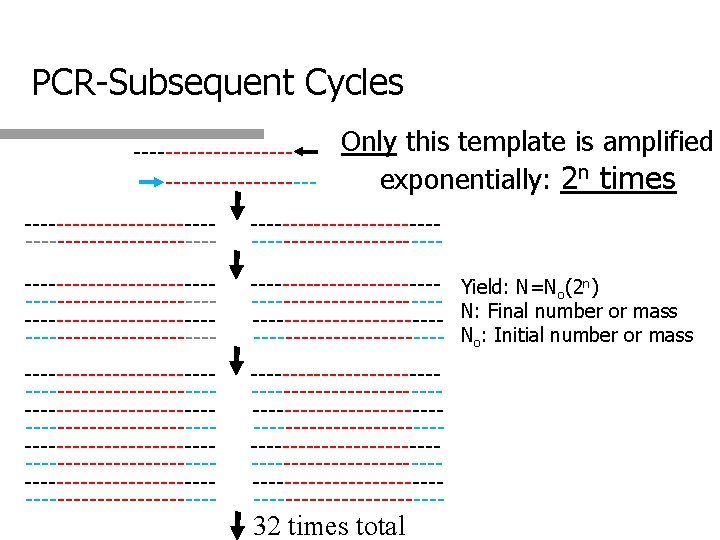

Problem q You wish to amplify a 1 kb sequence from 10 ng of a 10 kb single stranded template with primers spanning positions 12 -37 and 1007 -1037. How many cycles are required to obtain 1µg of the desired product?

Solution q Determine initial mass of desired product q q 10 kb = 10 ng therefore 1 kb = 1 ng or 10 -3µg (Single stranded) q Double stranded would be 2 X 10 -3 µg Determine number of cycles starting from initial mass of desired product q q Determine number of cycles to obtain double stranded desired product q q 1µg = 2 X 10 -3µg(2 n); solve for n – Approx. 9 3 cycles Total: 9+3 = 12

Review of PCR Cycles q PCR Primers: q Short single stranded nucleotide sequences complementary to the targets q 15 -30 nucleotides q Used in excess as compared to target to favor primer annealing rather than template self annealing

Review of PCR Cycles q Annealing: Temperature at which primers anneal to complementary target sequences q Must be below primer Tm q Must be a temperature that allows both primers to anneal q Usually between 55 -75 o. C q q Extension: Carried out at temperature optimum for DNA polymerase q Usually 72 -75 o. C for Taq polymerase q

Primers n Characteristics: n Short oligonucleotides complementary to sequences that flank the region of interest n Establish the point of initiation of replication n Establish the point of termination of replication 16
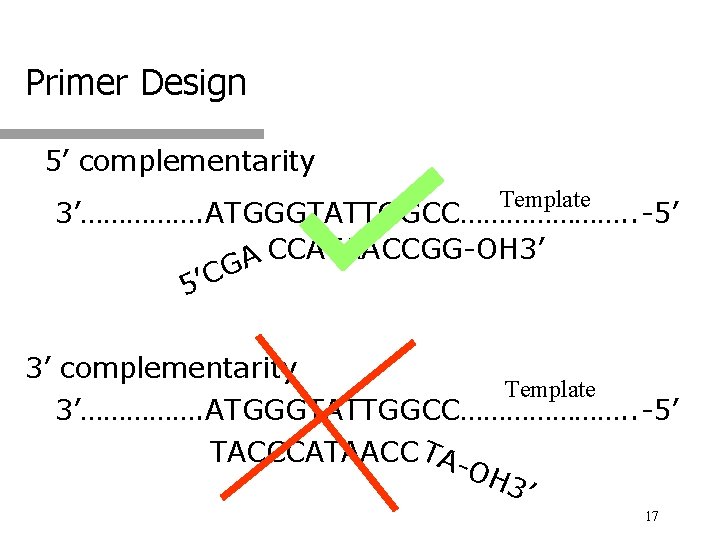
Primer Design 5’ complementarity Template 3’……………. ATGGGTATTGGCC…………………. . -5’ CCATAACCGG-OH 3’ A G C ’ 5 3’ complementarity Template 3’……………. ATGGGTATTGGCC…………………. . -5’ TACCCATAACC TAOH 3’ 17
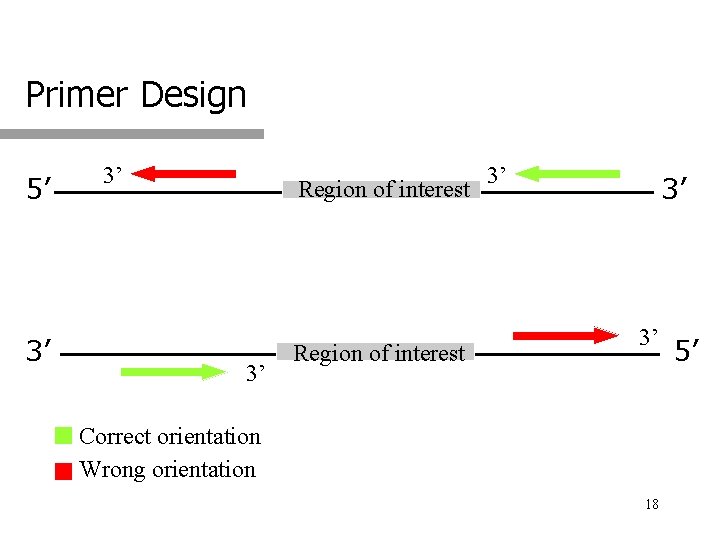
Primer Design 5’ 3’ 3’ Region of interest 3’ 3’ 3’ Correct orientation Wrong orientation 18 5’

Primers 2 primers are required for exponential amplification q Forward primer q q q Reverse primer q q Primer whose sequence is the same as that of the template indicated Primer whose sequence is complementary to that of the template indicated Example 3’-GCGGTT • • • • • ATCGTTA-5’ q q Forward primer: ATTGCTA Reverse primer: CGCCAA

Problem 5’-AAAAAA GGGGGGG-3’ n You wish to amplify the sequence represented by the box. Which primer pair represent the correct orientation to accomplish this? 1234 - AAAAAA TTTTTT GGGGGG CCCCCC 20

Utility of PCR q Amplification and isolation of a given region by changing its relative representation q Between 100 bp and 10 Kpb q Screening to determine the presence of a sequence of interest q Presence or absence of an amplification product q Site directed mutagenesis q Used to add or remove nucleotides from the original template 21
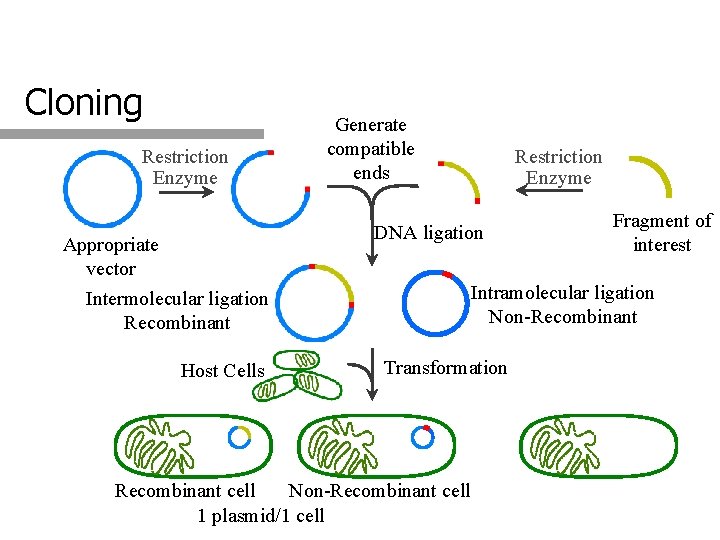
Cloning Restriction Enzyme Generate compatible ends Restriction Enzyme DNA ligation DNA Recombination Appropriate vector Intramolecular ligation Non-Recombinant Intermolecular ligation Recombinant Host Cells Fragment of interest Transformation Recombinant cell Non-Recombinant cell 1 plasmid/1 cell
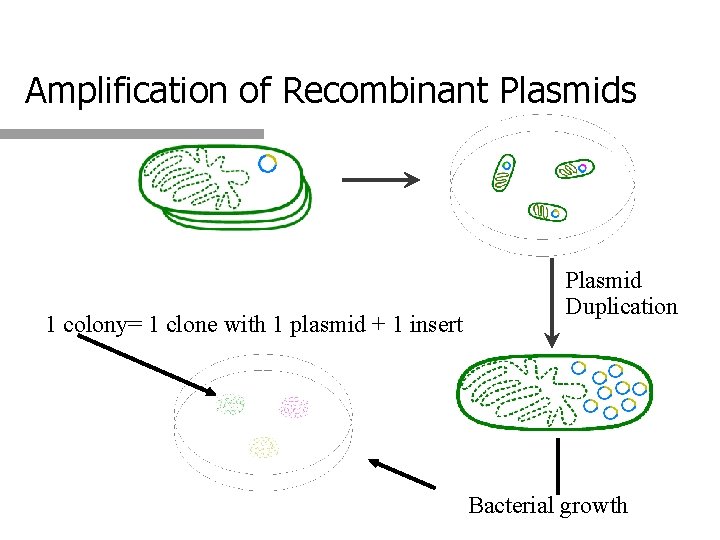
Amplification of Recombinant Plasmids 1 colony= 1 clone with 1 plasmid + 1 insert Plasmid Duplication Bacterial growth

Screening and Identification of Recombinant Plasmid clones q Restriction mapping q PCR
- Slides: 24