Molecular Classification of Cancer Class Discovery and Class





























- Slides: 39

Molecular Classification of Cancer Class Discovery and Class Prediction by Gene Expression Monitoring

Overview n Motivation n Microarray Background n Our Test Case n Class Prediction n Class Discovery

Motivation n Importance of cancer classification n Cancer classification has historically relied on specific biological insights n We will discuss a systematic and unbiased approach for recognizing tumor subtypes

Microarray Background n Microarrays enable simultaneous measurement of the expression levels of thousands of genes in a sample n Microarray: – Glass slide with a matrix of thousands of spots printed on to it – Each spot contains probes which bind to a specific gene

Microarray Background (cont. ) n The process: – DNA samples are taken from the test subjects – Samples are dyed with fluorescent colors and placed on the Microarray – Hybridization of DNA and c. DNA n The result: – Spots in the array are dyed in shades of red to green

Microarray Background (cont. ) n Sample 1 Sample 2 Gene 1 1. 04 2. 08 Gene 2 3. 2 10. 5 Gene 3 3. 34 1. 05 Gene 4 1. 85 0. 09 Microarray data is translated into an n x p table (p – number of genes, n – number of samples)

Demonstration http: //www. bio. davidson. edu/courses/genomics/chip. html

Our Test Case n 38 bone marrow samples from acute leukemia patients (27 ALL, 11 AML) n RNA from the samples was hybridized to microarrays containing probes for 6817 human genes n For each gene, an expression level was obtained

Class Prediction n Initial collection of samples belonging to known classes n Goal: create a “class predictor” to classify new samples – Look for “informative genes” – Make a prediction based on these genes – Test the validity of the predictor

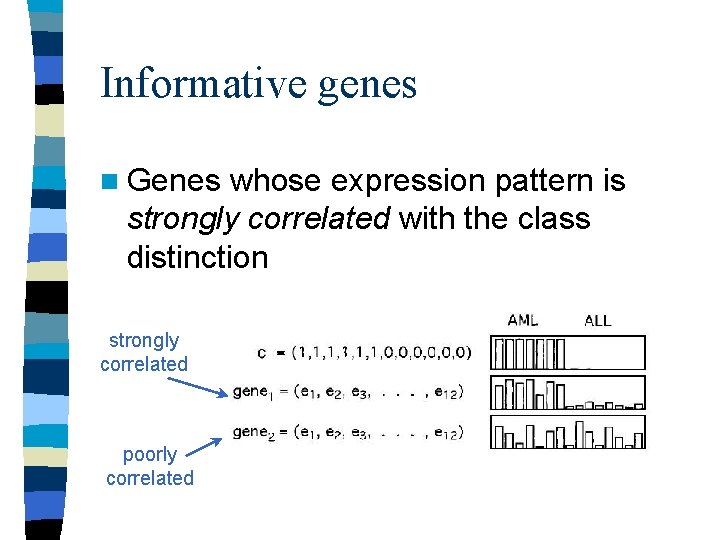
Informative genes n Genes whose expression pattern is strongly correlated with the class distinction strongly correlated poorly correlated

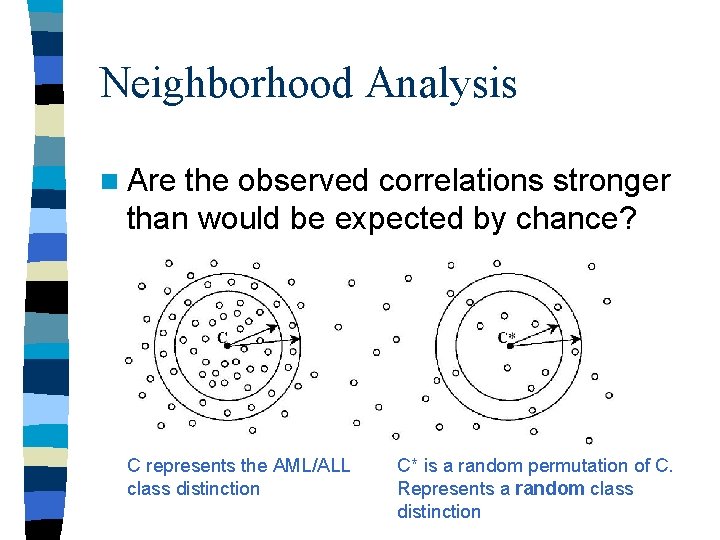
Neighborhood Analysis n Are the observed correlations stronger than would be expected by chance? C represents the AML/ALL class distinction C* is a random permutation of C. Represents a random class distinction

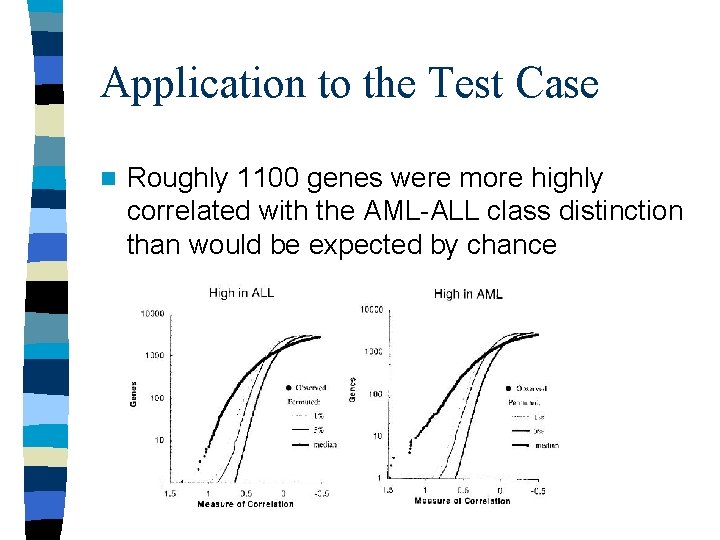
Application to the Test Case n Roughly 1100 genes were more highly correlated with the AML-ALL class distinction than would be expected by chance

Make a Prediction n Use a fixed subset of “informative genes” (most correlated with the class distinction) n Make a prediction on the basis of the expression level of these genes in a new sample

Prediction Algorithm Each gene Gi votes, depending on whether its expression level Xi in the sample is closer to µ or µ n The magnitude of the vote is Wi Vi n AML ALL – Wi reflects how well the gene is correlated with the class distinction – reflects the deviation of Xi from the average of µ and µ ALL AML

Prediction Algorithm (cont. ) n The votes for each class are summed to obtain total votes VAML and VALL

Prediction Algorithm (cont. ) n The prediction strength is calculated: n The sample is assigned to the winning class provided that the PS exceeds a predetermined threshold (0. 3 in the test case)

Testing the Validity of Class Predictors n Cross Validation – withhold a sample – build a predictor based on the remaining samples – predict the class of the withheld sample – repeat for each sample n Assess accuracy on an independent set of samples

Application to the Test Case 50 genes most highly correlated with the AML-ALL distinction were chosen n A class predictor based on these genes was built n

Application to the Test Case n Performance in cross validation: – Out of 38 samples there were 36 predictions and 2 uncertainties (PS < 0. 3) – 100% accuracy – PS median 0. 77

Application to the Test Case (cont. ) n Performance on an independent set of samples: – Out of 34 samples there were 29 predictions and 5 uncertainties (PS < 0. 3) – 100% accuracy – PS median 0. 73

Comments n Why 50 genes? – Large enough to be robust against noise – Small enough to be readily applied in a clinical setting – Predictors based on between 10 to 200 genes all performed well n Genes useful for cancer class prediction may also provide insight into cancer pathogenesis and pharmacology

Comments (cont. ) n Creation of a new predictor involves expression analysis of thousands of genes n Application of the predictor then requires only monitoring the expression level of few informative genes

Class Discovery n Cluster tumors by gene expression – Apply a clustering technique to produce presumed classes n Evaluation of the Classes: – Are the classes meaningful? – Do they reflect true structure?

Clustering Technique - SOMs n SOMs – Self Organizing Maps Well suited for identifying a small number of prominent classes – Find an optimal set of “centroids” – Partition the data set according to the centroids – Each centroid defines a cluster consisting of the data points nearest to it n We won't go into details about the calculation of SOMs

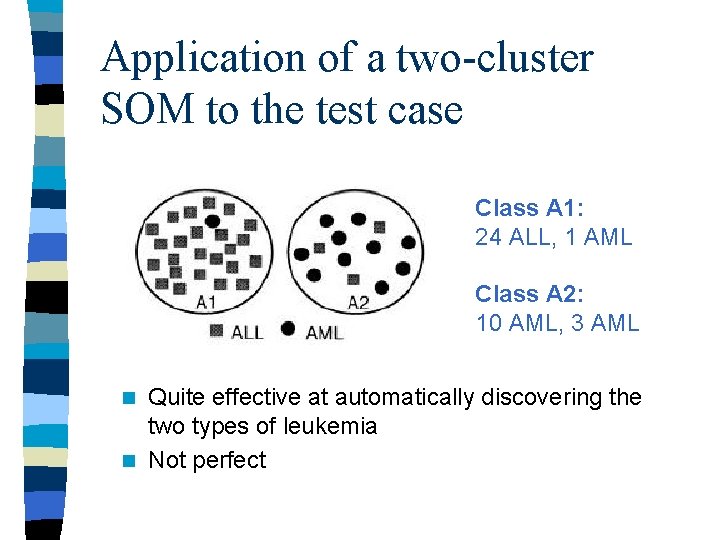
Application of a two-cluster SOM to the test case Class A 1: 24 ALL, 1 AML Class A 2: 10 AML, 3 AML Quite effective at automatically discovering the two types of leukemia n Not perfect n

Evaluation of the Classes n How can we evaluate such classes if the “right” answer is not already known? n Hypothesis: class discovery can be tested by class prediction – If the classes reflect true structure, then a class predictor based on them should perform well n Let’s test this hypothesis. . .

Validity of Predictors Based on A 1 and A 2 n Predictors based on different numbers of informative genes performed well n For example: a 20 -gene predictor

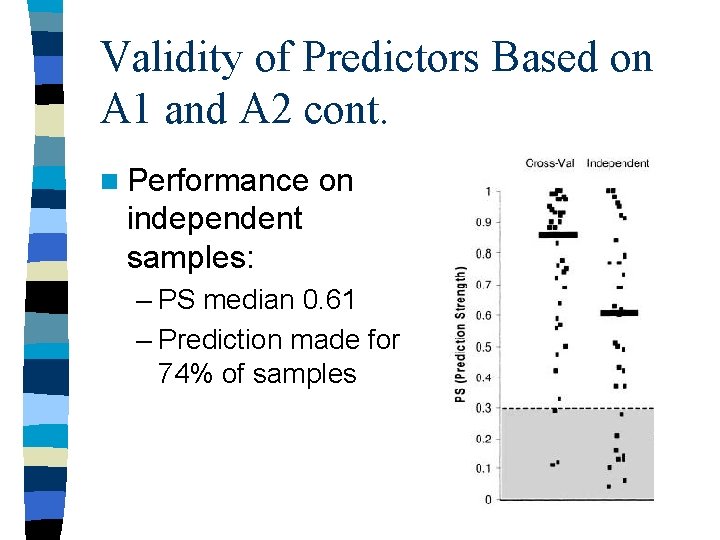
Validity of Predictors Based on A 1 and A 2 cont. n Performance on independent samples: – PS median 0. 61 – Prediction made for 74% of samples

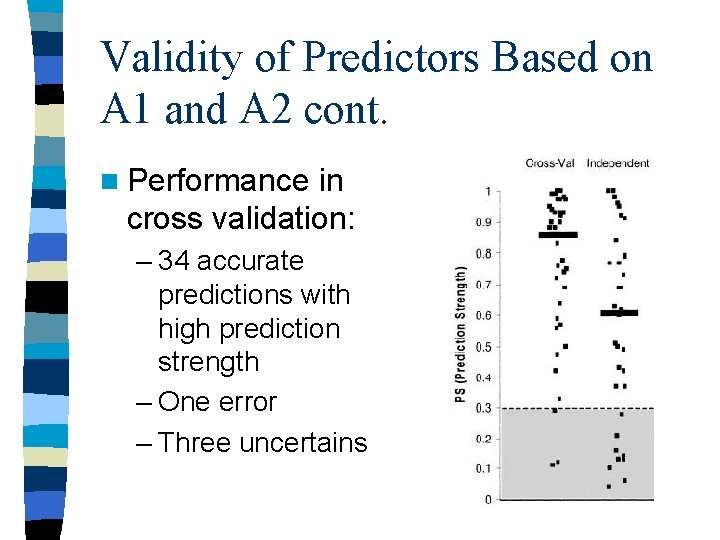
Validity of Predictors Based on A 1 and A 2 cont. n Performance in cross validation: – 34 accurate predictions with high prediction strength – One error – Three uncertains

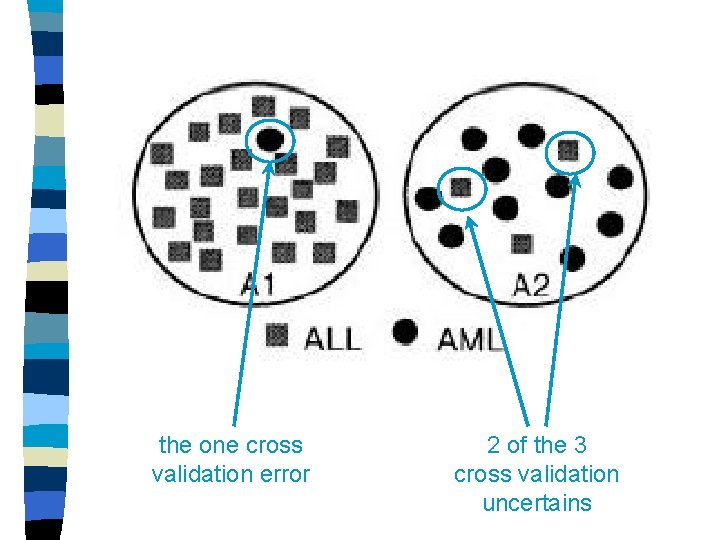
the one cross validation error 2 of the 3 cross validation uncertains

Iterative Procedure n Use a SOM to initially cluster the data n Construct a predictor n Remove samples that are not correctly predicted in cross-validation n Use the remaining samples to generate an improved predictor n Test on an independent data set

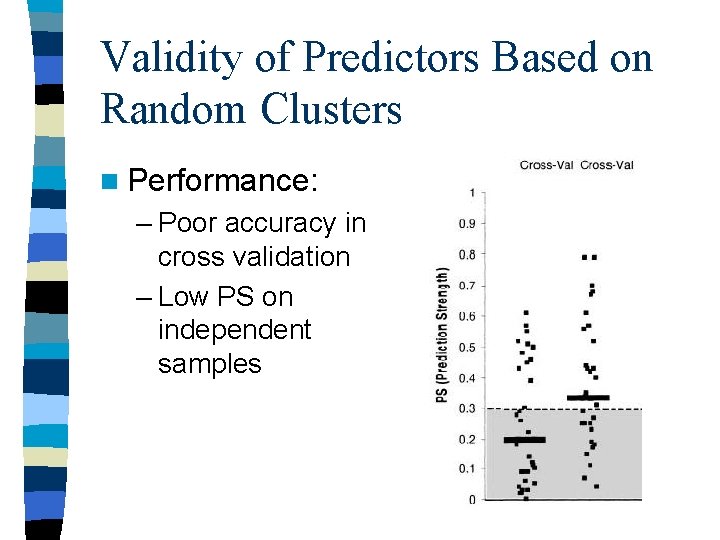
Validity of Predictors Based on Random Clusters n Performance: – Poor accuracy in cross validation – Low PS on independent samples

Conclusion n The AML-ALL distinction could have been automatically discovered and confirmed without previous biological knowledge

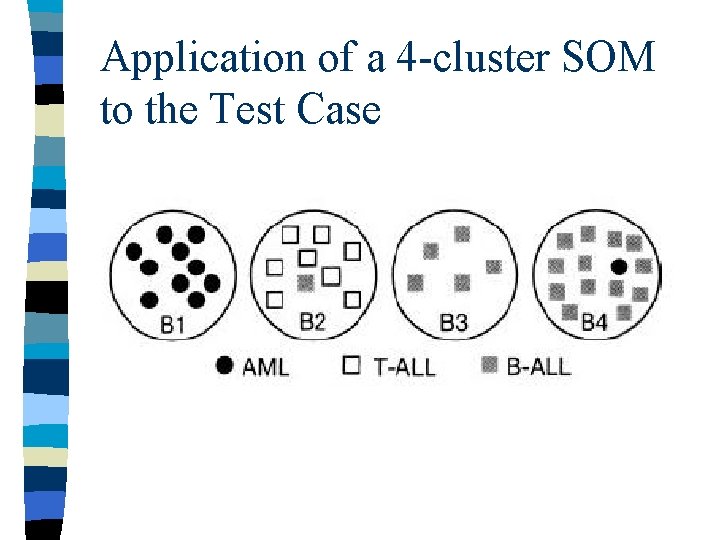
Application of a 4 -cluster SOM to the Test Case

Evaluation of the Classes n Complement approach: – Construct class predictors to distinguish each class from its complement n Pair-wise approach: – Construct class predictors to distinguish between each pair of classes Ci, Cj – Perform cross validation only on samples in Ci and Cj

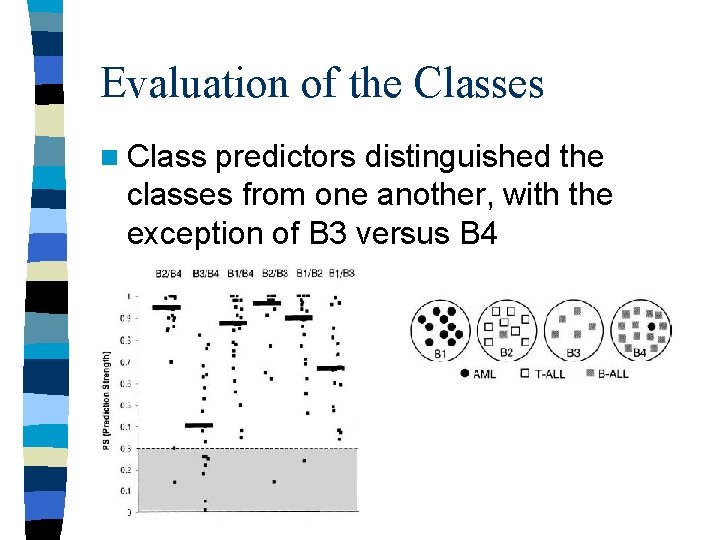
Evaluation of the Classes n Class predictors distinguished the classes from one another, with the exception of B 3 versus B 4

Conclusion n The results suggest the merging of classes B 3 and B 4 n The distinction corresponding to AML, B -ALL and T-ALL was confirmed

Uses of Class Discovery n Identify fundamental subtypes of any cancer n Search for fundamental mechanisms that cut across distinct types of cancers

Questions? n Thank you for listening