Molecular characterization of the Caenorhabditis elegans ALPEnigma gene

Molecular characterization of the Caenorhabditis elegans ALP/Enigma gene alp-1 Caroline R. Mc. Keown, Hsiao-Fen Han, and Mary C. Beckerle Developmental Dynamics Vol 235, Issue 2
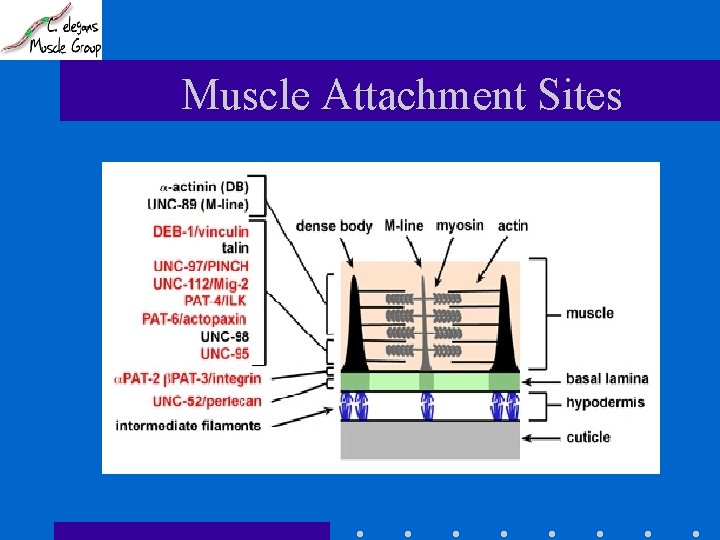
Muscle Attachment Sites
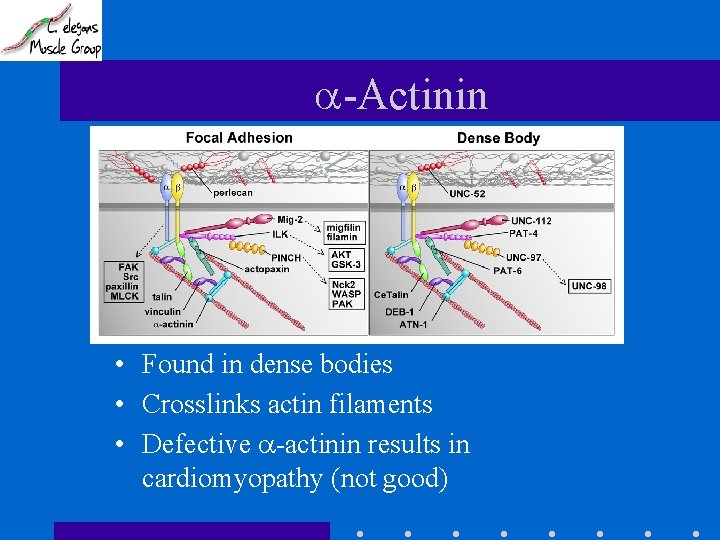
-Actinin • Found in dense bodies • Crosslinks actin filaments • Defective -actinin results in cardiomyopathy (not good)

Mammalian ALP • ALP ( -actinin associated LIM protein) • Family includes at least 7 proteins • ALP (which has subfamilies including ALP, RIL, CLP 36/Elfin, and Mystique) • Enigma • Cypher • All bind either -actinin or tropomyosin and localize to Z-lines

ALP family domains • LIM (LIN-11, Isl-1, MEC-3) • PDZ (PSD 95, Discs-large, ZO-1) • both are known protein binding interfaces

C. elegans ALP is eat-1 • First uncovered in a screen by Leon Avery for mutants with pharyngeal pumping defects • somewhat Unc, also has irregular pharyngeal pumping • SAGE data shows raised expression levels in larval stages • similar expression pattern for -actinin

C. elegans ALP is eat-1

C. elegans ALP is eat-1

ALP-1 Localization • Made GFP translational fusions for alp-1
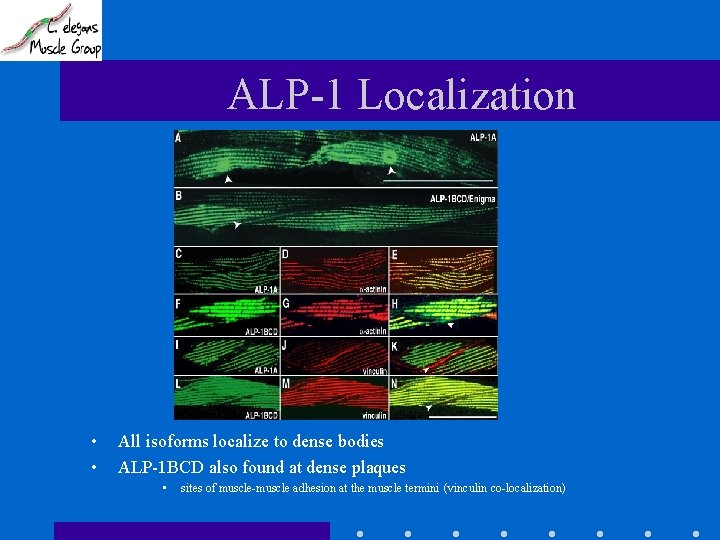
ALP-1 Localization • • All isoforms localize to dense bodies ALP-1 BCD also found at dense plaques • sites of muscle-muscle adhesion at the muscle termini (vinculin co-localization)

PDZ Domain • Sufficient for localization to dense bodies • not sufficient to localize to dense plaques • Made PDZ domain containing (and myc-tagged) construct under influence of heat shock promoter

Conclusions • Not essential for viability • PDZ domain localizes protein to sites of -actinin localization • LIM proteins enable additional protein interactions • many interactions in Y 2 H interactome project • More study needed for specific role

Layton getting CRAZY!!!
- Slides: 13