Molecular Cell Biology Bio 5068 Cell Cycle I

Molecular Cell Biology (Bio 5068) Cell Cycle I Ron Bose, MD Ph. D November 10, 2016
SI S M IT O CELL DIVISION CYCLE M G 2 S G 1


DISCOVERY AND NAMING OF CYCLINS A protein (called “cyclin”) was observed to increase as cells approached mitosis, peak in mitosis and then precipitously disappear as cells exited mitosis.
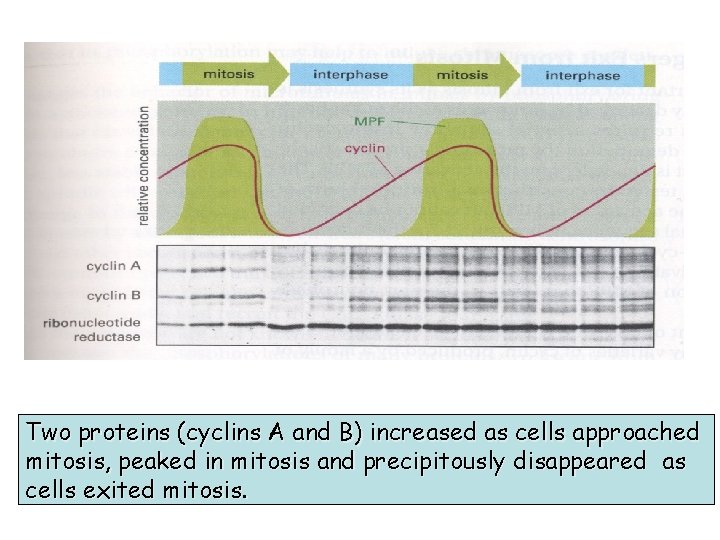
Two proteins (cyclins A and B) increased as cells approached mitosis, peaked in mitosis and precipitously disappeared as cells exited mitosis.
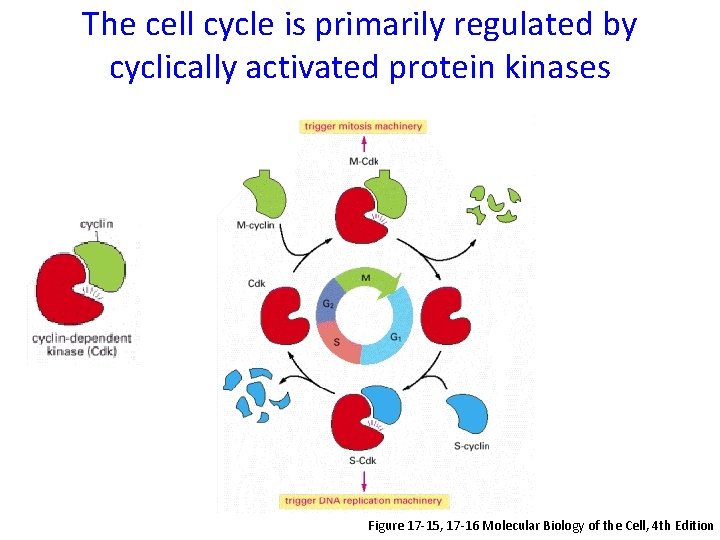
The cell cycle is primarily regulated by cyclically activated protein kinases Figure 17 -15, 17 -16 Molecular Biology of the Cell, 4 th Edition
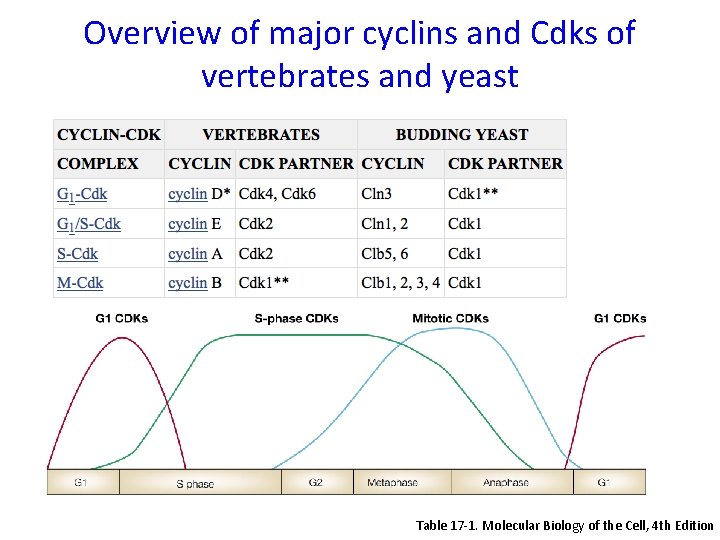
Overview of major cyclins and Cdks of vertebrates and yeast Table 17 -1. Molecular Biology of the Cell, 4 th Edition
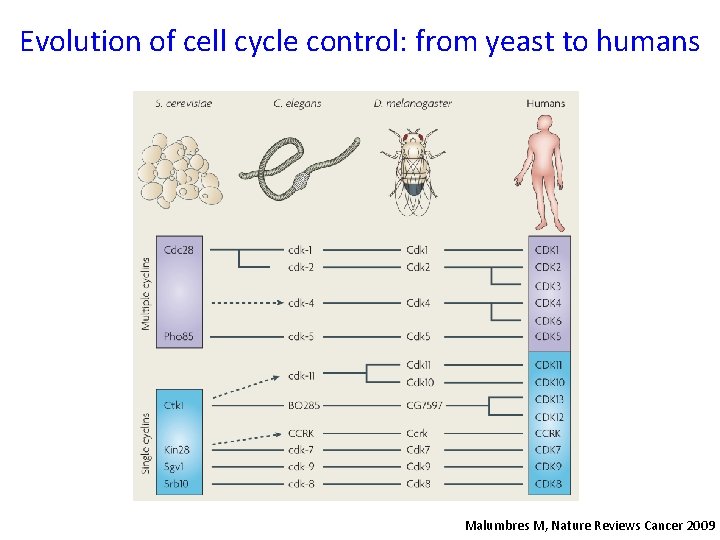
Evolution of cell cycle control: from yeast to humans Malumbres M, Nature Reviews Cancer 2009
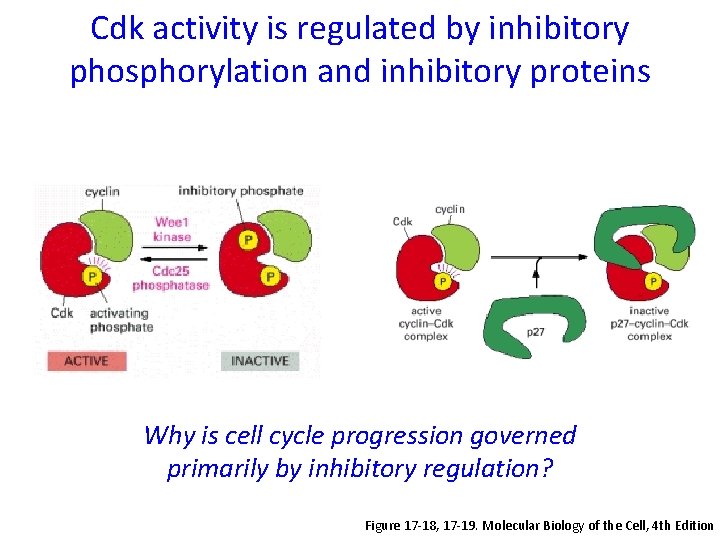
Cdk activity is regulated by inhibitory phosphorylation and inhibitory proteins Why is cell cycle progression governed primarily by inhibitory regulation? Figure 17 -18, 17 -19. Molecular Biology of the Cell, 4 th Edition
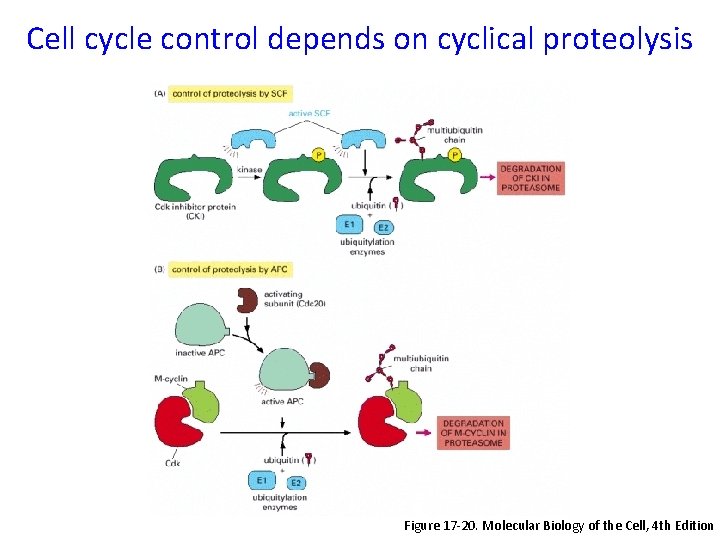
Cell cycle control depends on cyclical proteolysis Figure 17 -20. Molecular Biology of the Cell, 4 th Edition
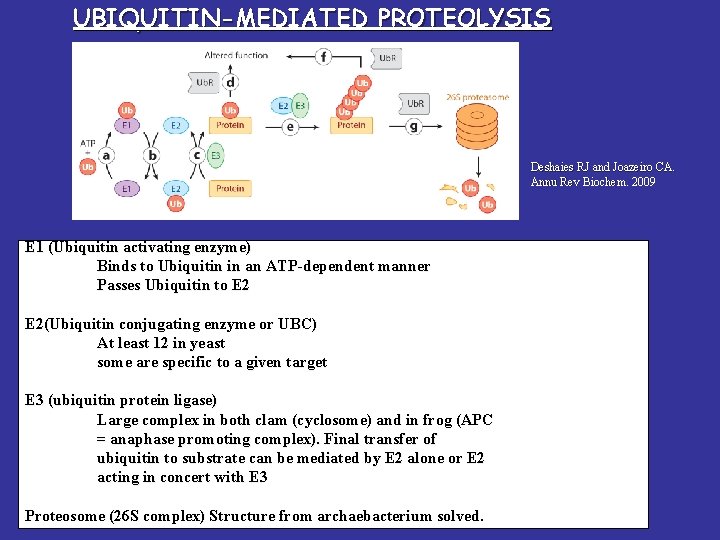
UBIQUITIN-MEDIATED PROTEOLYSIS Deshaies RJ and Joazeiro CA. Annu Rev Biochem. 2009 E 1 (Ubiquitin activating enzyme) Binds to Ubiquitin in an ATP-dependent manner Passes Ubiquitin to E 2(Ubiquitin conjugating enzyme or UBC) At least 12 in yeast some are specific to a given target E 3 (ubiquitin protein ligase) Large complex in both clam (cyclosome) and in. PROTEOLYIS frog (APC UBIQUITIN-MEDIATED = anaphase promoting complex). Final transfer of ubiquitin to substrate can be mediated by E 2 alone or E 2 acting in concert with E 3 Proteosome (26 S complex) Structure from archaebacterium solved.
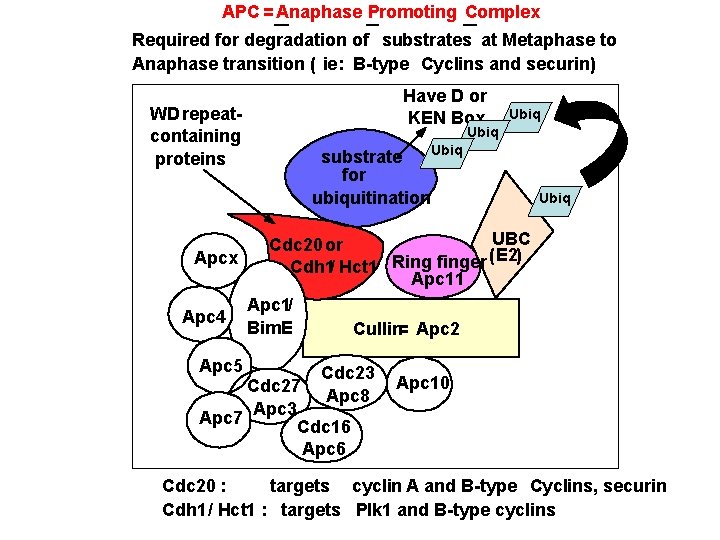
APC = Anaphase Promoting Complex Required for degradation of substrates at Metaphase to Anaphase transition ( ie: B-type Cyclins and securin) WD repeatcontaining proteins Have D or KEN Box Ubiq substrate for ubiquitination Ubiq UBC Cdc 20 or (E 2) Apcx Cdh 1/ Hct 1 Ring finger Apc 11 Apc 1/ Apc 4 Bim. E Cullin= Apc 2 Apc 5 Cdc 23 Apc 8 Cdc 27 Apc 3 Apc 7 Cdc 16 Apc 10 Cdc 20 : targets cyclin A and B-type Cyclins, securin Cdh 1/ Hct 1 : targets Plk 1 and B-type cyclins
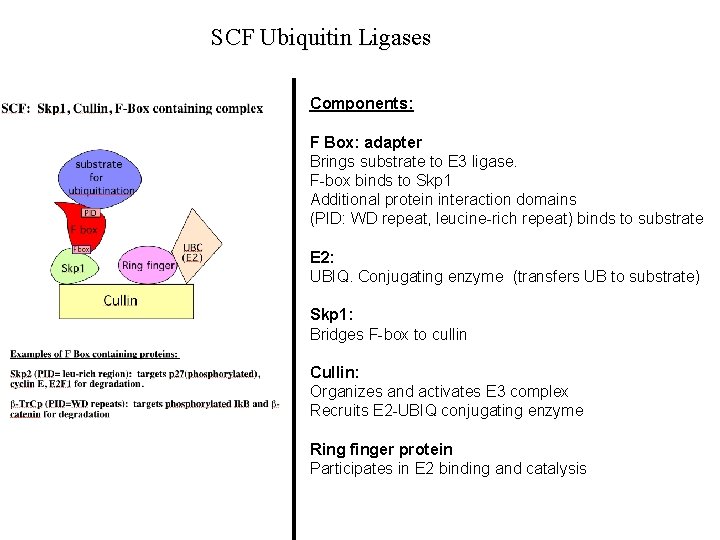
SCF Ubiquitin Ligases Components: F Box: adapter Brings substrate to E 3 ligase. F-box binds to Skp 1 Additional protein interaction domains (PID: WD repeat, leucine-rich repeat) binds to substrate E 2: UBIQ. Conjugating enzyme (transfers UB to substrate) Skp 1: Bridges F-box to cullin Cullin: Organizes and activates E 3 complex Recruits E 2 -UBIQ conjugating enzyme Ring finger protein Participates in E 2 binding and catalysis
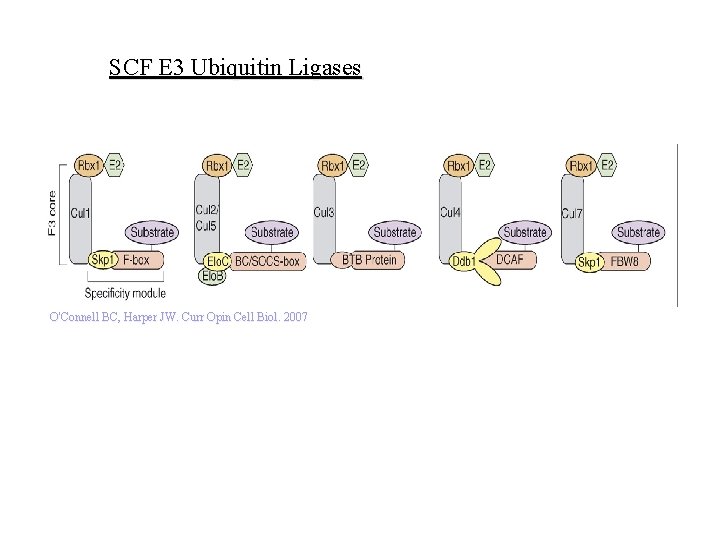
SCF E 3 Ubiquitin Ligases O'Connell BC, Harper JW. Curr Opin Cell Biol. 2007
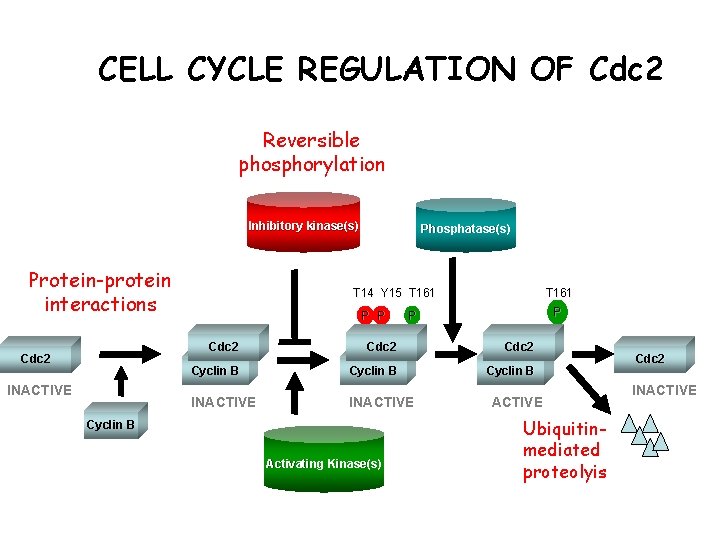
CELL CYCLE REGULATION OF Cdc 2 Reversible phosphorylation Inhibitory kinase(s) Protein-protein interactions Cdc 2 INACTIVE Phosphatase(s) T 14 Y 15 T 161 P P Cdc 2 Cyclin B INACTIVE Cyclin B Activating Kinase(s) ACTIVE Ubiquitinmediated proteolyis Cdc 2 INACTIVE

Cyclin-dependent Kinase Inhibitor Proteins (CKI’s) 1. CIP/KIP family (p 21 Cip 1, p 27 Kip 1, p 57 Kip 2): a. Binds to Cdk 2 and inhibits activity. b. Binds Cdk 4/6 and helps assemble complexes with cyclins. 2. INK 4 family (p 16, p 15, p 18, p 19). a. Specific for Cdk 4 and Cdk 6. b. Binds Cdk subunit alone and prevents cyclin binding c. Bind and inhibit Cdk 4/6 -Cyclin D heterodimers.
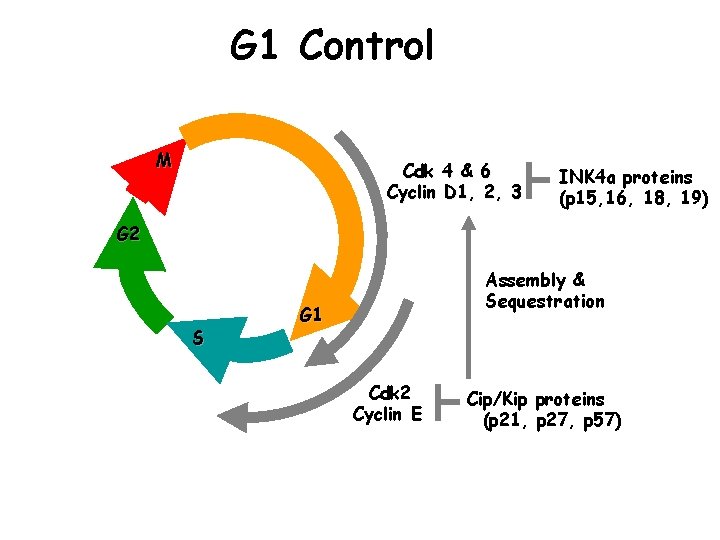
G 1 Control M Cdk 4 & 6 Cyclin D 1, 2, 3 INK 4 a proteins (p 15, 16, 18, 19) G 2 S Assembly & Sequestration G 1 Cdk 2 Cyclin E Cip/Kip proteins (p 21, p 27, p 57)
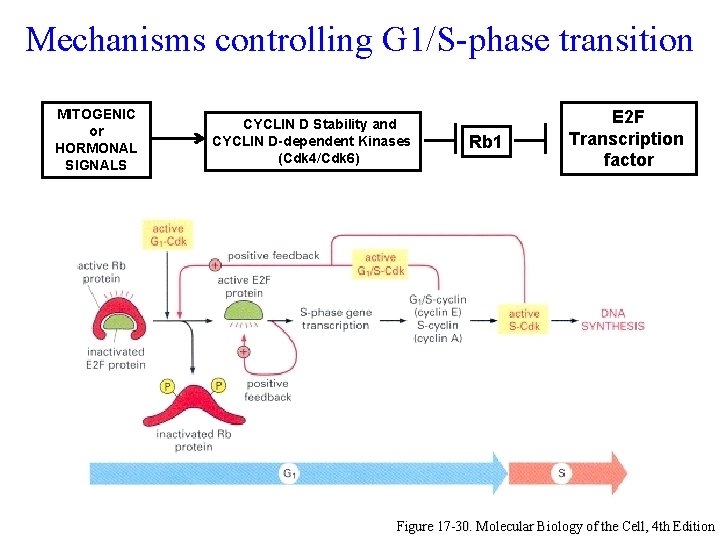
Mechanisms controlling G 1/S-phase transition MITOGENIC or HORMONAL SIGNALS CYCLIN D Stability and CYCLIN D-dependent Kinases (Cdk 4/Cdk 6) Rb 1 E 2 F Transcription factor Figure 17 -30. Molecular Biology of the Cell, 4 th Edition
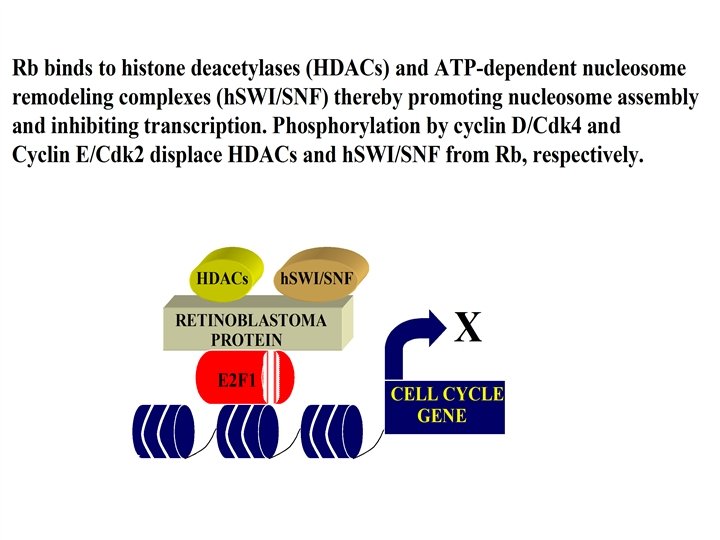
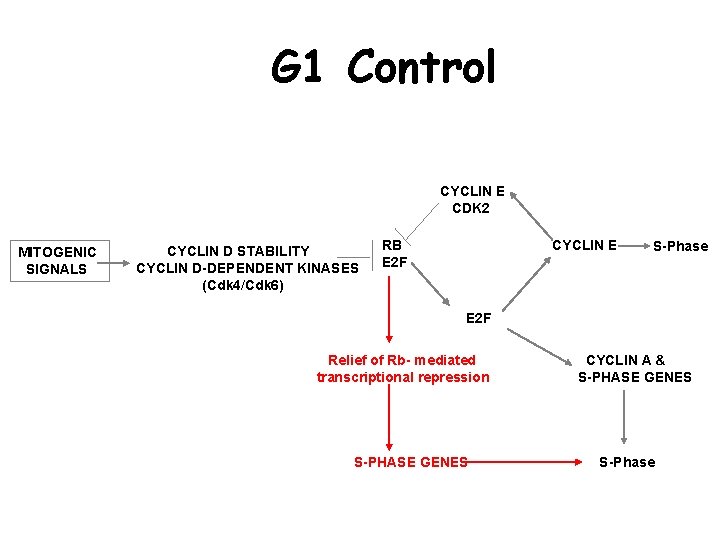
G 1 Control CYCLIN E CDK 2 MITOGENIC SIGNALS CYCLIN D STABILITY CYCLIN D-DEPENDENT KINASES (Cdk 4/Cdk 6) RB E 2 F CYCLIN E S-Phase E 2 F Relief of Rb- mediated transcriptional repression S-PHASE GENES CYCLIN A & S-PHASE GENES S-Phase
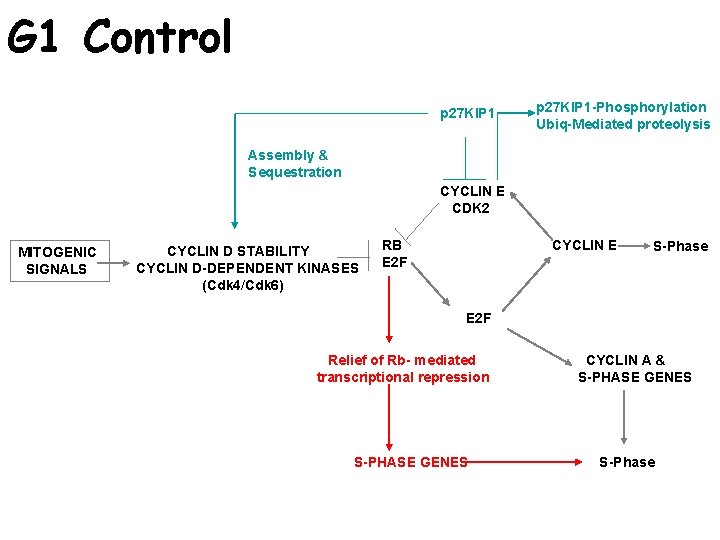
G 1 Control p 27 KIP 1 -Phosphorylation Ubiq-Mediated proteolysis Assembly & Sequestration CYCLIN E CDK 2 MITOGENIC SIGNALS CYCLIN D STABILITY CYCLIN D-DEPENDENT KINASES (Cdk 4/Cdk 6) RB E 2 F CYCLIN E S-Phase E 2 F Relief of Rb- mediated transcriptional repression S-PHASE GENES CYCLIN A & S-PHASE GENES S-Phase

Checkpoints What are they? How were they defined? How does their derailment contribute to cancer?
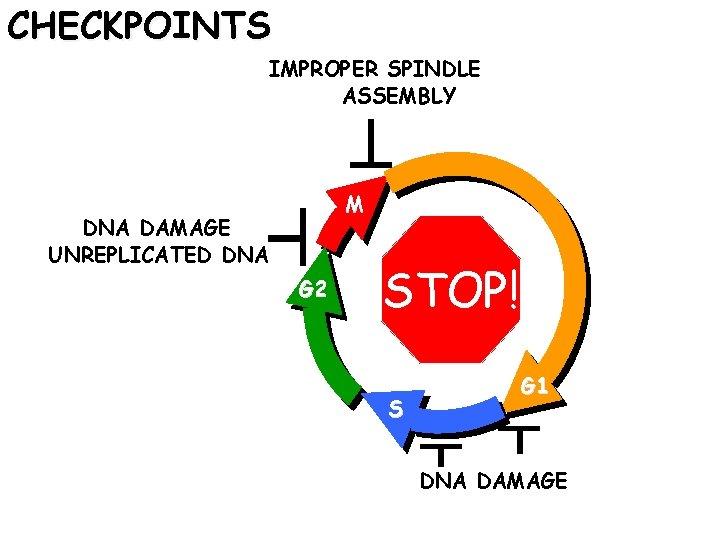
CHECKPOINTS IMPROPER SPINDLE ASSEMBLY M DNA DAMAGE UNREPLICATED DNA G 2 STOP! S G 1 DNA DAMAGE
Checkpoints: intracellular signaling pathways that determine if previous steps are complete before proceeding onto the next stage (complete DNA synthesis before entering mitosis; spindles must be assembled before exiting metaphase and entering into anaphase) and whethere has been any damage to the DNA damage checkpoint: integrity of DNA damage is repaired before entering S, completing S or entering M. DNA replication checkpoint: replication state of DNA Complete DNA synthesis before mitosis. Spindle assembly checkpoint: integrity of spindles must be assembled before exiting metaphase into anaphase.
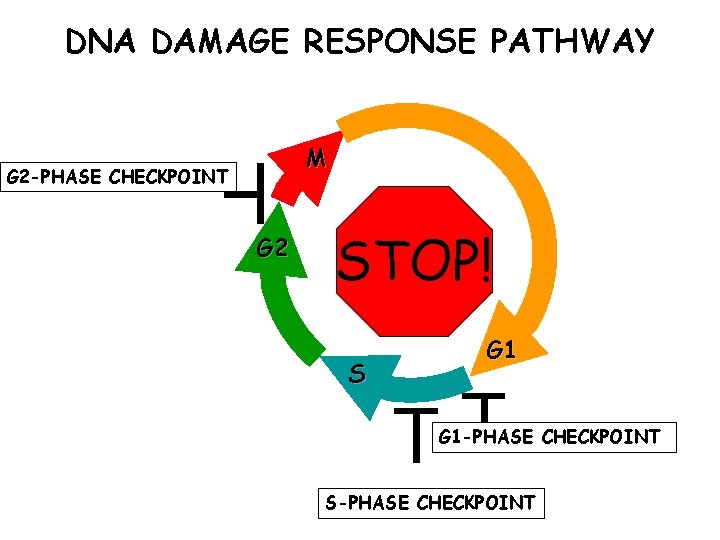
DNA DAMAGE RESPONSE PATHWAY M G 2 -PHASE CHECKPOINT G 2 STOP! S G 1 -PHASE CHECKPOINT S-PHASE CHECKPOINT
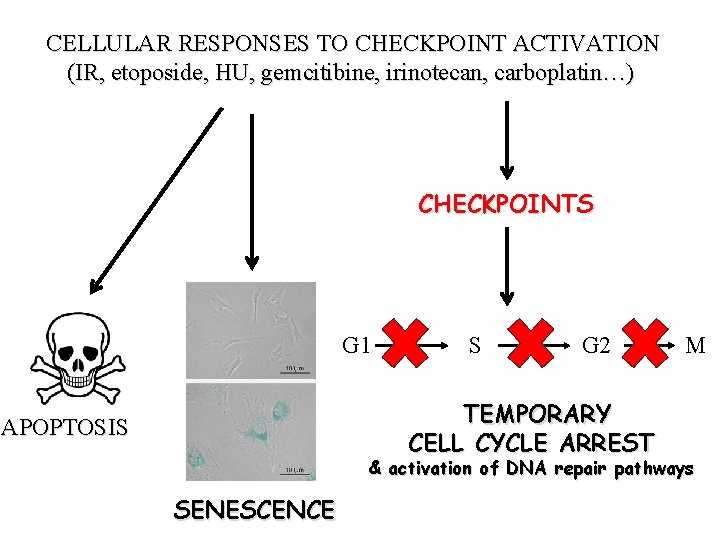
CELLULAR RESPONSES TO CHECKPOINT ACTIVATION (IR, etoposide, HU, gemcitibine, irinotecan, carboplatin…) CHECKPOINTS G 1 S G 2 TEMPORARY CELL CYCLE ARREST APOPTOSIS M & activation of DNA repair pathways SENESCENCE
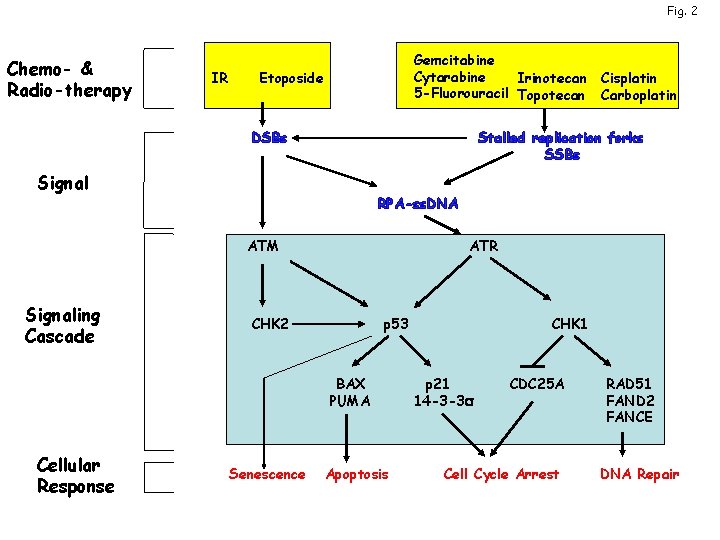
Fig. 2 Chemo- & Radio-therapy IR Gemcitabine Cytarabine Irinotecan 5 -Fluorouracil Topotecan Etoposide DSBs Stalled replication forks SSBs Signal RPA-ss. DNA ATM Signaling Cascade ATR CHK 2 p 53 BAX PUMA Cellular Response Cisplatin Carboplatin Senescence Apoptosis CHK 1 p 21 14 -3 -3 s CDC 25 A Cell Cycle Arrest RAD 51 FAND 2 FANCE DNA Repair
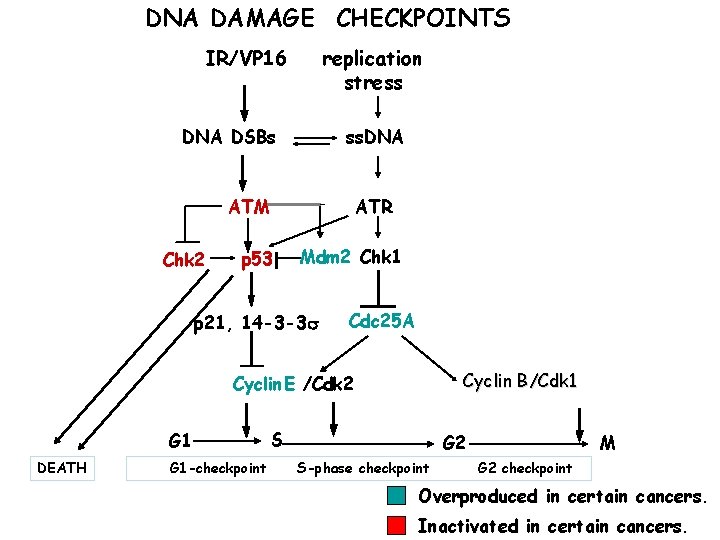
DNA DAMAGE CHECKPOINTS IR/VP 16 replication stress DNA DSBs ss. DNA ATR ATM Chk 2 p 53 Mdm 2 Chk 1 p 21, 14 -3 -3 s Cdc 25 A Cyclin B/Cdk 1 Cyclin E /Cdk 2 G 1 DEATH G 1 -checkpoint S G 2 S-phase checkpoint M G 2 checkpoint Overproduced in certain cancers. Inactivated in certain cancers.
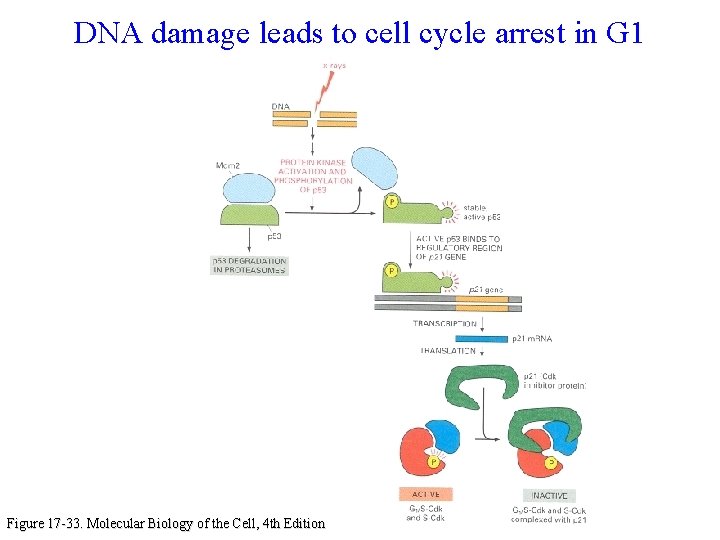
DNA damage leads to cell cycle arrest in G 1 Figure 17 -33. Molecular Biology of the Cell, 4 th Edition
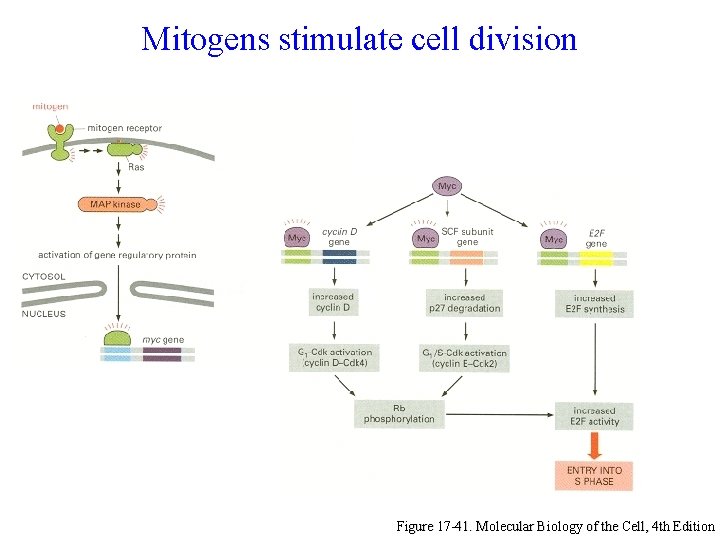
Mitogens stimulate cell division Figure 17 -41. Molecular Biology of the Cell, 4 th Edition
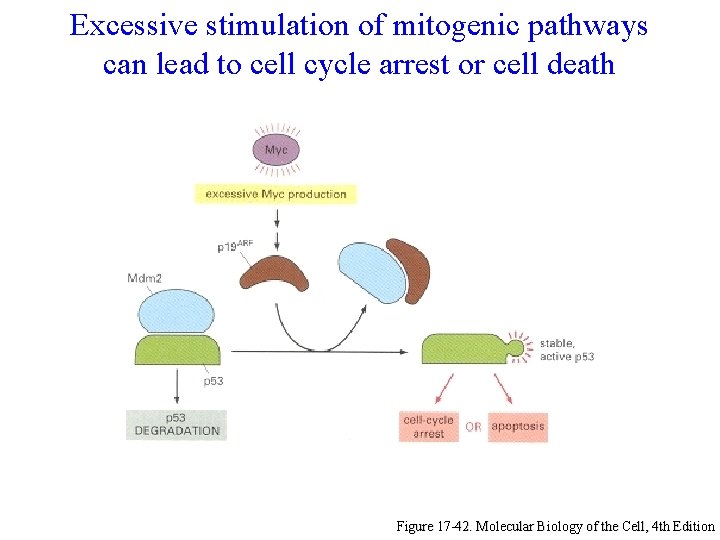
Excessive stimulation of mitogenic pathways can lead to cell cycle arrest or cell death Figure 17 -42. Molecular Biology of the Cell, 4 th Edition
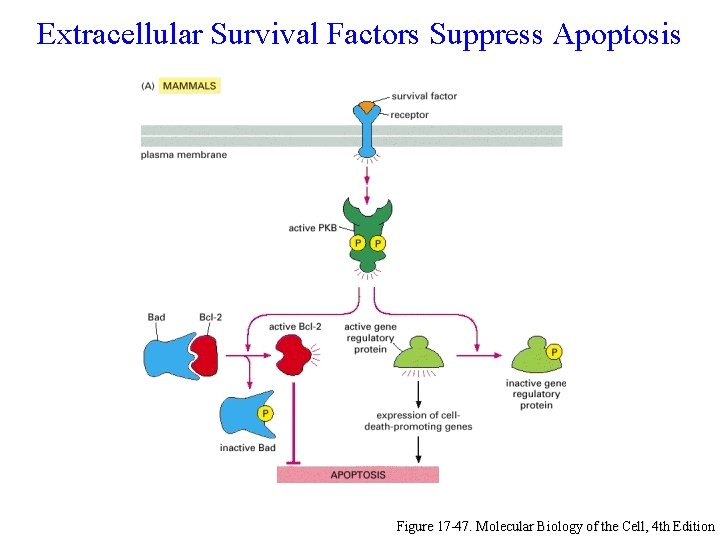
Extracellular Survival Factors Suppress Apoptosis Figure 17 -47. Molecular Biology of the Cell, 4 th Edition
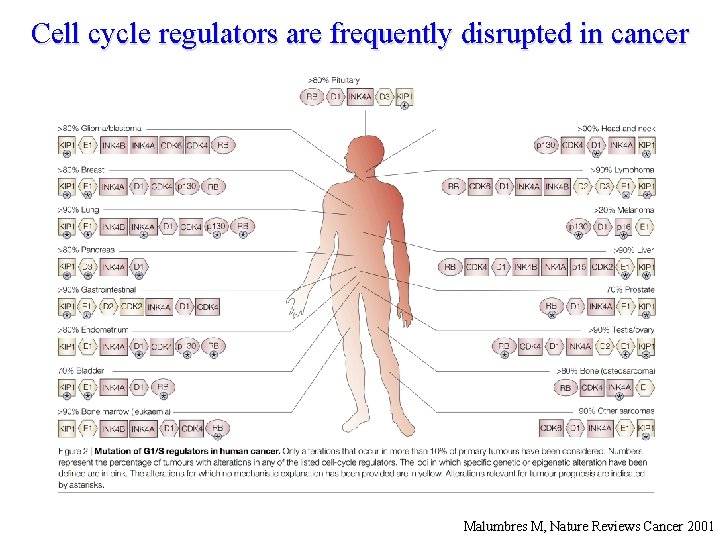
Cell cycle regulators are frequently disrupted in cancer Malumbres M, Nature Reviews Cancer 2001
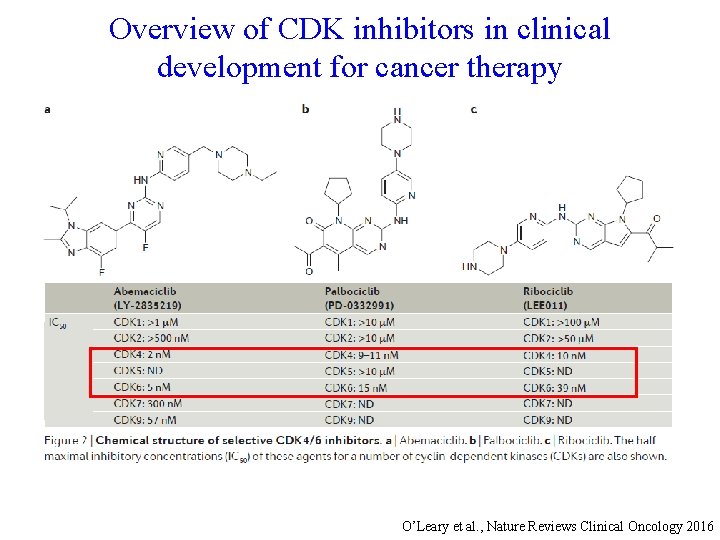
Overview of CDK inhibitors in clinical development for cancer therapy O’Leary et al. , Nature Reviews Clinical Oncology 2016
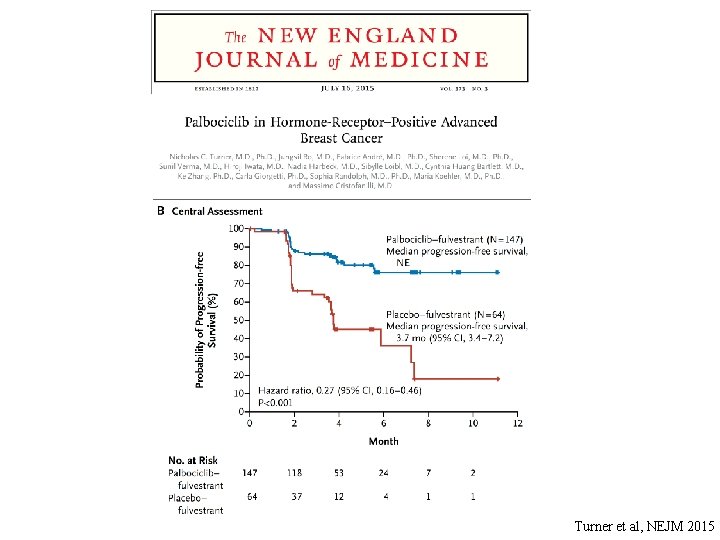
Turner et al, NEJM 2015
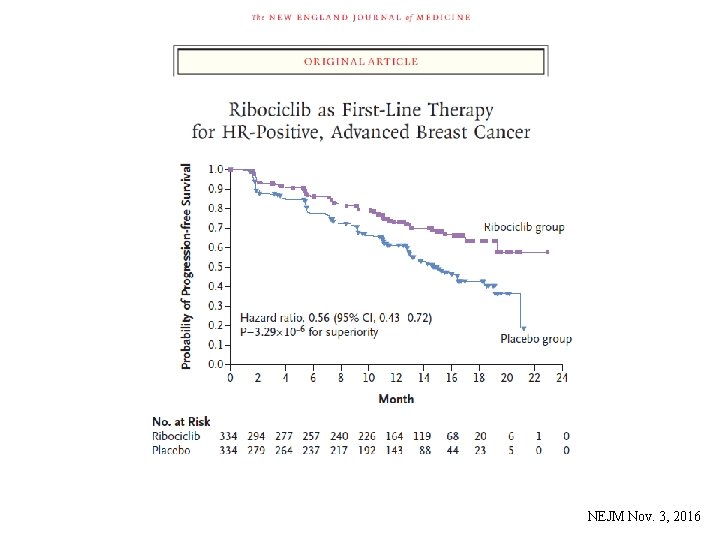
NEJM Nov. 3, 2016

Conclusions 1. The cell cycle is a coordinated and tightly organized process to ensure the successful replication of the cell. 2. Activity of CDK-Cyclins is determined by: 1. Synthesis of Cyclins. 2. Reversible phosphorylation/dephosphorylation of stimulatory and inhibitory sites on CDK. 3. Ubiquitin mediated degradation of Cyclins. 4. CDK inhibitors – INK 4 and CIP/KIP families. 3. Checkpoints can halt the cell cycle if all steps have not been properly completed. 4. Cancers have many alterations in cell cycle proteins and selective CDK 4/6 inhibitors are now used in cancer treatment.
- Slides: 37