Molecular Biology TRANSCRIPTION 1 v Definition v When

Molecular Biology TRANSCRIPTION 1
v. Definition v. When it occurs in the cell v. Types of RNA & Differences with DNA v. Requirements for Transcription TRANSCRIPTION v. Stages of Transcription v. Events of Transcription v. Post transcriptional modfications v. Inhibitors of transcription 2
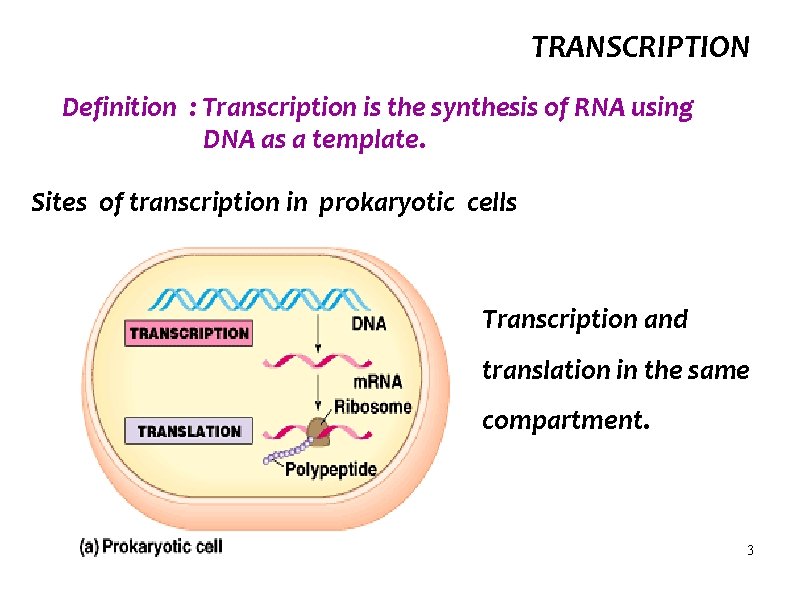
TRANSCRIPTION Definition : Transcription is the synthesis of RNA using DNA as a template. Sites of transcription in prokaryotic cells Transcription and translation in the same compartment. 3
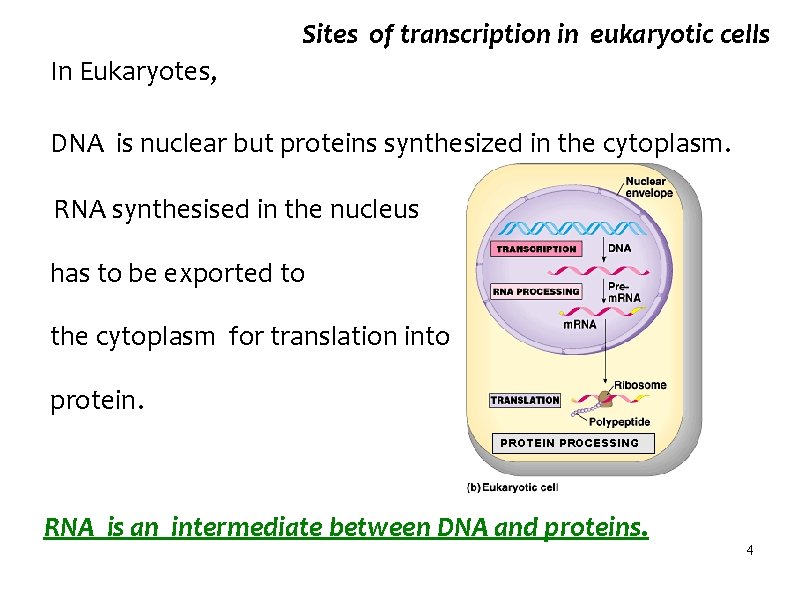
Sites of transcription in eukaryotic cells In Eukaryotes, DNA is nuclear but proteins synthesized in the cytoplasm. RNA synthesised in the nucleus has to be exported to the cytoplasm for translation into protein. PROTEIN PROCESSING RNA is an intermediate between DNA and proteins. 4

Types of RNA m. RNA Messenger RNA carries information on how to construct a protein. r. RNA Ribosomal RNA is not a code carrier but a structural part of ribosomes. t. RNA Transfer RNA has a coding section and an amino acid carrying section. 5

Types of RNA. . . . hn. RNA Heterogenous nuclear RNA – high MW – nascent RNA synthesised from DNA. Micro. RNA Mi. RNA - a short, non-coding RNA - regulate gene expression - bind to m. RNA - inhibit protein translation. sn. RNA Small nuclear RNA. si. RNA Small interfering RNA 6
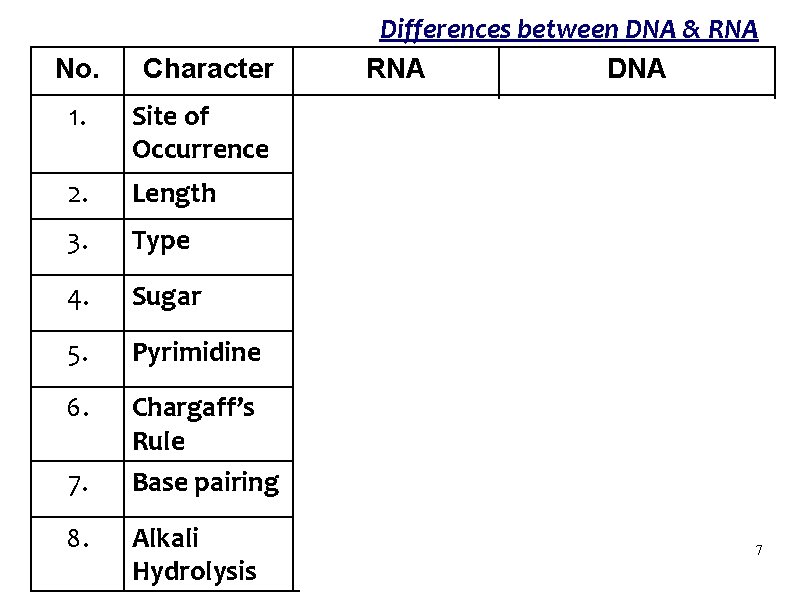
Differences between DNA & RNA DNA No. Character 1. Site of Occurrence Cytoplasm and Nucleus 2. Length 100 – 500 bases Millions of base pairs 3. Type Single stranded Double stranded 4. Sugar Ribose Deoxyribose 5. Pyrimidine Uracil Thymine 6. Chargaff’s Rule Base pairing Does not Follow Intrastrand Follows Interstrand Easily destroyed Alkali resistant 7. 8. Alkali Hydrolysis Nucleus 7

Salient features of Transcription v. Only the gene-segments of the DNA molecule are transcribed. (Gene : Sequence in the DNA coding for a protein) v. The region on the strand of DNA that is transcribed acts as template and is called template strand. v. Complimentary region of the template on the other strand of DNA is called coding strand. Reason : The base sequence of this strand is identical to that of m. RNA, except U replacing T. 8

v Direction of reading DNA template - 3’ 5’ synthesis of RNA - 5’ 3’ v. There is no proof reading and editing mechanism in transcription - is less serious than replication. Reason : any error in reading the bases will produce a) only a few abnormal protein molecules without affecting the protein function b) error will not be transferred to offspring. v. Transcription does not involve an RNA primer to initiate RNA synthesis. 9

Strands of DNA and its relationship with RNA m. RNA The sequence of the m. RNA is : the same as that on the anti template strand except the thymine residues ar replaced by uracil. 10

Each RNA is complementary to one strand of the DNA double helix - Template strand. The opposite strand is called Non-template Anti-template coding strand m. RNA-like strand Sense strand : Complementary to the template. the strand whose base sequence is m. RNA-like. 11

Requirements for Transcription: 1. DNA template : One strand of the DNA is the “Template strand” that is copied into m. RNA as per the base pair rule 2. Topoisomerases I and II: Unwind & Rewind the DNA Topoisomerase I Unwind ahead of transcription and Topoisomerase II rewind behind the transcription. 12

Requirements for Transcription. . . 3. Substrates : Ribonucleotide triphosphates – ATP, GTP, CTP, UTP 4. Enzymes : a) RNA Polymerase - RNAP Types of RNAP – 3 types (Prokaryotes) RNAP I - synthesizes RNAP III - r. RNA m. RNA t. RNA 13
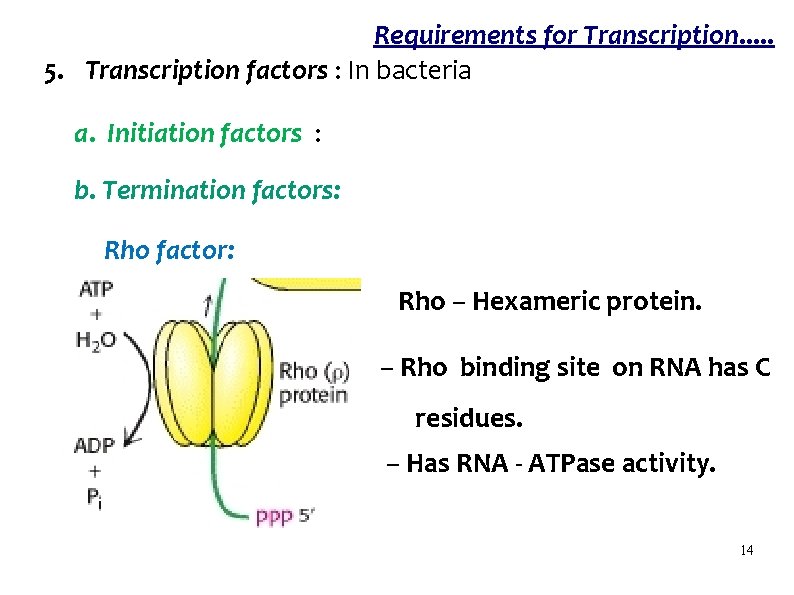
Requirements for Transcription. . . 5. Transcription factors : In bacteria a. Initiation factors : b. Termination factors: Rho factor: Rho – Hexameric protein. – Rho binding site on RNA has C residues. – Has RNA - ATPase activity. 14
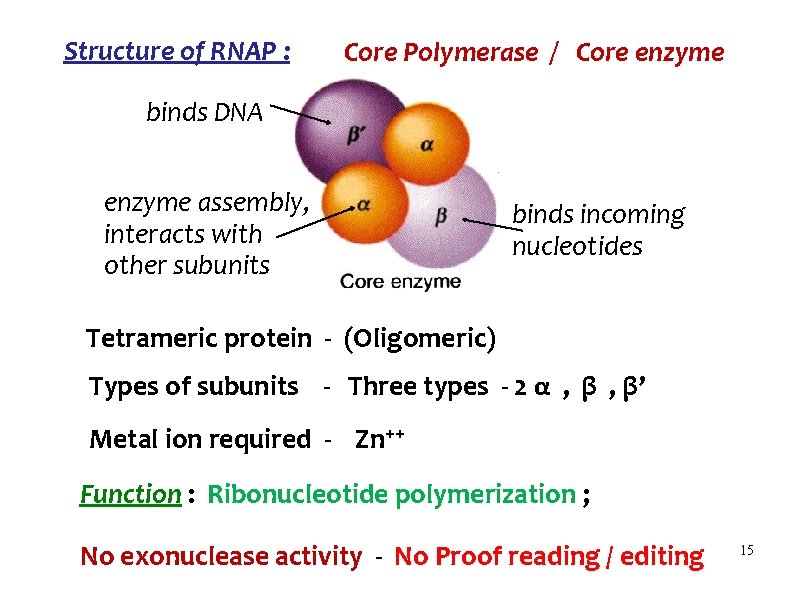
Structure of RNAP : Core Polymerase / Core enzyme binds DNA enzyme assembly, interacts with other subunits binds incoming nucleotides Tetrameric protein - (Oligomeric) Types of subunits - Three types - 2 α , β’ Metal ion required - Zn++ Function : Ribonucleotide polymerization ; No exonuclease activity - No Proof reading / editing 15
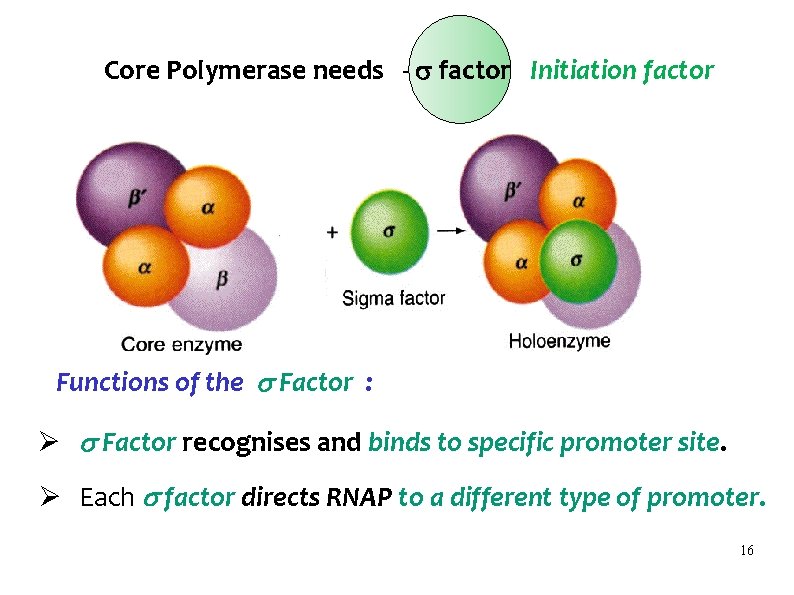
Core Polymerase needs - factor Initiation factor Functions of the Factor : Ø Factor recognises and binds to specific promoter site. Ø Each factor directs RNAP to a different type of promoter. 16

Transcription Regulatory Sites on the DNA In addition to transcription site (gene), DNA contains specific base sequences, “transcription regulatory sites” Function : involved in regulation of transcription. Examples : ü Promoter site – for initiation of transcription üTermination site – for termination of transcription ü Enhancer üSilencer and – for modifying the rate of transcription üHormone responsive elements (HRE) 17
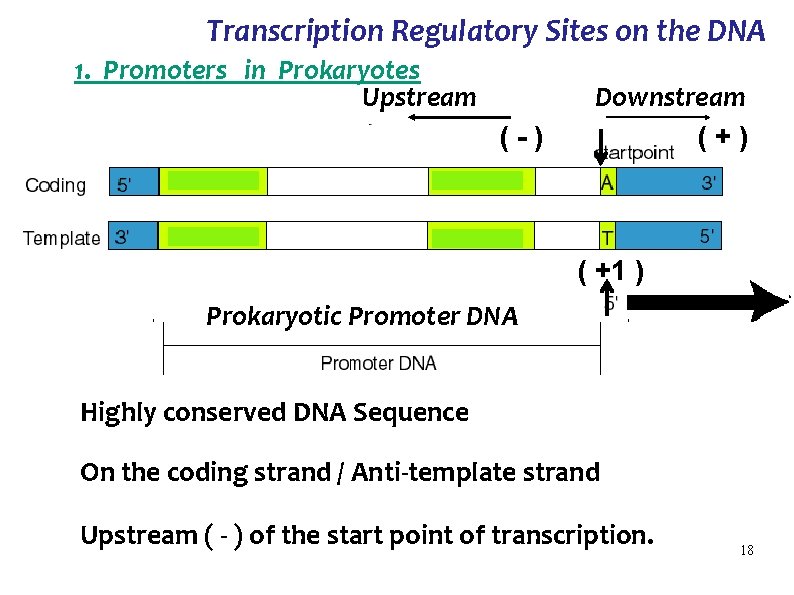
Transcription Regulatory Sites on the DNA 1. Promoters in Prokaryotes Upstream (-) Downstream (+) ( +1 ) Prokaryotic Promoter DNA Highly conserved DNA Sequence On the coding strand / Anti-template strand Upstream ( - ) of the start point of transcription. 18
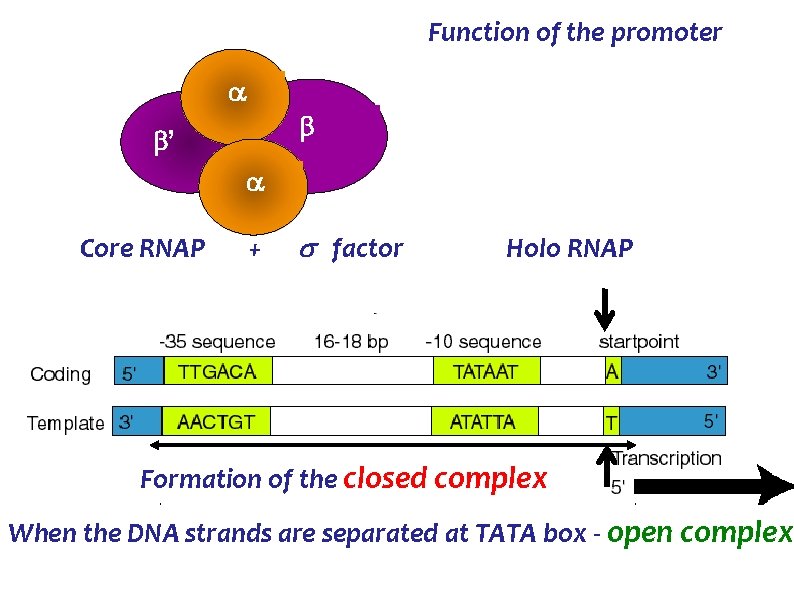
Function of the promoter ’ Core RNAP + factor Holo RNAP Formation of the closed complex When the DNA strands are separated at TATA box - open complex

In Prokaryotes, Promoters are : Sequence TATAAT Name Location - TATA box TTGACA box - 10 bases -35 bases In Eukaryotes, Promoters are : Sequence Name Location TATAAA - Goldberg Hogness box- -25 to -30 bases GGCCAATCT - CAAT box -70 to -80 bases
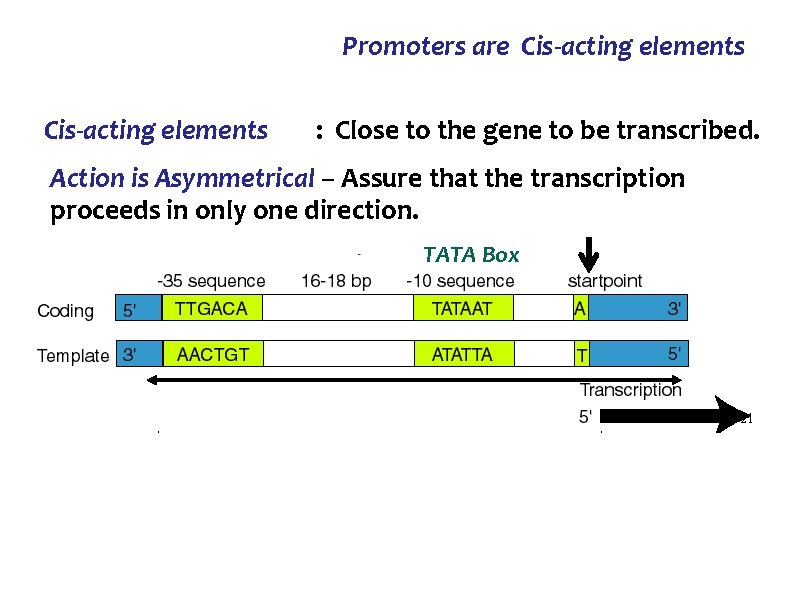
Promoters are Cis-acting elements : Close to the gene to be transcribed. Action is Asymmetrical – Assure that the transcription proceeds in only one direction. TATA Box 21
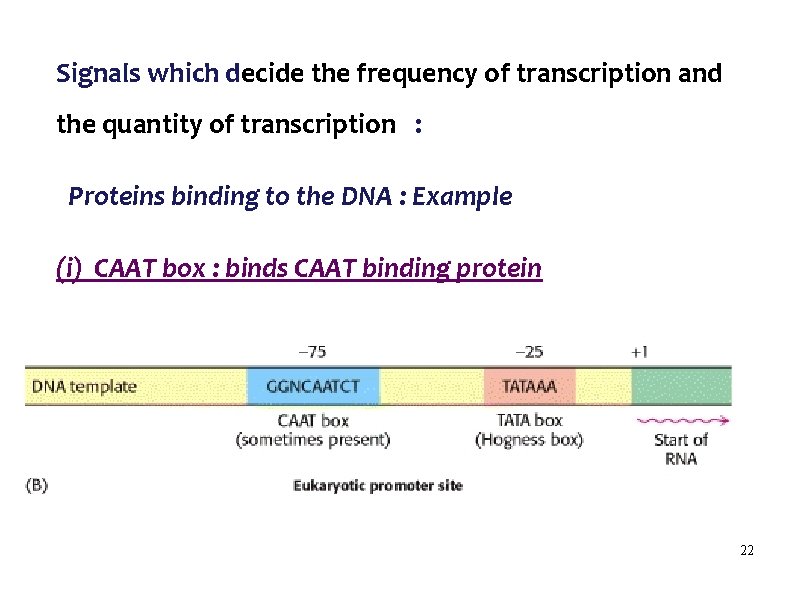
Signals which decide the frequency of transcription and the quantity of transcription : Proteins binding to the DNA : Example (i) CAAT box : binds CAAT binding protein 22
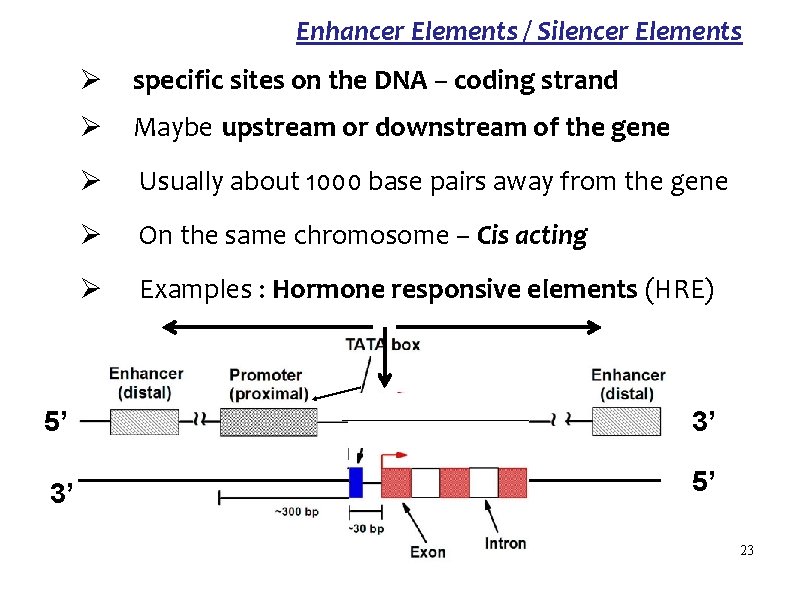
Enhancer Elements / Silencer Elements Ø specific sites on the DNA – coding strand Ø Maybe upstream or downstream of the gene Ø Usually about 1000 base pairs away from the gene Ø On the same chromosome – Cis acting Ø Examples : Hormone responsive elements (HRE) 5’ 3’ 3’ 5’ 23
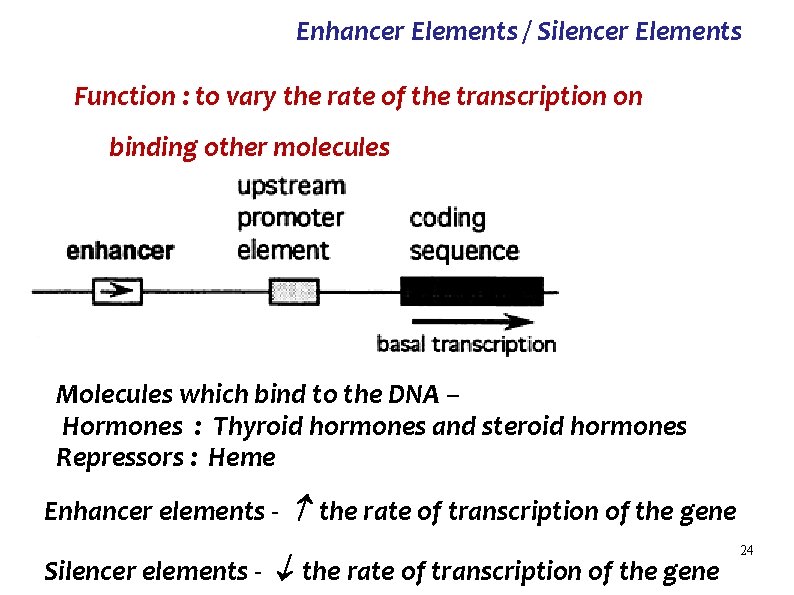
Enhancer Elements / Silencer Elements Function : to vary the rate of the transcription on binding other molecules Molecules which bind to the DNA – Hormones : Thyroid hormones and steroid hormones Repressors : Heme Enhancer elements - the rate of transcription of the gene Silencer elements - the rate of transcription of the gene 24
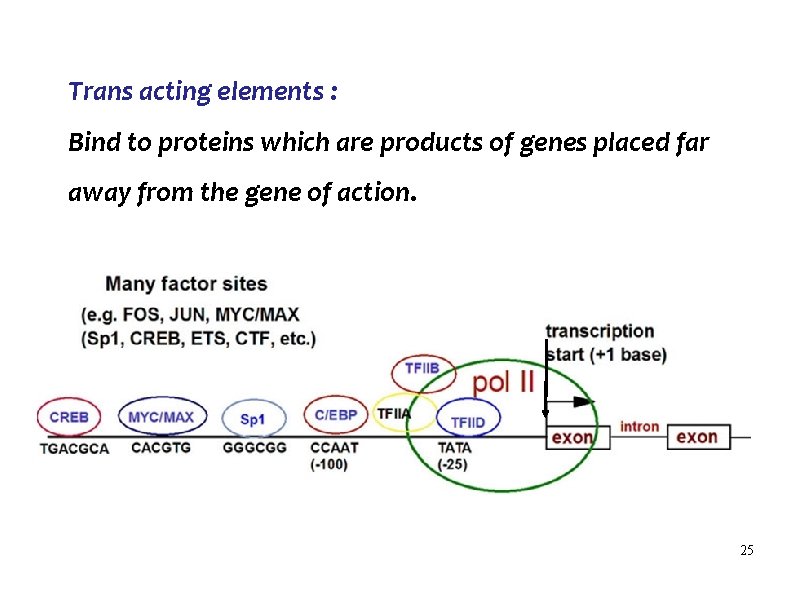
Trans acting elements : Bind to proteins which are products of genes placed far away from the gene of action. 25
Stages of transcription Transcription has 4 stages I stage : Initiation of transcription II stage : Elongation III stage : Termination IV stage : Post-transcriptional modifications 26
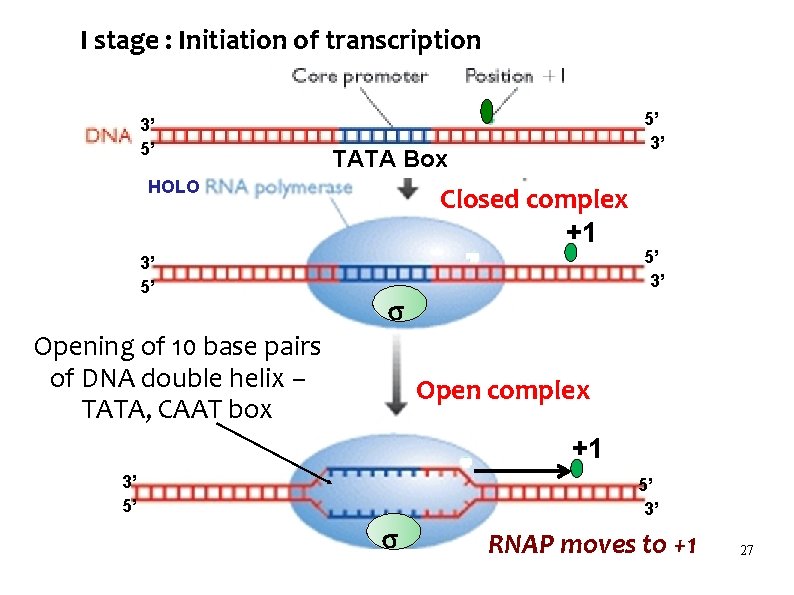
I stage : Initiation of transcription 3’ 5’ TATA Box HOLO 3’ 5’ 5’ 3’ Closed complex +1 5’ 3’ Opening of 10 base pairs of DNA double helix – TATA, CAAT box Open complex +1 3’ 5’ 5’ 3’ RNAP moves to +1 27
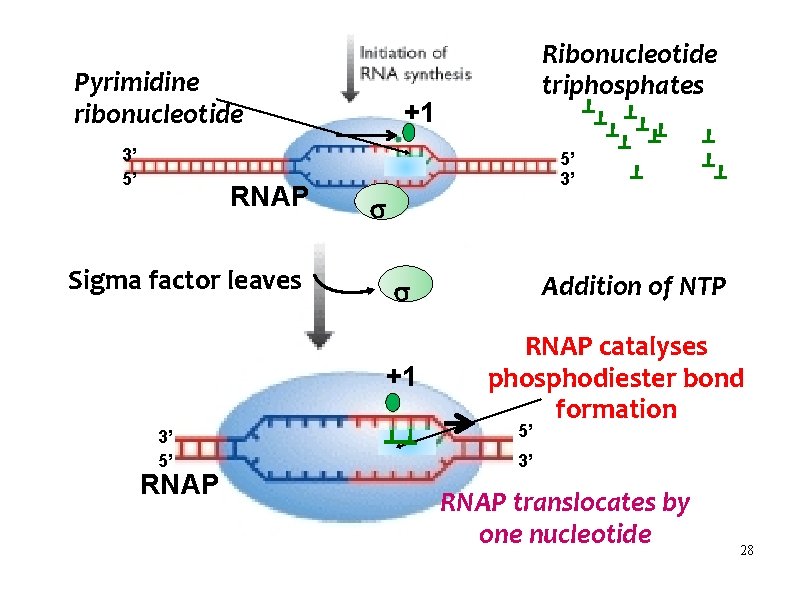
Pyrimidine ribonucleotide 3’ 5’ RNAP Sigma factor leaves +1 5’ 3’ RNAP Addition of NTP +1 3’ 5’ Ribonucleotide triphosphates RNAP catalyses phosphodiester bond formation 5’ 3’ RNAP translocates by one nucleotide 28
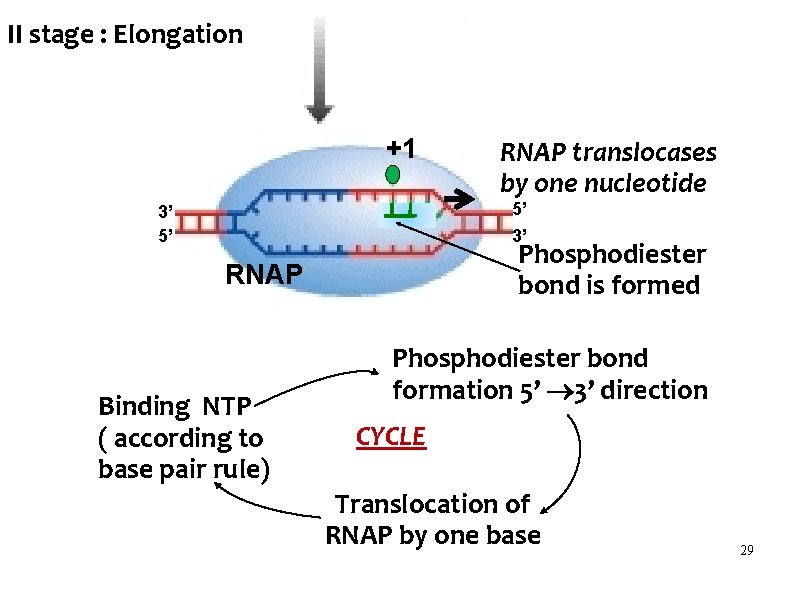
II stage : Elongation +1 RNAP translocases by one nucleotide 5’ 3’ 3’ 5’ Phosphodiester bond is formed RNAP Binding NTP ( according to base pair rule) Phosphodiester bond formation 5’ 3’ direction CYCLE Translocation of RNAP by one base 29

II stage : Elongation Cycle of : Binding NTP – Phosphodiester bond formation – translocation occurs Rate : 40 nucleotides / second RNAP moves downstream - 3’ 5’ direction on the DNA template Nucleotides are added according to base pair rule DNA template - RNA Hybrid is formed in 5’ 3’ direction Transcription bubble is formed 30
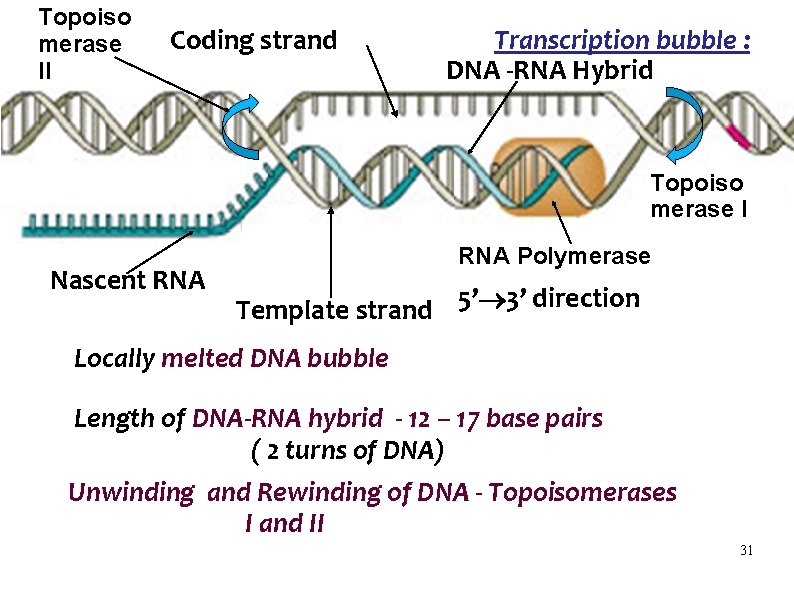
Topoiso merase II Coding strand Transcription bubble : DNA -RNA Hybrid Topoiso merase I Nascent RNA Polymerase Template strand 5’ 3’ direction Locally melted DNA bubble Length of DNA-RNA hybrid - 12 – 17 base pairs ( 2 turns of DNA) Unwinding and Rewinding of DNA - Topoisomerases I and II 31
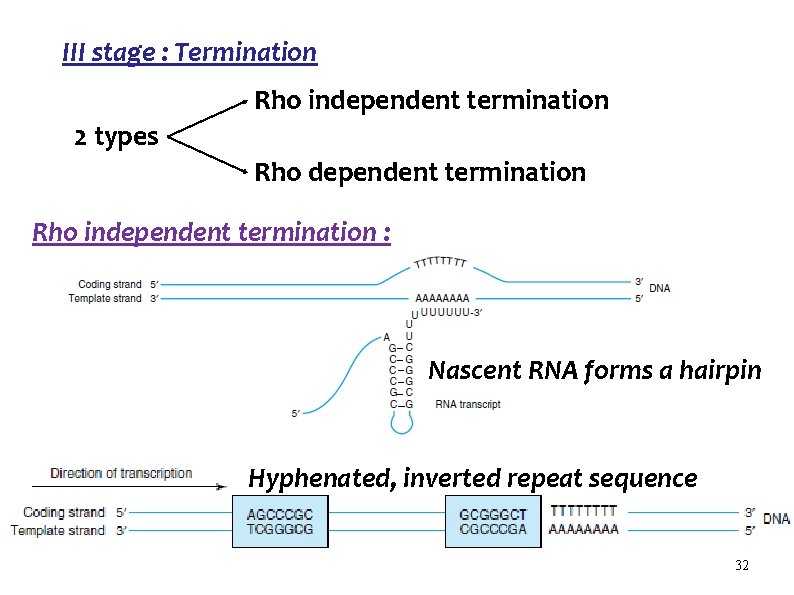
III stage : Termination Rho independent termination 2 types Rho dependent termination Rho independent termination : Nascent RNA forms a hairpin Hyphenated, inverted repeat sequence 32
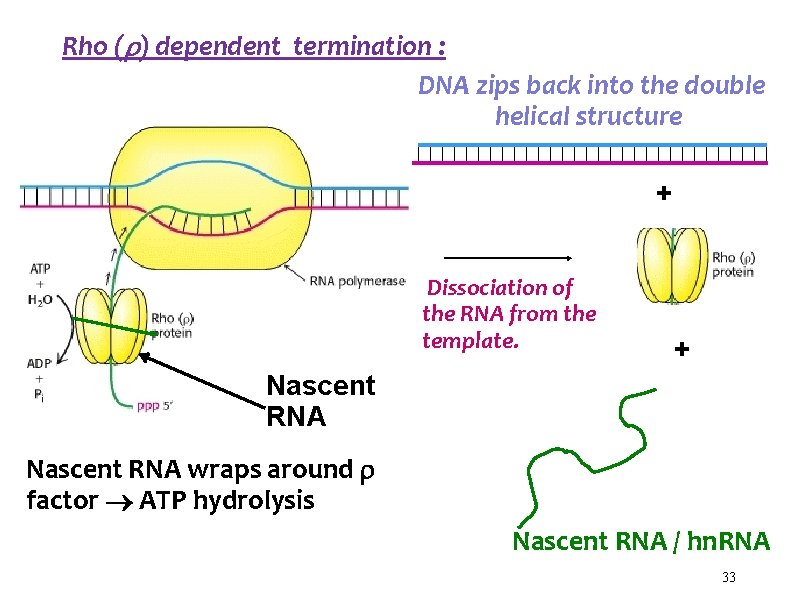
Rho ( ) dependent termination : DNA zips back into the double helical structure + Dissociation of the RNA from the template. + Nascent RNA wraps around factor ATP hydrolysis Nascent RNA / hn. RNA 33
Nascent RNA : is newly synthesized DNA - Sequence from Transcription start site to the termination site -Also called as Precusor RNA - Primary RNA - Heterogenous RNA (hn. RNA; pre RNA) Post Transcriptional Modifications – In the nucleus editing hn. RNA Mature m. RNA 34
Why does it need processing 1. hn. RNA has Exons : Coding sequences for protein Introns : Intervening sequences (non coding) 2. Removal of introns and joining of exons to form mature RNA ready for translation --- splicing. 3. Protection from nucleases 3’ end by Tailing with a Poly A tail 4. Attaching to ribosome : 5’ end by Capping 7 methyl GTP 5. Modification of certain bases 35

Post Transcriptional Modifications 1. Capping at 5’ end - 7 methyl GTP 2. Splicing : Excision of introns and joining of exons 3. Removal of extra RNA from the 3’ end 4. Tailing at the 3’ end – Poly A tail 5. Modification of certain bases 36
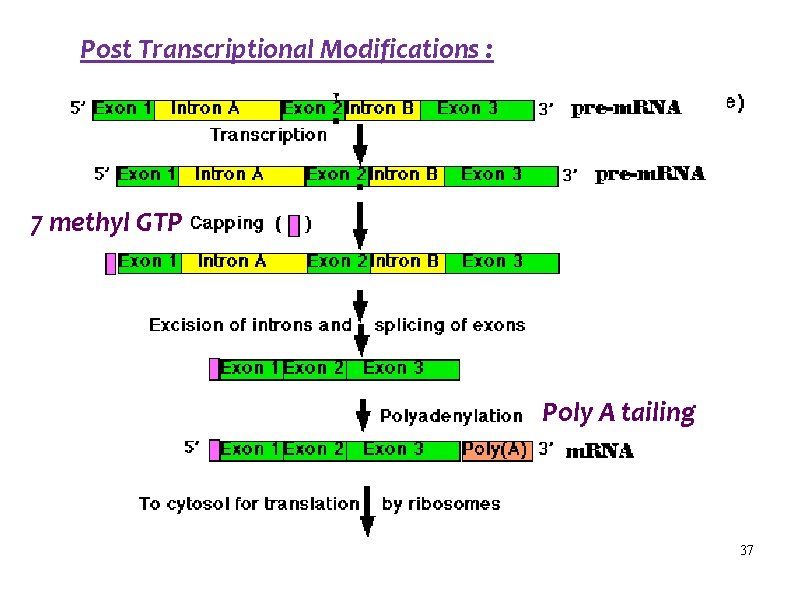
Post Transcriptional Modifications : 7 methyl GTP Poly A tailing 37

Splicing : Exons : Coding sequence Introns : intervening sequence Introns are excised (Deleted) and exons are spliced (joined). Splicing : by spliceosomes = small nuclear RNA + Small nuclear ribonucleoprotein (sn. RNA) (snurps) Systemic Lupus Erythematosus (SLE) Autoimmune disease – antibodies against snurps 38

Inhibitors of Transcription : Actinomycin D : Binds to DNA template and blocks the movement of the RNAP. Α – Amanitin : Poison from the mushrron Amanita phylloids. Strongly inhibits RNAP II Rifampicin : Antibacterial drug. Inhibits initiation of transcription – binds to the β subunit Daunomycin : Anticancer drug. Intercalates in the double stranded DNA. 39
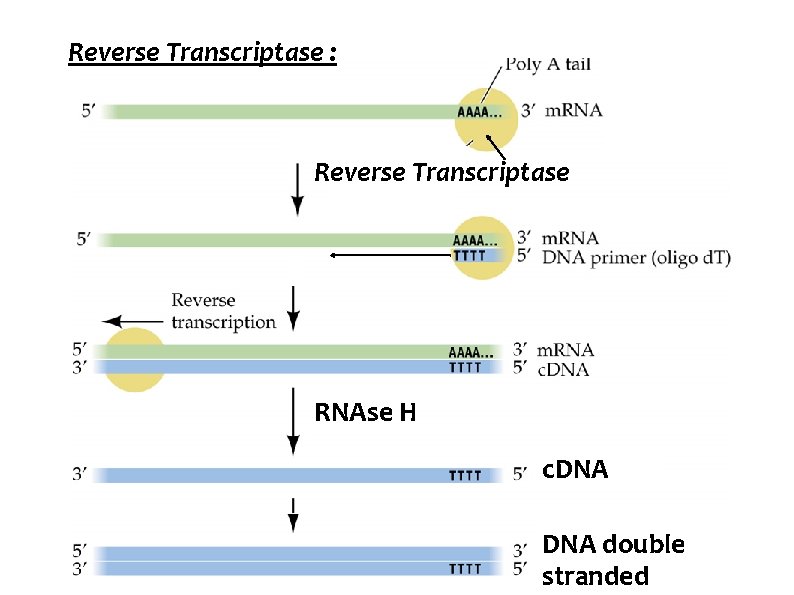
Reverse Transcriptase : Reverse Transcriptase RNAse H c. DNA double stranded

Reverse Transcriptase : Ø RNA dependant DNA Polymerase Ø Present in RNA containing viruses - HIV causative agent for AIDS. Ø Ø Ø Retroviruses - contain Reverse transcriptase and integrase enzymes. Function : Made use of in recombinant technology to make DNA sequence from RNA genome of viruses. Protooncogenes and Oncogenes are derived from retroviruses. Ø 41
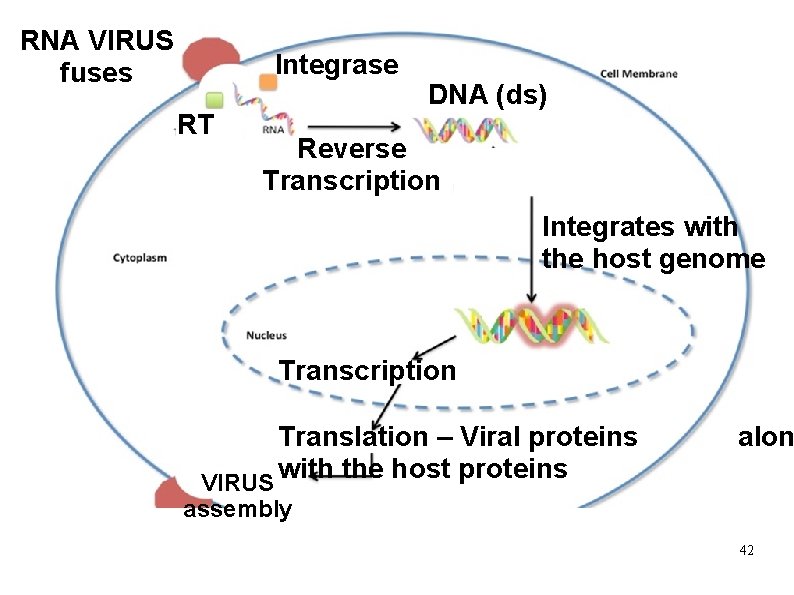
RNA VIRUS fuses Integrase RT DNA (ds) Reverse Transcription Integrates with the host genome Transcription Translation – Viral proteins with the host proteins VIRUS alon assembly 42

1. Transcription (10) 2. What is transcription? Describe the steps of transcription. (10) 3. Give a detailed account of transcription process. How is it regulated? Name the inhibitors of transcription (10) 4. Synthesis of m. RNA (5) 5. Name two inhibitors of RNA synthesis. (3) 6. Posttranscriptional modification of m. RNA (3) 43

THANK YOU 44
- Slides: 44