Molecular biology Session III Cytogenetics Ali Barzegar May

Molecular biology Session III Cytogenetics Ali Barzegar May 2020
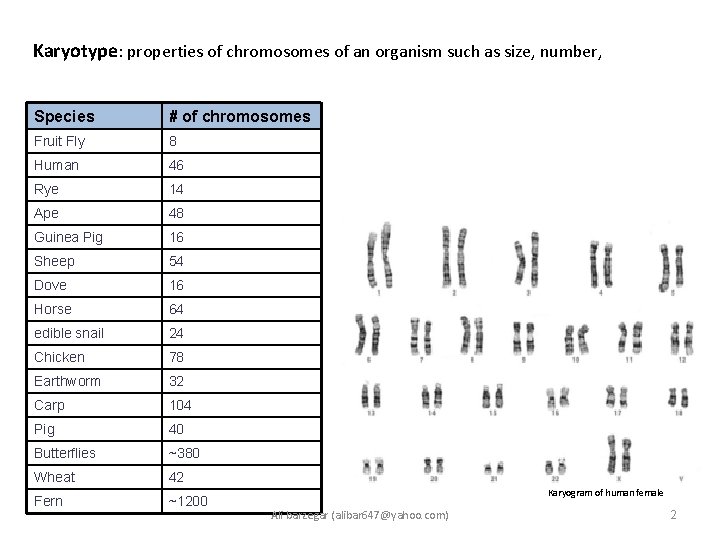
Karyotype: properties of chromosomes of an organism such as size, number, Species # of chromosomes Fruit Fly 8 Human 46 Rye (Roggen) 14 Ape 48 Guinea Pig 16 Sheep 54 Dove (Taube) 16 Horse 64 edible snail 24 Chicken 78 Earthworm 32 Carp (Karpfen) 104 Pig 40 Butterflies ~380 Wheat 42 Fern (Farn) ~1200 Karyogram of human female Ali barzegar (alibar 647@yahoo. com) 2
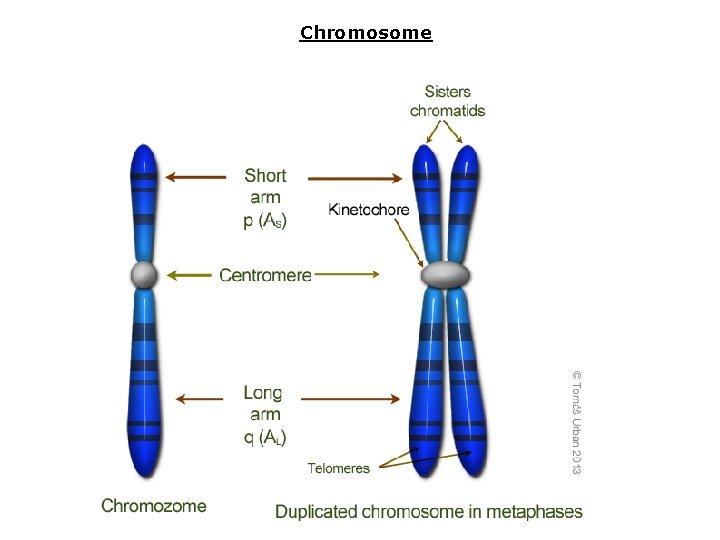
Chromosome
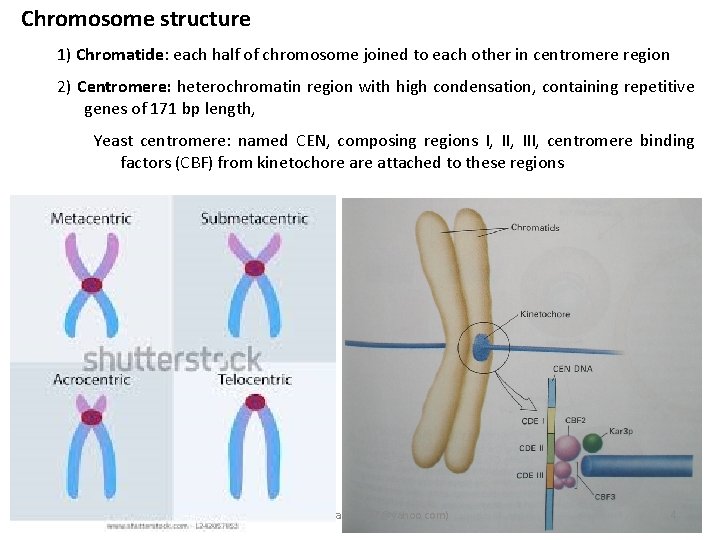
Chromosome structure 1) Chromatide: each half of chromosome joined to each other in centromere region 2) Centromere: heterochromatin region with high condensation, containing repetitive genes of 171 bp length, Yeast centromere: named CEN, composing regions I, III, centromere binding factors (CBF) from kinetochore attached to these regions Ali barzegar (alibar 647@yahoo. com) 4

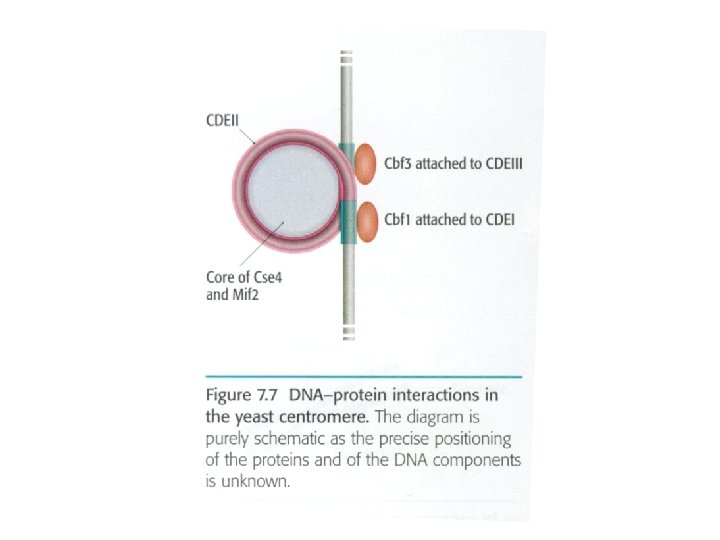
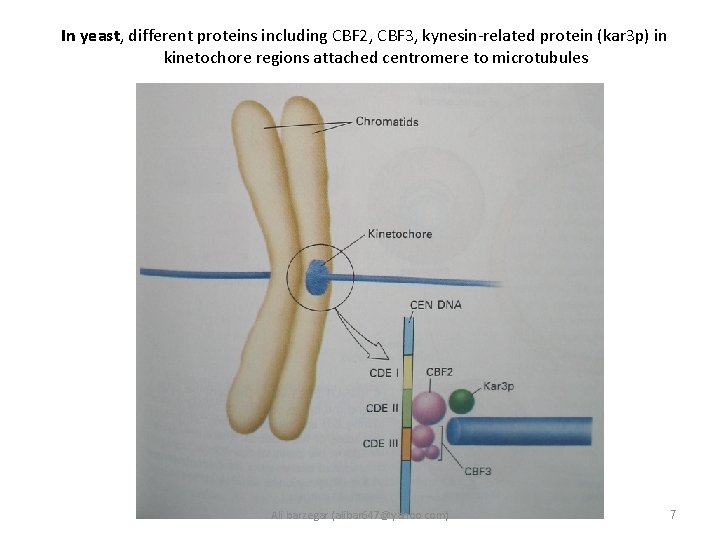
In yeast, different proteins including CBF 2, CBF 3, kynesin-related protein (kar 3 p) in kinetochore regions attached centromere to microtubules Ali barzegar (alibar 647@yahoo. com) 7
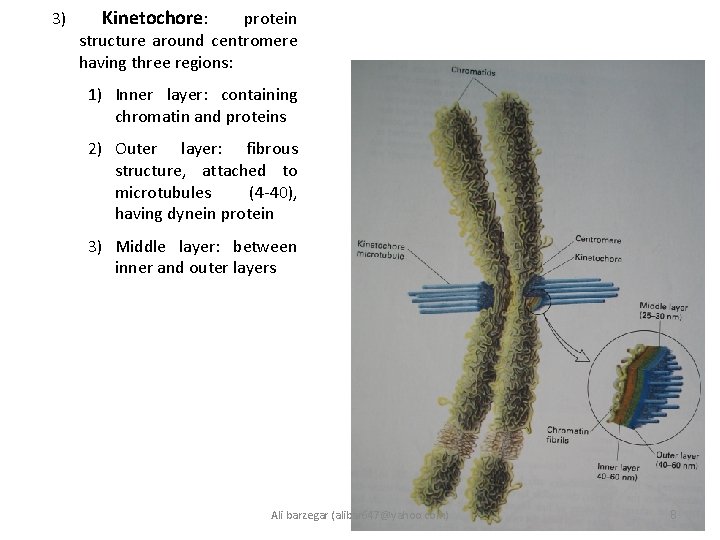
3) Kinetochore: protein structure around centromere having three regions: 1) Inner layer: containing chromatin and proteins 2) Outer layer: fibrous structure, attached to microtubules (4 -40), having dynein protein 3) Middle layer: between inner and outer layers Ali barzegar (alibar 647@yahoo. com) 8
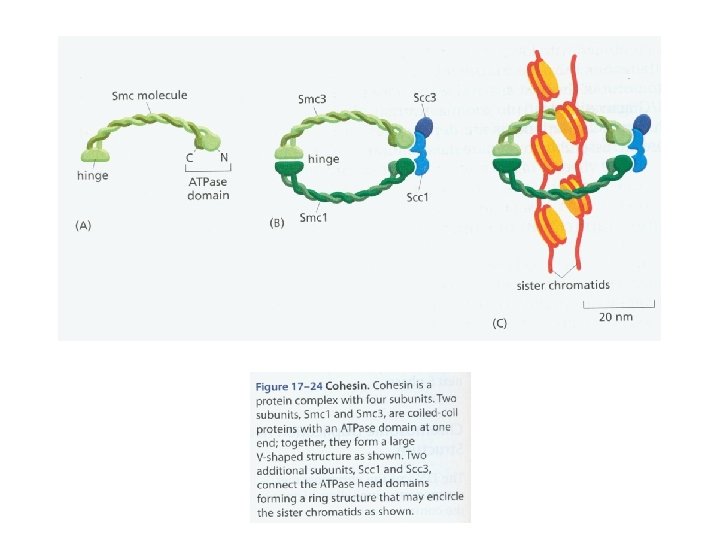
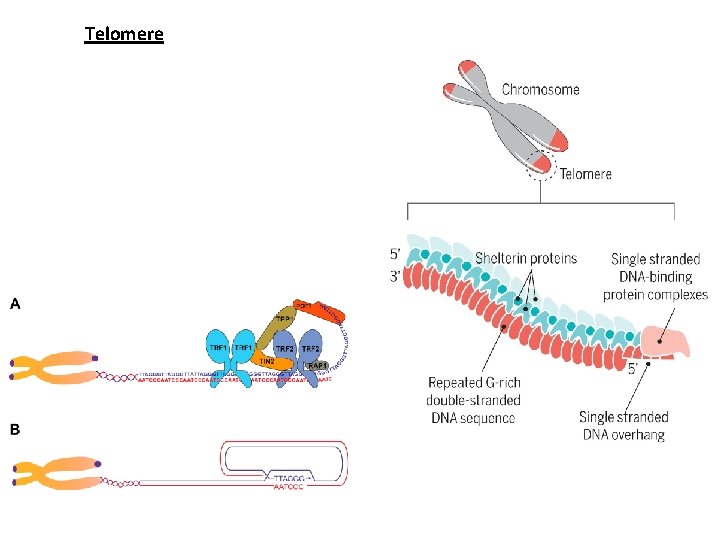
Telomere
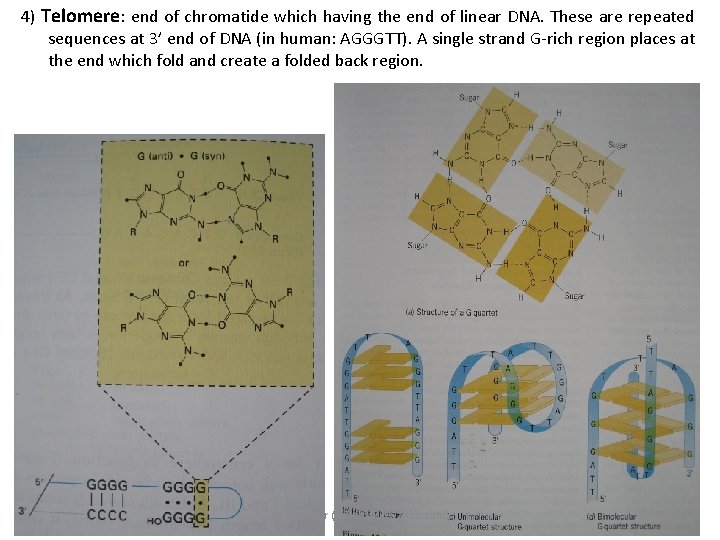
4) Telomere: end of chromatide which having the end of linear DNA. These are repeated sequences at 3’ end of DNA (in human: AGGGTT). A single strand G-rich region places at the end which fold and create a folded back region. Ali barzegar (alibar 647@yahoo. com) 11
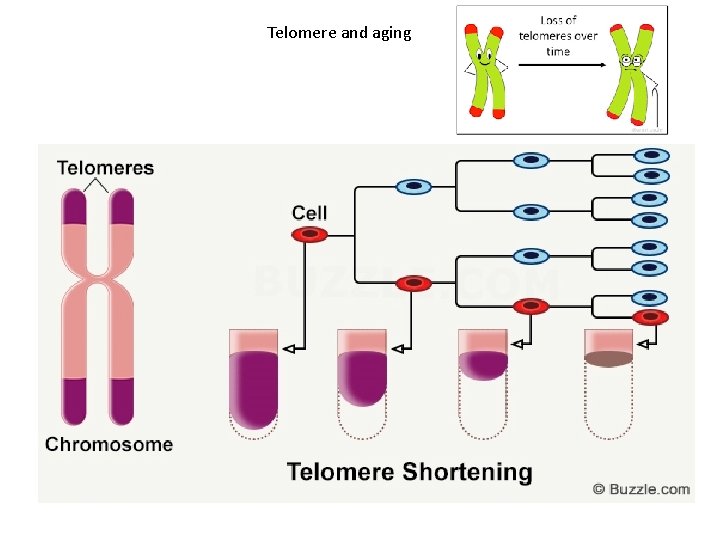
Telomere and aging
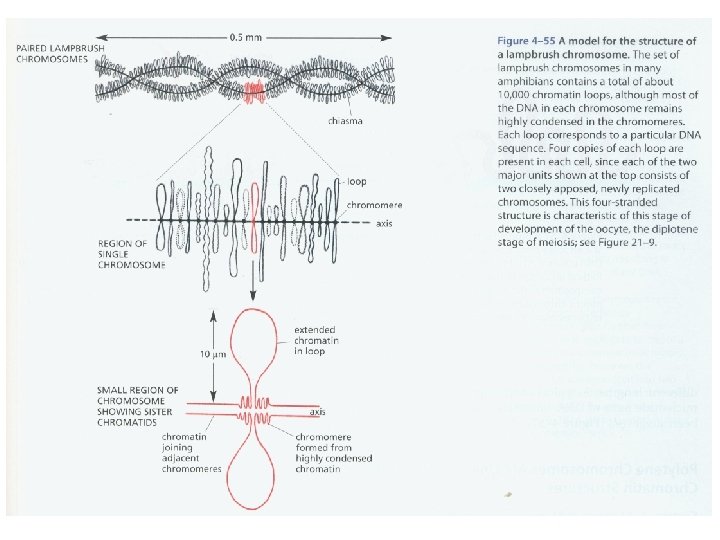
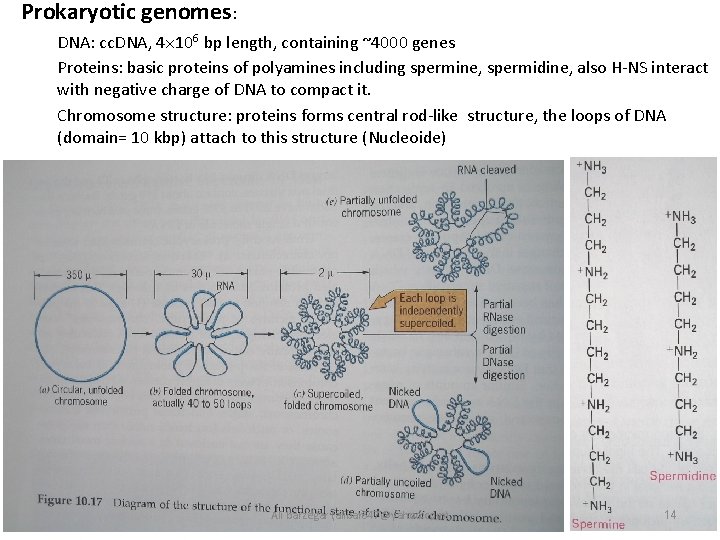
Prokaryotic genomes: DNA: cc. DNA, 4 106 bp length, containing ~4000 genes Proteins: basic proteins of polyamines including spermine, spermidine, also H-NS interact with negative charge of DNA to compact it. Chromosome structure: proteins forms central rod-like structure, the loops of DNA (domain= 10 kbp) attach to this structure (Nucleoide) Ali barzegar (alibar 647@yahoo. com) 14
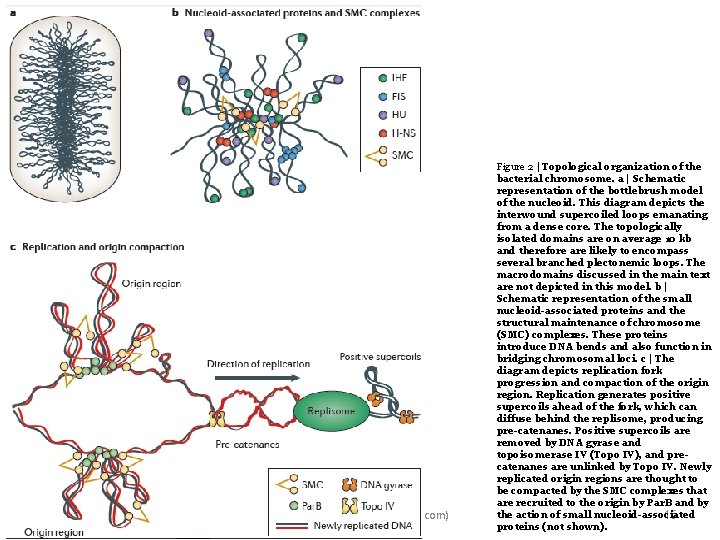
Ali barzegar (alibar 647@yahoo. com) Figure 2 | Topological organization of the bacterial chromosome. a | Schematic representation of the bottlebrush model of the nucleoid. This diagram depicts the interwound supercoiled loops emanating from a dense core. The topologically isolated domains are on average 10 kb and therefore are likely to encompass several branched plectonemic loops. The macrodomains discussed in the main text are not depicted in this model. b | Schematic representation of the small nucleoid-associated proteins and the structural maintenance of chromosome (SMC) complexes. These proteins introduce DNA bends and also function in bridging chromosomal loci. c | The diagram depicts replication fork progression and compaction of the origin region. Replication generates positive supercoils ahead of the fork, which can diffuse behind the replisome, producing pre-catenanes. Positive supercoils are removed by DNA gyrase and topoisomerase IV (Topo IV), and precatenanes are unlinked by Topo IV. Newly replicated origin regions are thought to be compacted by the SMC complexes that are recruited to the origin by Par. B and by the action of small nucleoid-associated 15 proteins (not shown).
- Slides: 15