Molecular Biology II Recombinant DNA Technology Biotechnology terminology

Molecular Biology II Recombinant DNA Technology • Biotechnology terminology • Common hosts and experimental organisms • Transcription and translation • Prokaryotic gene organization & expression

Terminology • Molecular biology-The study of biology on a molecular level including the structure, function, and makeup of biologically important molecules such as DNA, RNA, and proteins • Recombinant DNA technology-a set of techniques for manipulating DNA, including: the identification and cloning of genes; the study of the expression of cloned genes; and the production of large quantities of gene product • Genetic engineering-the process of transferring DNA from one organism into another that results in a genetic modification • Biotechnology-production of goods and services using biological organisms, systems, and processes • Molecular biotechnology-r. DNA technology + biotechnology
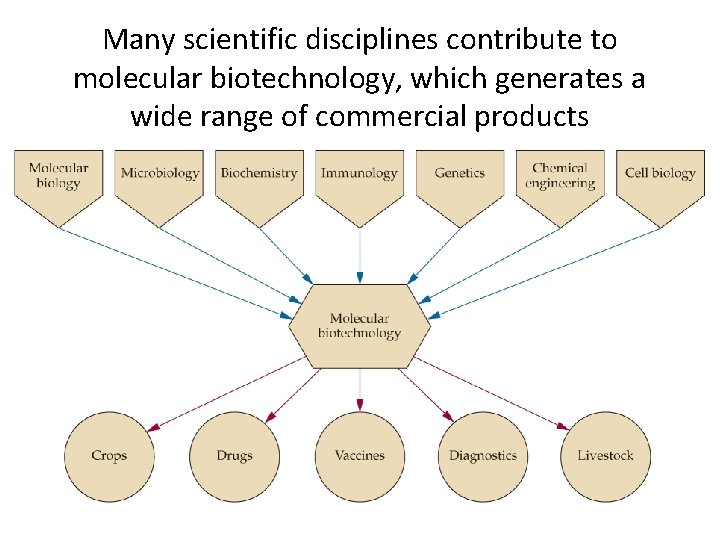
Many scientific disciplines contribute to molecular biotechnology, which generates a wide range of commercial products

Common host organisms used in molecular biotechnology • • • E. coli Yeast (Saccharomyces cerevisiae) Insect cell lines Plant cell lines Animal cell lines
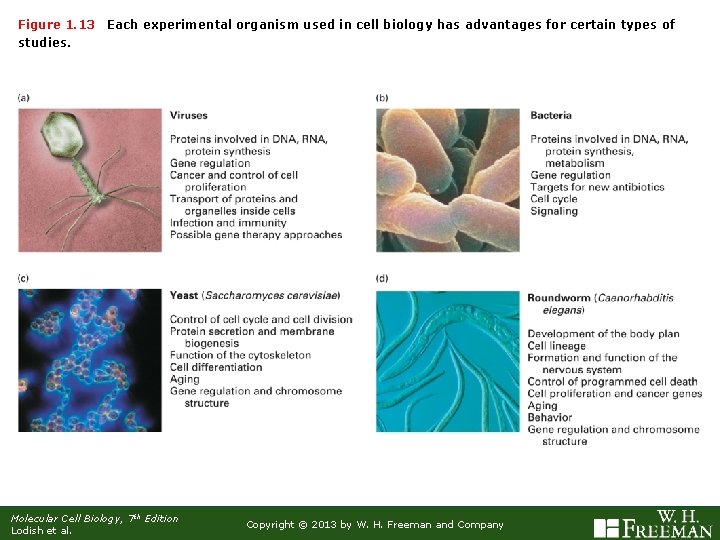
Figure 1. 13 studies. Each experimental organism used in cell biology has advantages for certain types of Molecular Cell Biology, 7 th Edition Lodish et al. Copyright © 2013 by W. H. Freeman and Company

Figure 1. 13 (Continued) Each experimental organism used in cell biology has advantages for certain types of studies. Listen to the podcast! Molecular Cell Biology, 7 th Edition Lodish et al. Copyright © 2013 by W. H. Freeman and Company
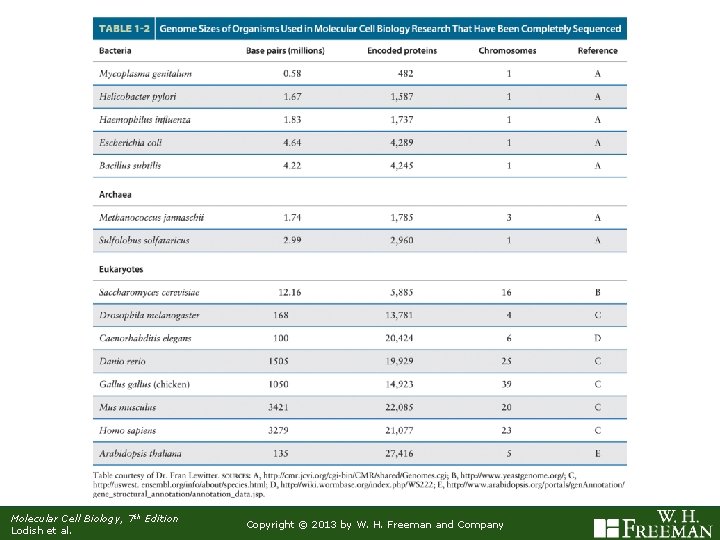
Molecular Cell Biology, 7 th Edition Lodish et al. Copyright © 2013 by W. H. Freeman and Company
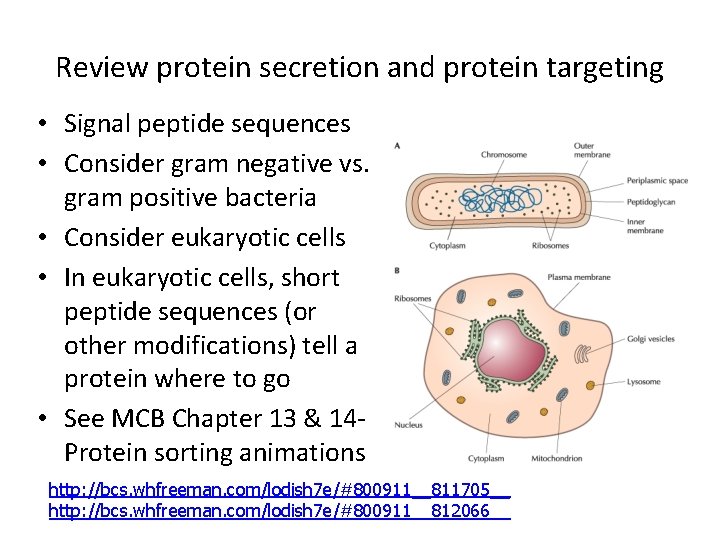
Review protein secretion and protein targeting • Signal peptide sequences • Consider gram negative vs. gram positive bacteria • Consider eukaryotic cells • In eukaryotic cells, short peptide sequences (or other modifications) tell a protein where to go • See MCB Chapter 13 & 14 Protein sorting animations http: //bcs. whfreeman. com/lodish 7 e/#800911__811705__ http: //bcs. whfreeman. com/lodish 7 e/#800911__812066__
Central Dogma of Biology DNA transcription RNA reverse transcription DNA replication translation Protein $$$ Molecular Biotechnology
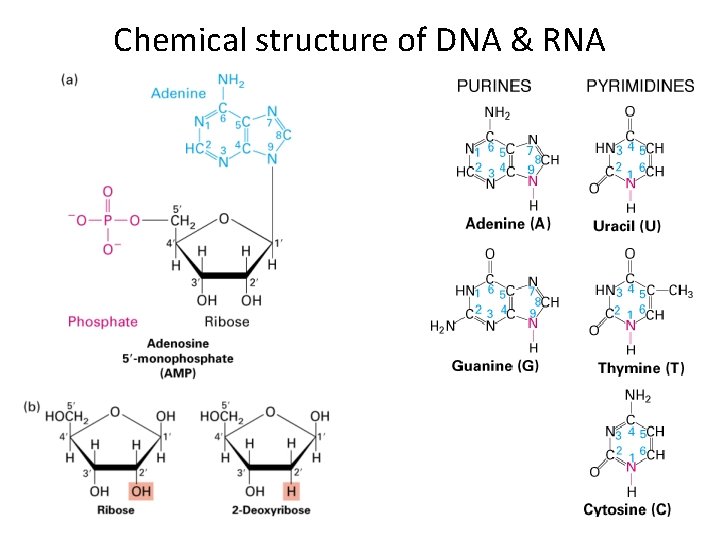
Chemical structure of DNA & RNA
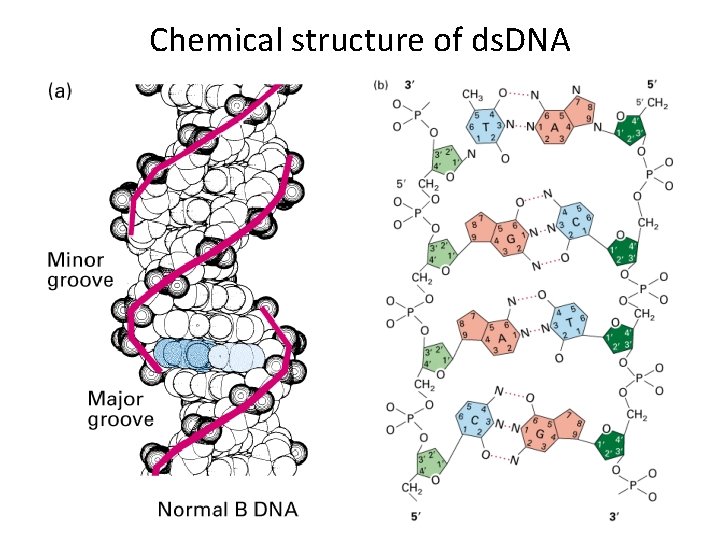
Chemical structure of ds. DNA
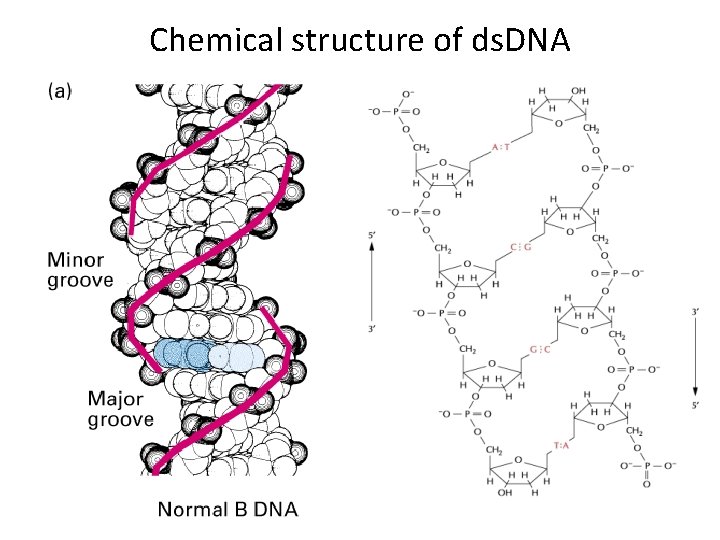
Chemical structure of ds. DNA
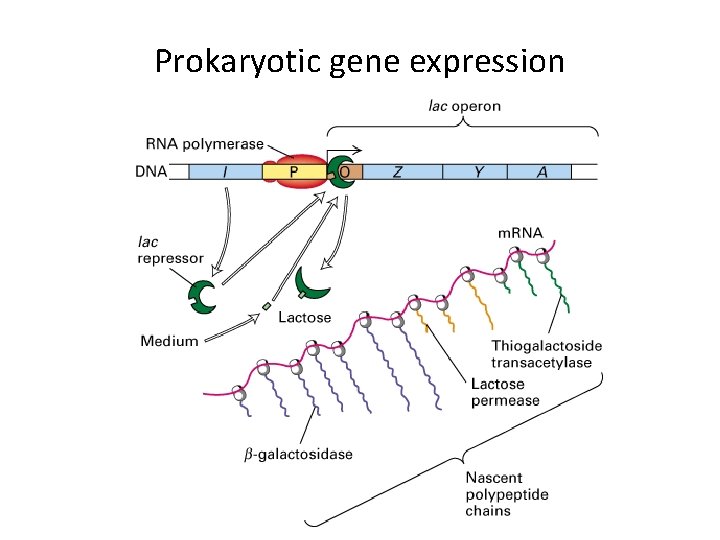
Prokaryotic gene expression
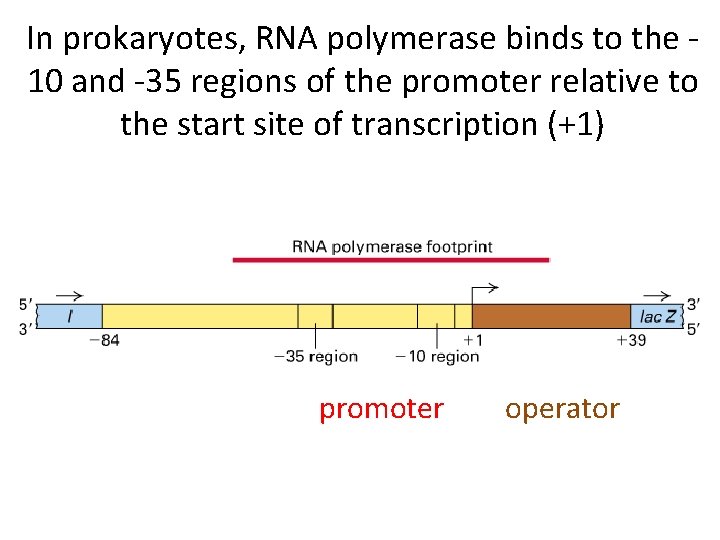
In prokaryotes, RNA polymerase binds to the 10 and -35 regions of the promoter relative to the start site of transcription (+1) promoter operator

• Eukaryotic gene organization • Restriction enzymes • Cloning vectors
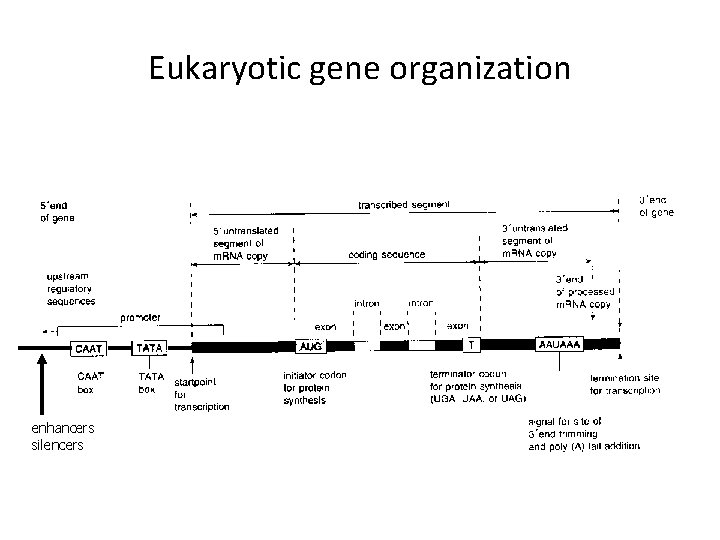
Eukaryotic gene organization enhancers silencers
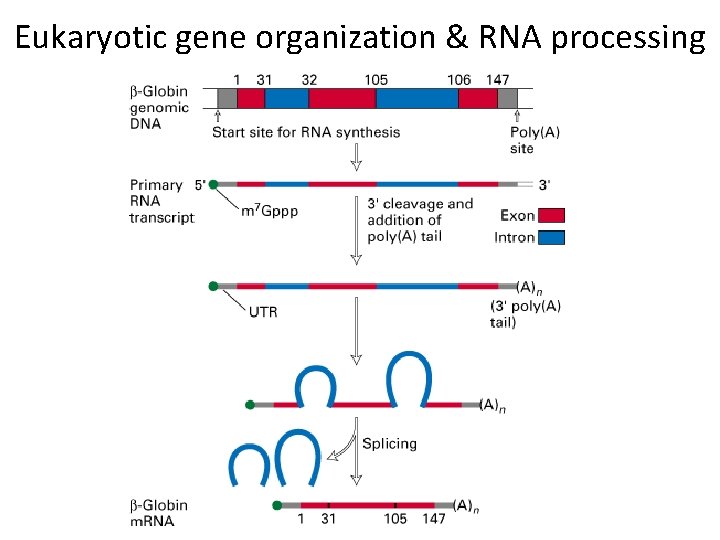
Eukaryotic gene organization & RNA processing
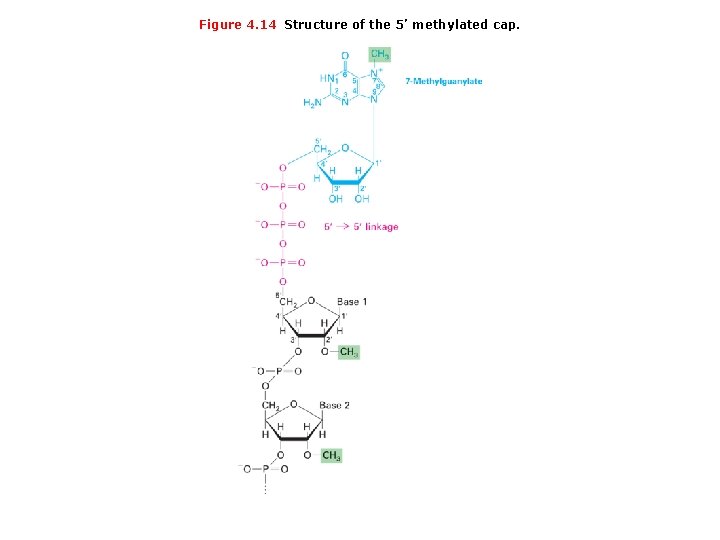
Figure 4. 14 Structure of the 5’ methylated cap.

Basic Transcriptional Mechanisms and m. RNA Splicing Animations Basic Molecular Genetic Mechanisms (animations) • Life Cycle of m. RNA u http: //bcs. whfreeman. com/lodish 7 e/#800911__812036__ • Basic Transcriptional Mechanisms u http: //bcs. whfreeman. com/lodish 7 e/#800911__812037__ Post-transcriptional Gene Control (animation) • m. RNA Splicing u http: //bcs. whfreeman. com/lodish 7 e/#800911__812057__
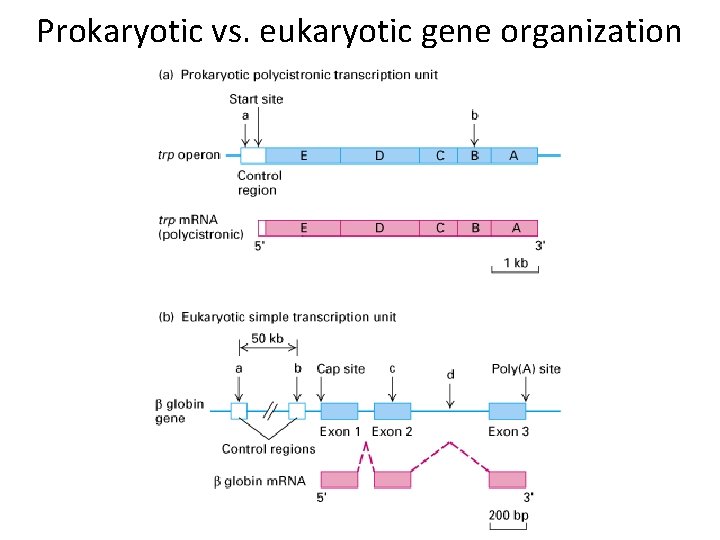
Prokaryotic vs. eukaryotic gene organization
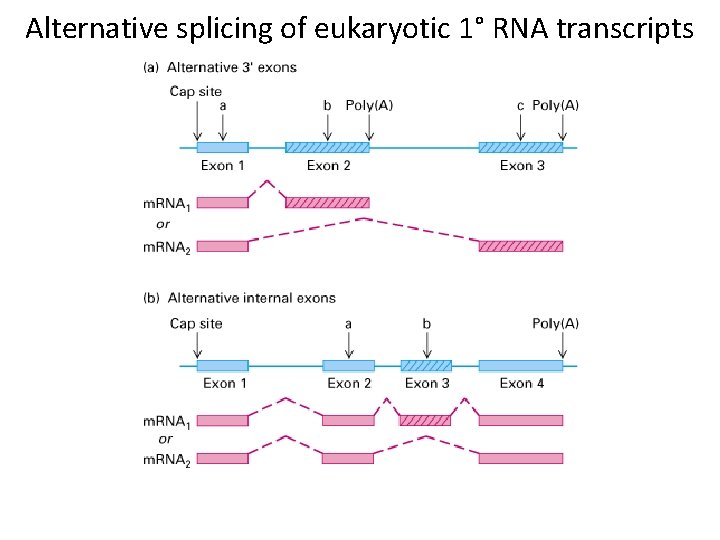
Alternative splicing of eukaryotic 1° RNA transcripts
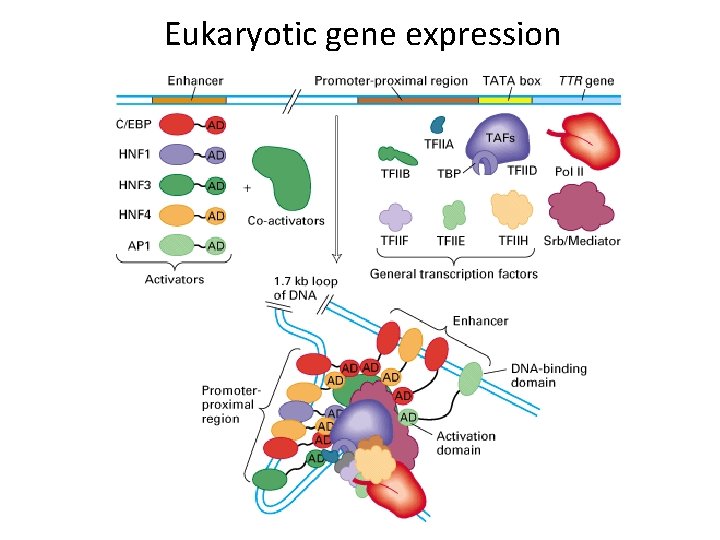
Eukaryotic gene expression

Life Cycle of m. RNA Basic Molecular Genetic Mechanisms (animation) • Life Cycle of m. RNA u http: //bcs. whfreeman. com/lodish 7 e/#800911__812036__

Translation Or. R
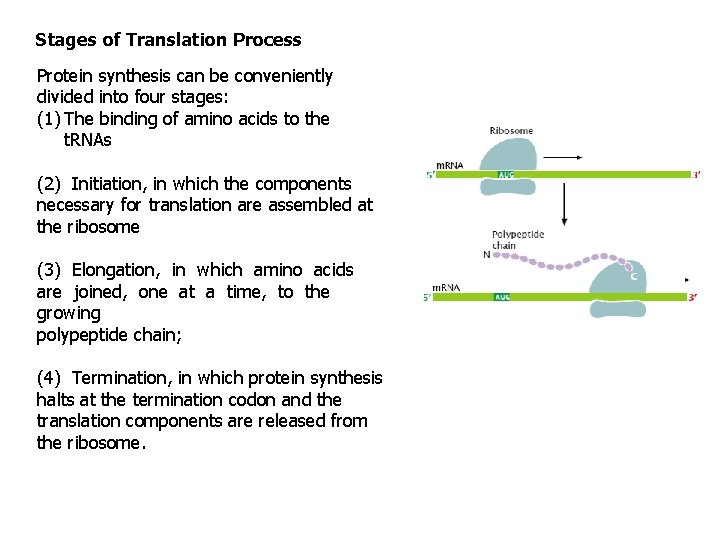
Stages of Translation Process Protein synthesis can be conveniently divided into four stages: (1) The binding of amino acids to the t. RNAs (2) Initiation, in which the components necessary for translation are assembled at the ribosome (3) Elongation, in which amino acids are joined, one at a time, to the growing polypeptide chain; (4) Termination, in which protein synthesis halts at the termination codon and the translation components are released from the ribosome.
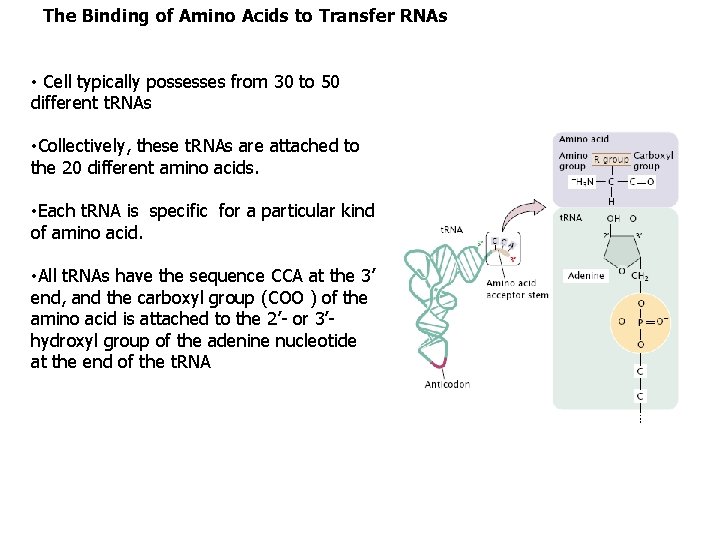
The Binding of Amino Acids to Transfer RNAs • Cell typically possesses from 30 to 50 different t. RNAs • Collectively, these t. RNAs are attached to the 20 different amino acids. • Each t. RNA is specific for a particular kind of amino acid. • All t. RNAs have the sequence CCA at the 3’ end, and the carboxyl group (COO ) of the amino acid is attached to the 2’- or 3’hydroxyl group of the adenine nucleotide at the end of the t. RNA

If each t. RNA is specific for a particular amino acid but all amino acids are attached to the same nucleotide (A) at the 3’ end of a t. RNA, how does a t. RNA link up with its appropriate amino acid? The key to specificity between an amino acid and its t. RNA is a set of enzymes called aminoacyl-t. RNA synthetases. A cell has 20 different aminoacyl-t. RNA synthetases, one for each of the 20 amino acids. Each synthetase recognizes a particular amino acid, as well as all the t. RNAs that accept that amino acid. Recognition of the appropriate amino acid by a synthetase is based on the different sizes, charges, and R groups of the amino acids.
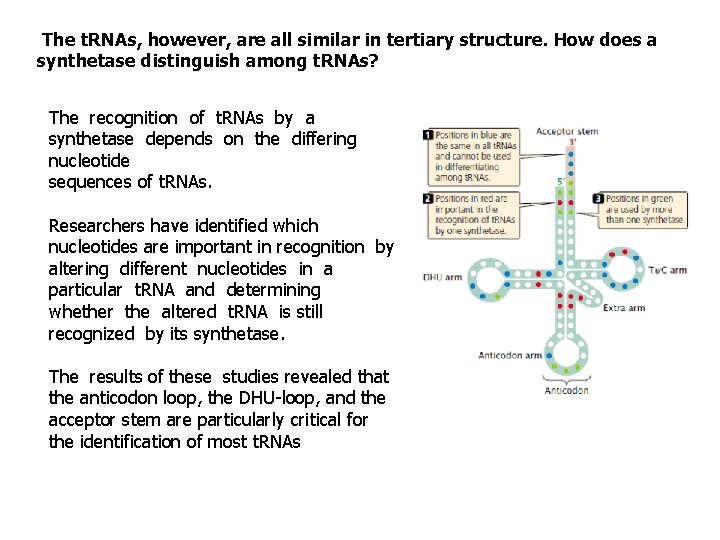
The t. RNAs, however, are all similar in tertiary structure. How does a synthetase distinguish among t. RNAs? The recognition of t. RNAs by a synthetase depends on the differing nucleotide sequences of t. RNAs. Researchers have identified which nucleotides are important in recognition by altering different nucleotides in a particular t. RNA and determining whether the altered t. RNA is still recognized by its synthetase. The results of these studies revealed that the anticodon loop, the DHU-loop, and the acceptor stem are particularly critical for the identification of most t. RNAs
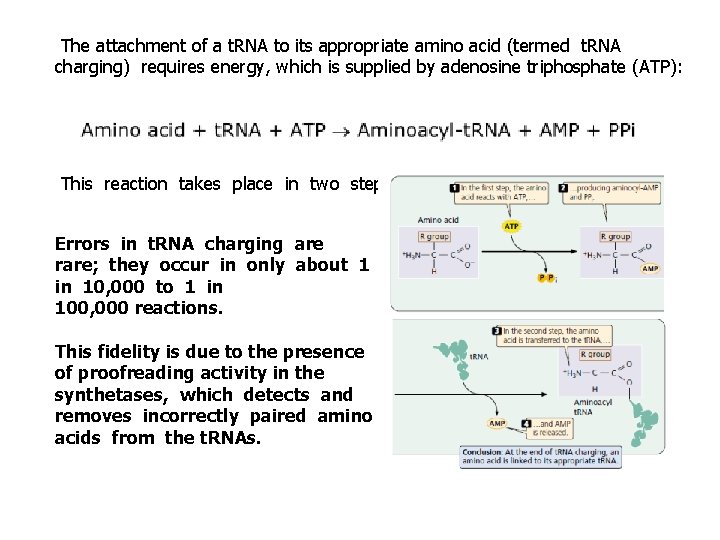
The attachment of a t. RNA to its appropriate amino acid (termed t. RNA charging) requires energy, which is supplied by adenosine triphosphate (ATP): This reaction takes place in two steps: Errors in t. RNA charging are rare; they occur in only about 1 in 10, 000 to 1 in 100, 000 reactions. This fidelity is due to the presence of proofreading activity in the synthetases, which detects and removes incorrectly paired amino acids from the t. RNAs.

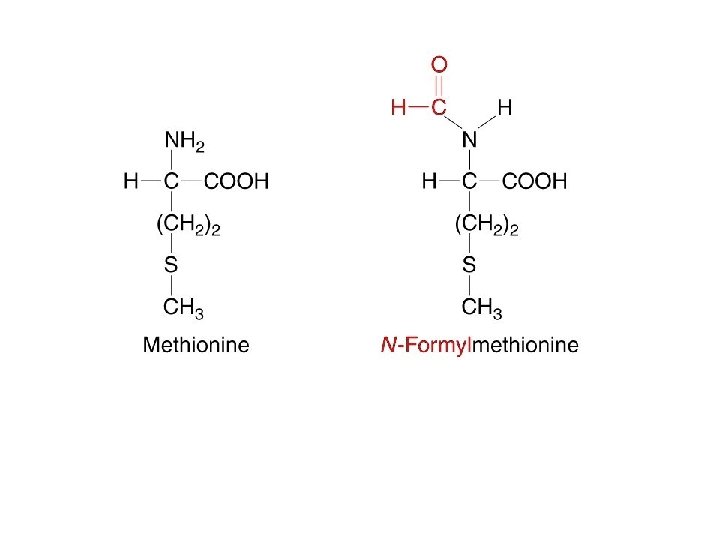
The Initiation of Translation Prokaryotic Initiation System The second stage in the process of protein synthesis is initiation. During initiation, all the components necessary for protein synthesis assemble: (1) m. RNA (2) the small and large subunits of the ribosome (3) a set of three proteins called initiation factors (4) initiator t. RNA with N-formylmethionine attached (f. Met-t. RNA f. Met ) (5) guanosine triphosphate (GTP). Initiation comprises three major steps.

The Initiation of Translation Prokaryotic Initiation System Initiation comprises three major steps. First, m. RNA binds to the small subunit of the ribosome. Second, initiator t. RNA binds to the m. RNA through base pairing between the codon and anticodon. Third, the large ribosome joins the initiation complex.
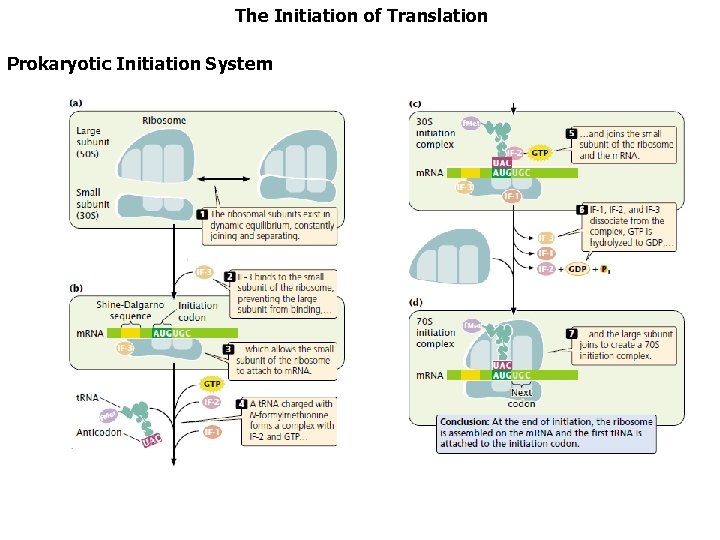
The Initiation of Translation Prokaryotic Initiation System
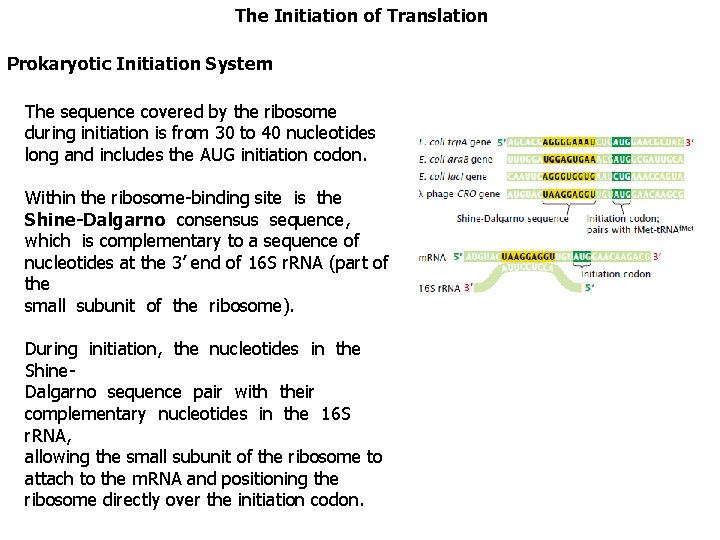
The Initiation of Translation Prokaryotic Initiation System The sequence covered by the ribosome during initiation is from 30 to 40 nucleotides long and includes the AUG initiation codon. Within the ribosome-binding site is the Shine-Dalgarno consensus sequence, which is complementary to a sequence of nucleotides at the 3’ end of 16 S r. RNA (part of the small subunit of the ribosome). During initiation, the nucleotides in the Shine. Dalgarno sequence pair with their complementary nucleotides in the 16 S r. RNA, allowing the small subunit of the ribosome to attach to the m. RNA and positioning the ribosome directly over the initiation codon.

The Initiation of Translation Eukaryotic Initiation System The small subunit of the eukaryotic ribosome, with the help of initiation factors, recognizes the cap and binds there; the small subunit then migrates along (scans) the m. RNA until it locates the first AUG codon. The identification of the start codon is facilitated by the presence of a consensus sequence (called the Kozak sequence) that surrounds the start codon:

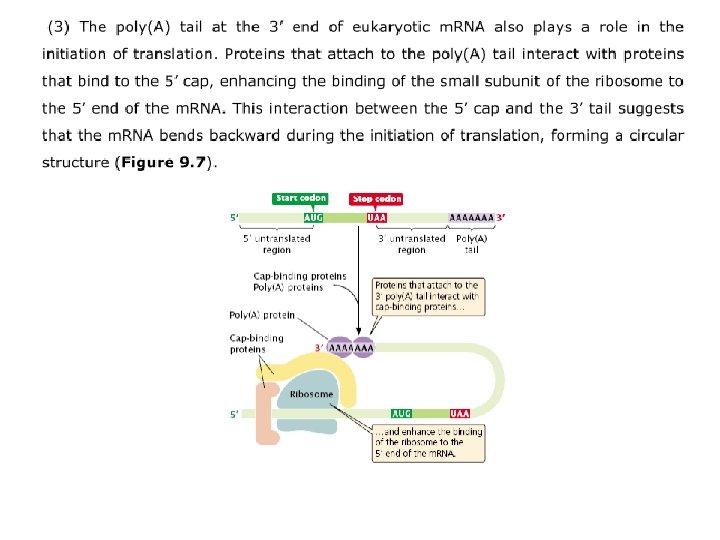

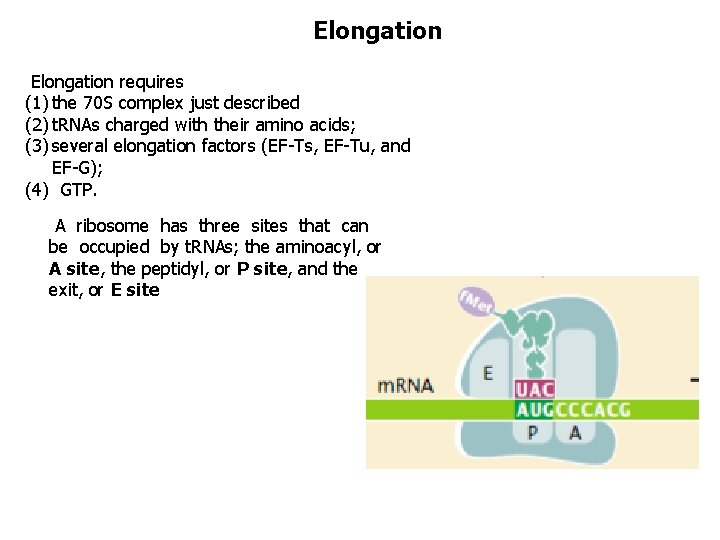
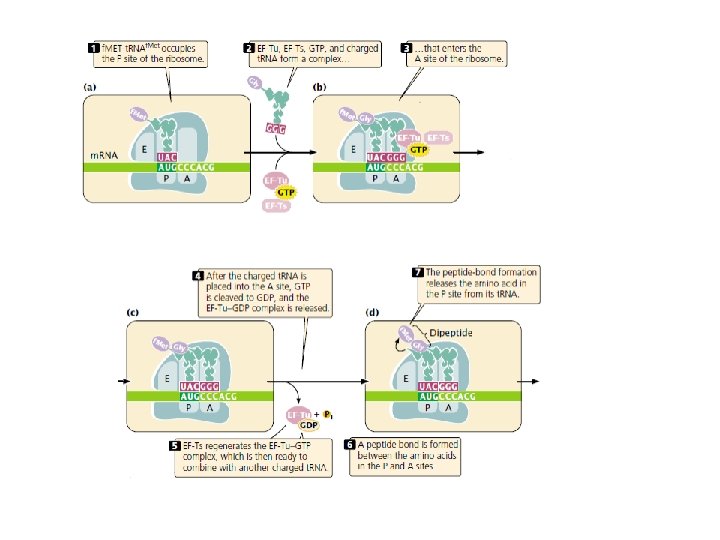
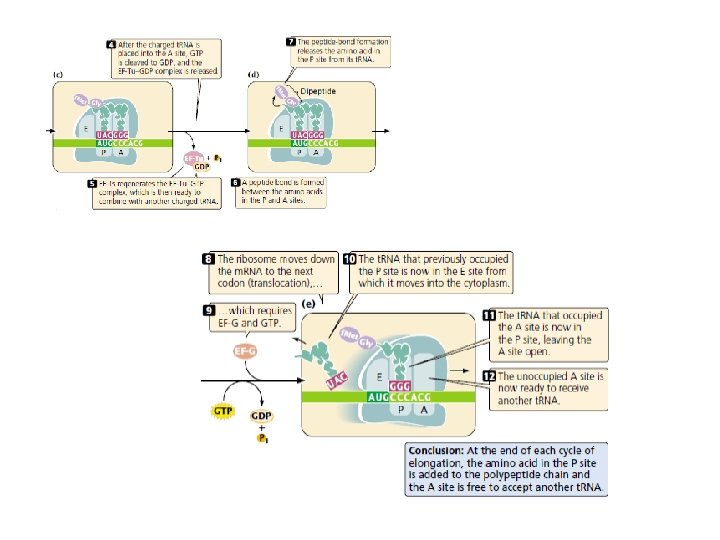
Elongation requires (1) the 70 S complex just described (2) t. RNAs charged with their amino acids; (3) several elongation factors (EF-Ts, EF-Tu, and EF-G); (4) GTP. A ribosome has three sites that can be occupied by t. RNAs; the aminoacyl, or A site, the peptidyl, or P site, and the exit, or E site
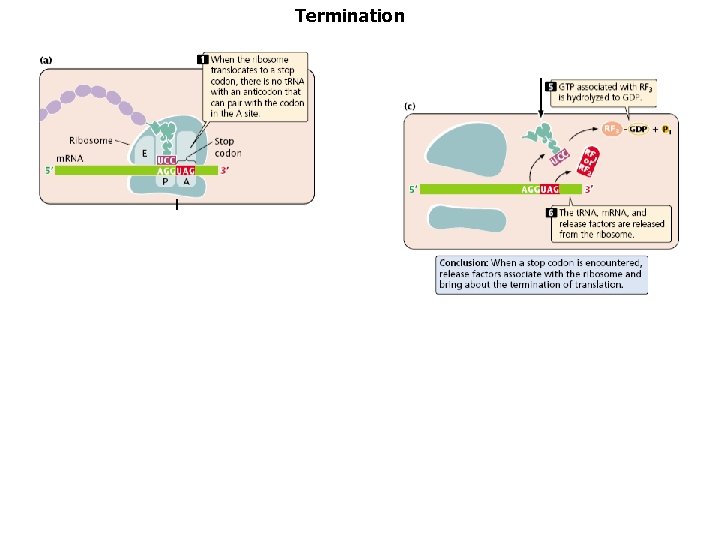
Termination
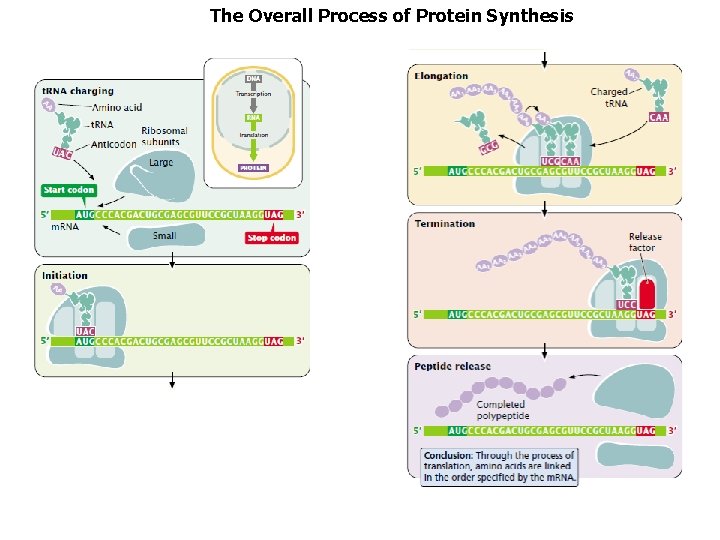
The Overall Process of Protein Synthesis
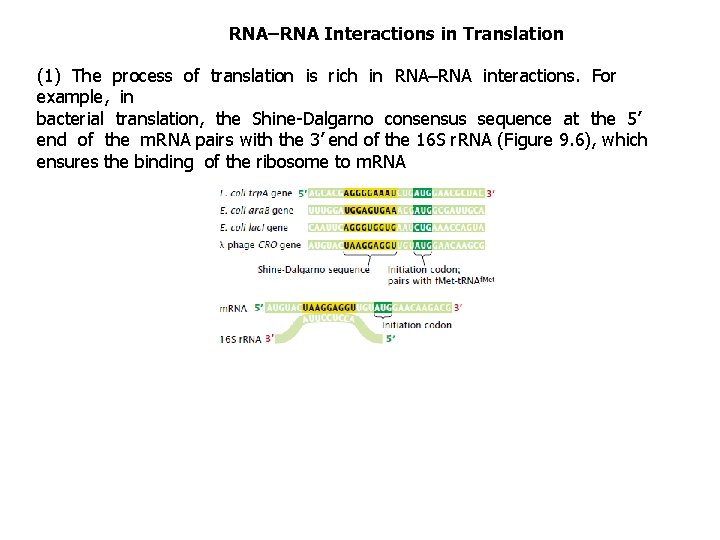
RNA–RNA Interactions in Translation (1) The process of translation is rich in RNA–RNA interactions. For example, in bacterial translation, the Shine-Dalgarno consensus sequence at the 5’ end of the m. RNA pairs with the 3’ end of the 16 S r. RNA (Figure 9. 6), which ensures the binding of the ribosome to m. RNA


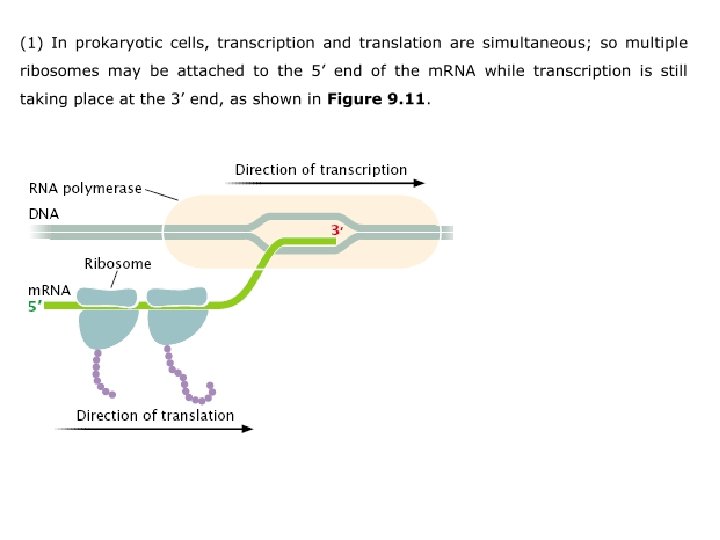
Polyribosomes In both prokaryotic and eukaryotic cells, m. RNA molecules are translated simultaneously by multiple ribosomes. The resulting structure—an m. RNA with several ribosomes attached—is called a polyribosome. Each ribosome successively attaches to the ribosome- binding site at the 5’ end of the m. RNA and moves toward the 3’ end; the polypeptide associated with each ribosome becomes progressively longer as the ribosome moves along the m. RNA.

A Comparison of Bacterial and Eukaryotic Translation 1. Initiator Amino Acid 2. Temporal and Spatial Differences of Gene Exp 3. Longevity of m. RNA 4. Sizes and Compositions Ribosomal Subunit 5. Initiation of Transcription 6. Elongation and Termination

The Posttranslational Modifications of Proteins

Translation and Antibiotics

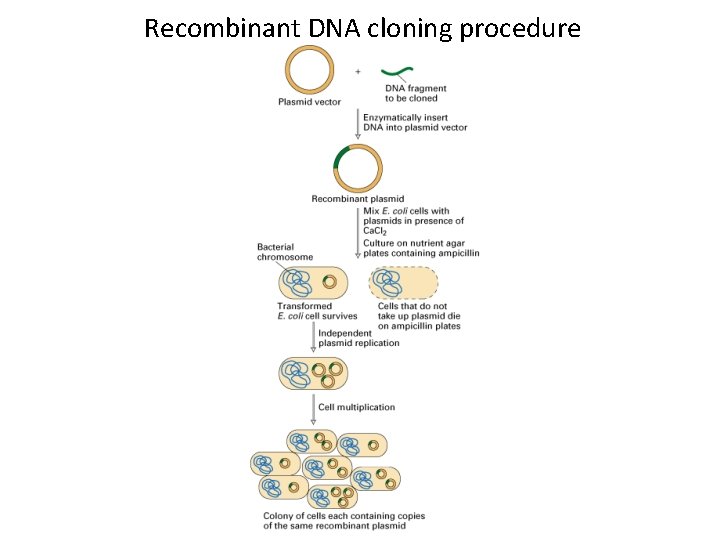
Recombinant DNA cloning procedure

Recombinant DNA cloning procedure MCB Chapter 5 - Molecular Genetic Techniques (animation) • Plasmid Cloning u http: //bcs. whfreeman. com/lodish 7 e/#800911__812047__
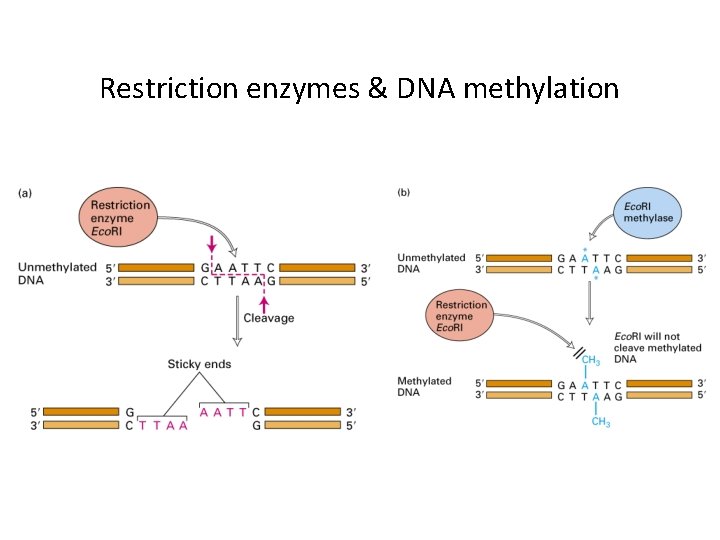
Restriction enzymes & DNA methylation
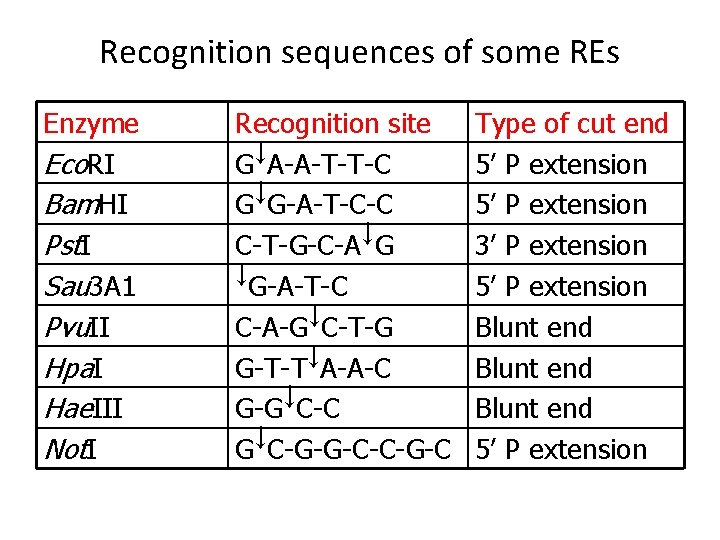
Recognition sequences of some REs Enzyme Eco. RI Bam. HI Pst. I Sau 3 A 1 Pvu. II Hpa. I Hae. III Not. I Recognition site G↓A-A-T-T-C G↓G-A-T-C-C C-T-G-C-A↓G ↓G-A-T-C C-A-G↓C-T-G G-T-T↓A-A-C G-G↓C-C G↓C-G-G-C-C-G-C Type of cut end 5’ P extension 3’ P extension 5’ P extension Blunt end 5’ P extension
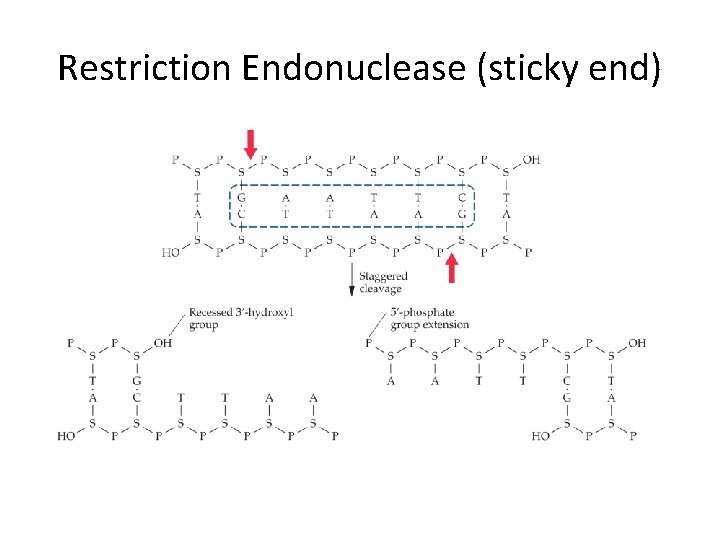
Restriction Endonuclease (sticky end)
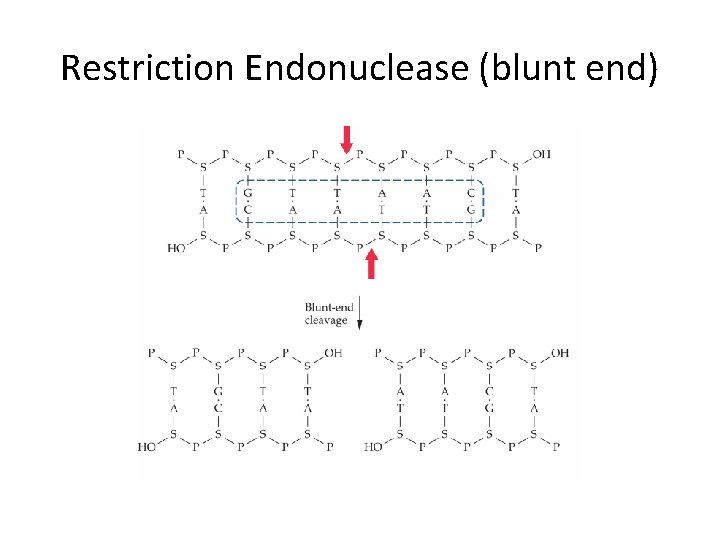
Restriction Endonuclease (blunt end)

Restriction Endonuclease Type II

Restriction Endonuclease (Type IIS)
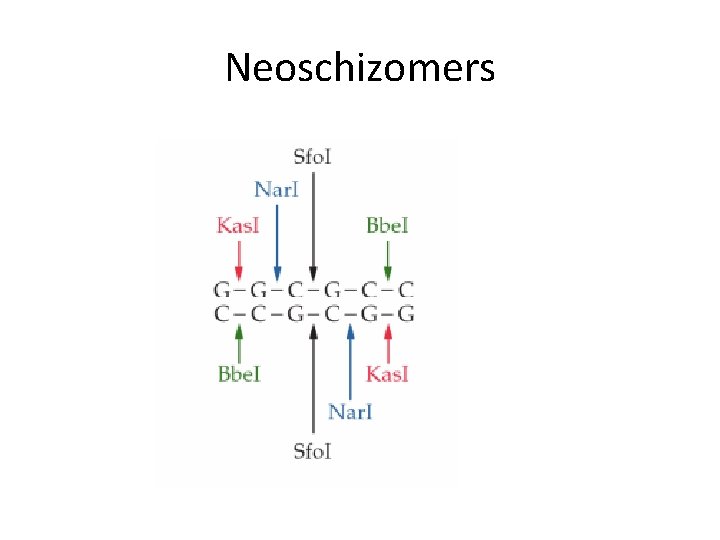
Neoschizomers
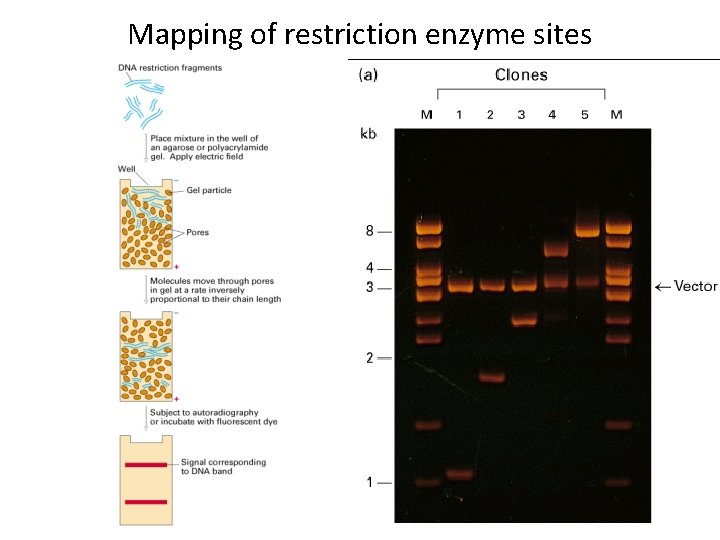
Mapping of restriction enzyme sites
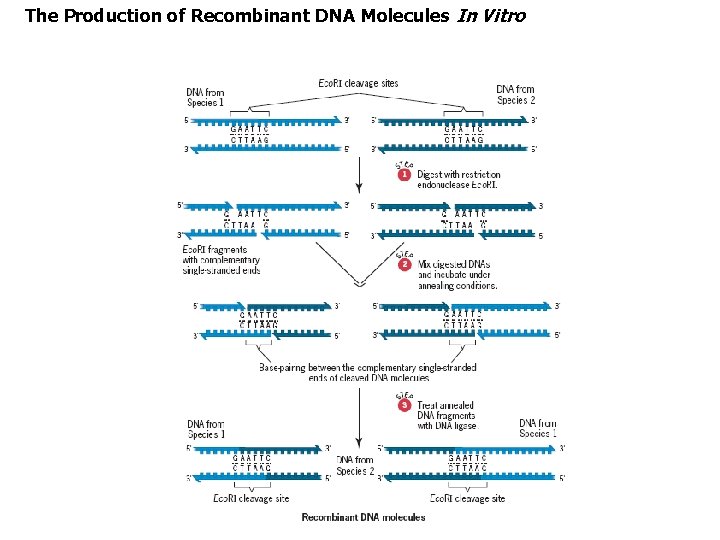
The Production of Recombinant DNA Molecules In Vitro
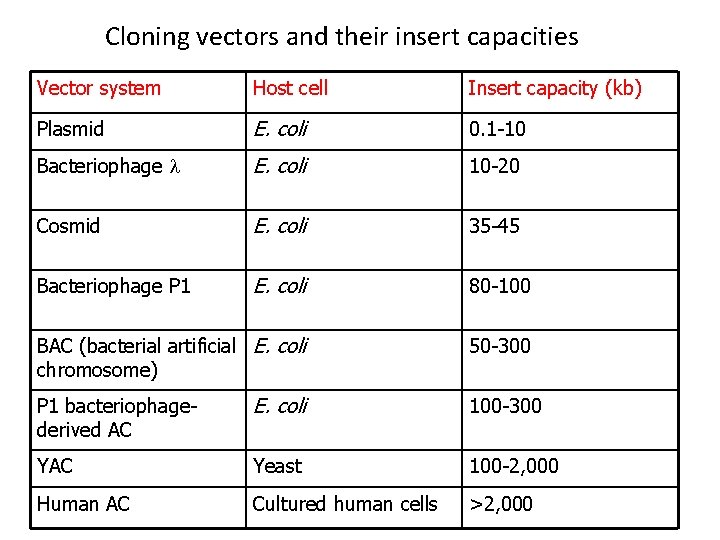
Cloning vectors and their insert capacities Vector system Host cell Insert capacity (kb) Plasmid E. coli 0. 1 -10 Bacteriophage l E. coli 10 -20 Cosmid E. coli 35 -45 Bacteriophage P 1 E. coli 80 -100 BAC (bacterial artificial E. coli chromosome) 50 -300 P 1 bacteriophagederived AC E. coli 100 -300 YAC Yeast 100 -2, 000 Human AC Cultured human cells >2, 000
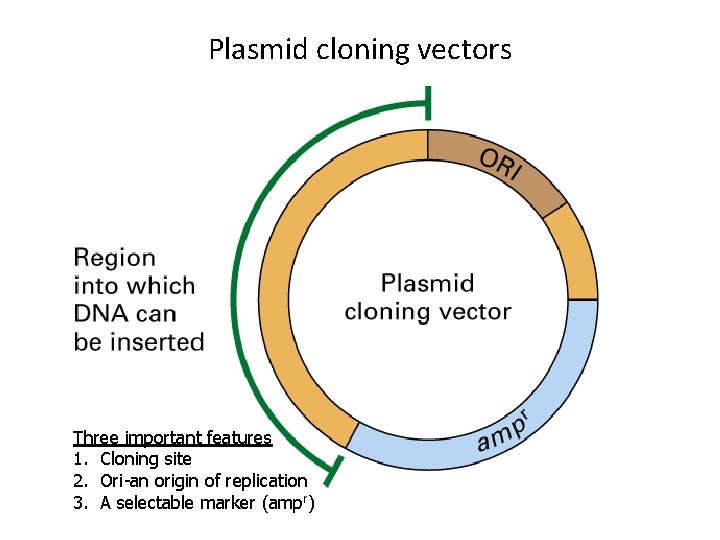
Plasmid cloning vectors Three important features 1. Cloning site 2. Ori-an origin of replication 3. A selectable marker (ampr)
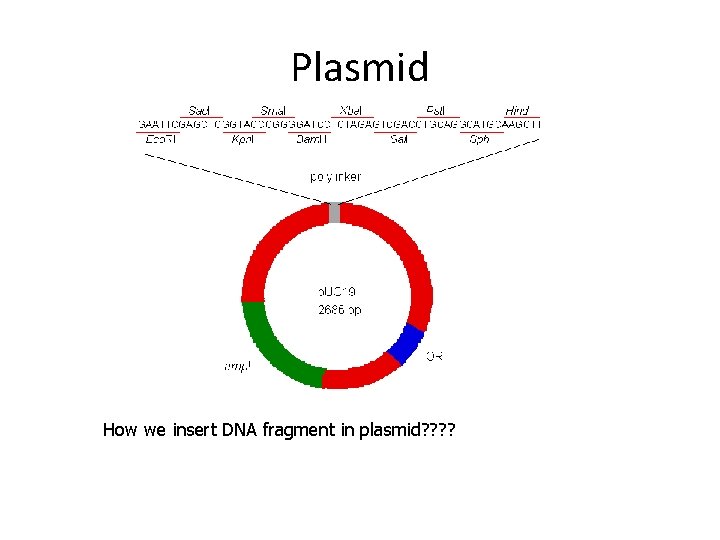
Plasmid How we insert DNA fragment in plasmid? ?
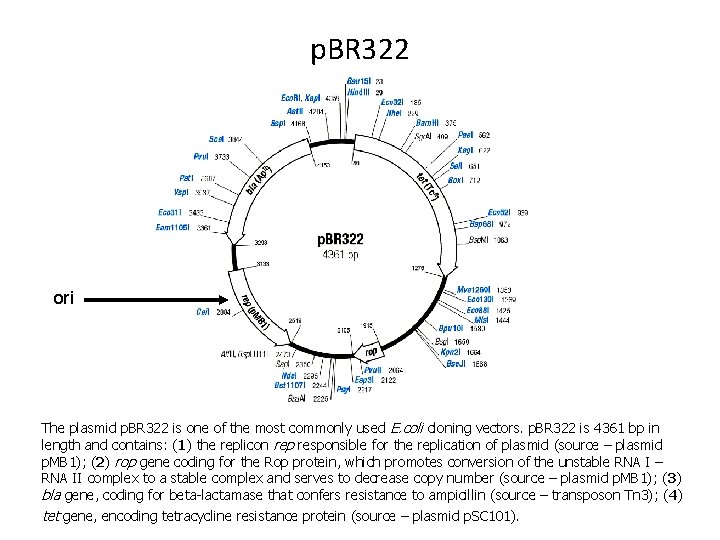
p. BR 322 ori The plasmid p. BR 322 is one of the most commonly used E. coli cloning vectors. p. BR 322 is 4361 bp in length and contains: (1) the replicon rep responsible for the replication of plasmid (source – plasmid p. MB 1); (2) rop gene coding for the Rop protein, which promotes conversion of the unstable RNA I – RNA II complex to a stable complex and serves to decrease copy number (source – plasmid p. MB 1); (3) bla gene, coding for beta-lactamase that confers resistance to ampicillin (source – transposon Tn 3); (4) tet gene, encoding tetracycline resistance protein (source – plasmid p. SC 101).

p. UC 18/19 p. UC 18 and p. UC 19 vectors are small, high copy number, E. coli plasmids, 2686 bp in length. They are identical except that they contain multiple cloning sites (MCS) arranged in opposite orientations. p. UC 18/19 plasmids contain: (1) the p. MB 1 replicon rep responsible for the replication of plasmid (source – plasmid p. BR 322). The high copy number of p. UC plasmids is a result of the lack of the rop gene and a single point mutation in rep of p. MB 1; (2) bla gene, coding for beta-lactamase that confers resistance to ampicillin (source – plasmid p. BR 322); (3) region of E. coli operon lac containing CAP protein binding site, promoter Plac, lac repressor binding site and 5’-terminal part of the lac. Z gene encoding the N-terminal fragment of beta-galactosidase (source – M 13 mp 18/19). This fragment, whose synthesis can be induced by IPTG, is capable of intra-allelic (alfa) complementation with a defective form of beta-galactosidase encoded by host (mutation lac. ZDM 15). In the presence of IPTG, bacteria synthesize both fragments of the enzyme and form blue colonies on media with X-Gal. Insertion of DNA into the MCS located within the lac. Z gene (codons 6 -7 of lac. Z are replaced by MCS) inactivates the N-terminal fragment of betagalactosidase and abolishes alfa-complementation. Bacteria carrying recombinant plasmids therefore give rise to white colonies.
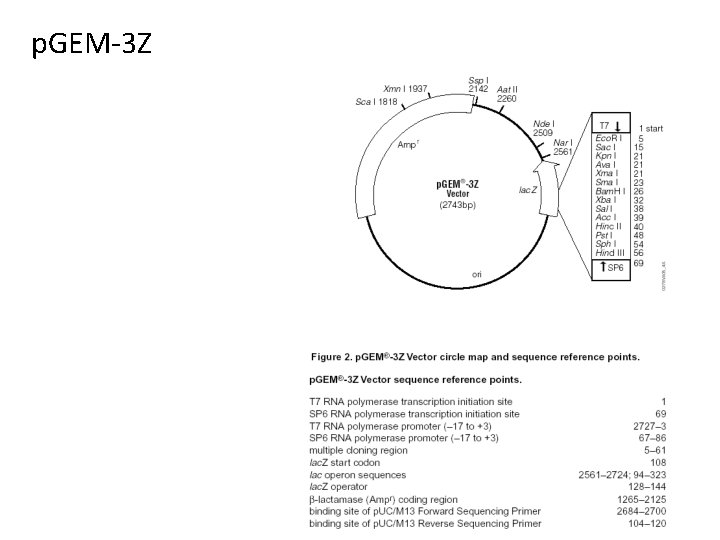
p. GEM-3 Z
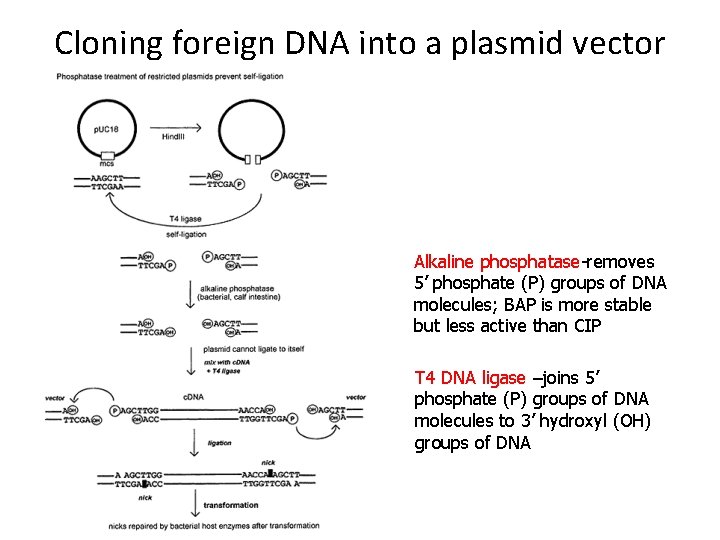
Cloning foreign DNA into a plasmid vector Alkaline phosphatase-removes 5’ phosphate (P) groups of DNA molecules; BAP is more stable but less active than CIP T 4 DNA ligase –joins 5’ phosphate (P) groups of DNA molecules to 3’ hydroxyl (OH) groups of DNA
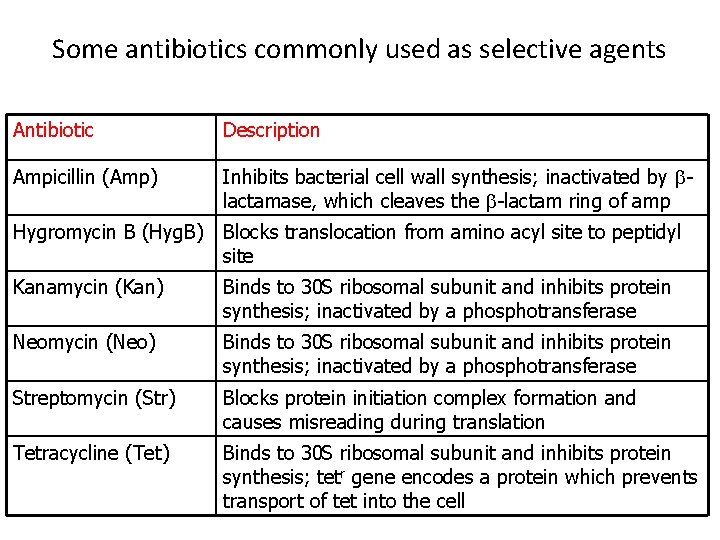
Some antibiotics commonly used as selective agents Antibiotic Description Ampicillin (Amp) Inhibits bacterial cell wall synthesis; inactivated by blactamase, which cleaves the b-lactam ring of amp Hygromycin B (Hyg. B) Blocks translocation from amino acyl site to peptidyl site Kanamycin (Kan) Binds to 30 S ribosomal subunit and inhibits protein synthesis; inactivated by a phosphotransferase Neomycin (Neo) Binds to 30 S ribosomal subunit and inhibits protein synthesis; inactivated by a phosphotransferase Streptomycin (Str) Blocks protein initiation complex formation and causes misreading during translation Tetracycline (Tet) Binds to 30 S ribosomal subunit and inhibits protein synthesis; tetr gene encodes a protein which prevents transport of tet into the cell

Construction and screening of Library • Why required? • What is this library content? • How to find the book from this library if it is huge collection of books?

• Gene libraries • c. DNA libraries • Library screening
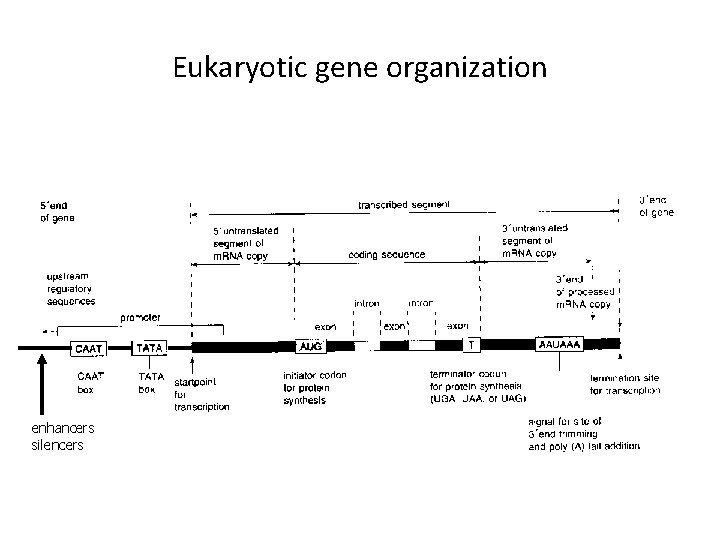
Eukaryotic gene organization enhancers silencers
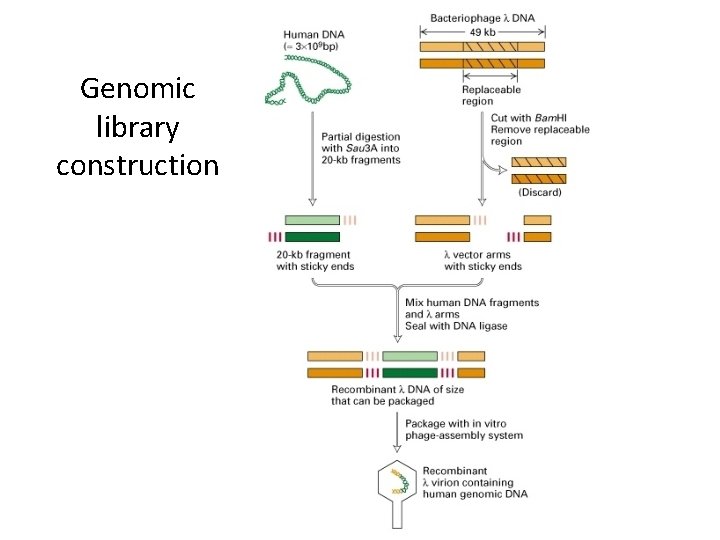
Genomic library construction
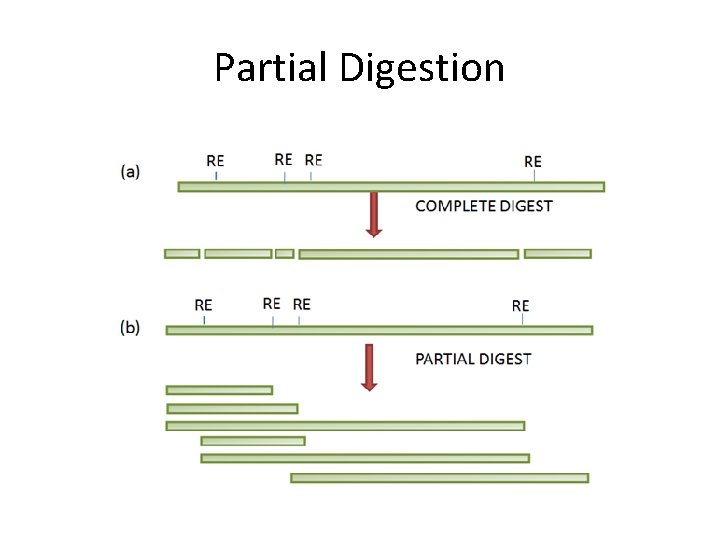
Partial Digestion
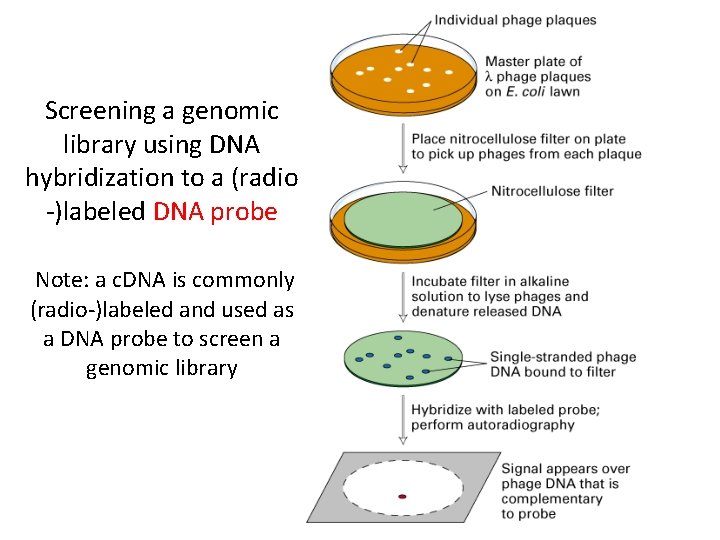
Screening a genomic library using DNA hybridization to a (radio -)labeled DNA probe Note: a c. DNA is commonly (radio-)labeled and used as a DNA probe to screen a genomic library
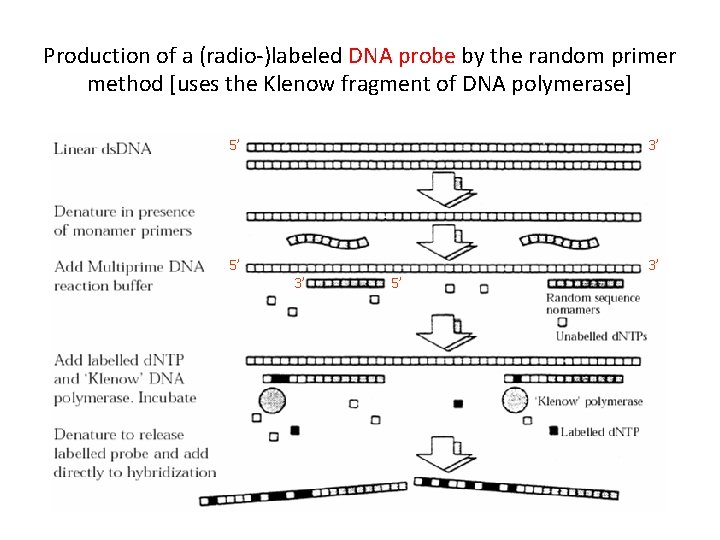
Production of a (radio-)labeled DNA probe by the random primer method [uses the Klenow fragment of DNA polymerase] 5’ 3’ 3’ 5’
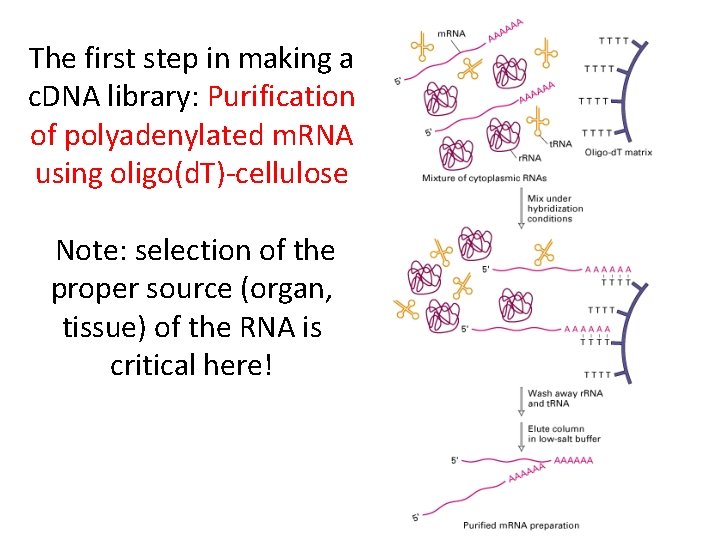
The first step in making a c. DNA library: Purification of polyadenylated m. RNA using oligo(d. T)-cellulose Note: selection of the proper source (organ, tissue) of the RNA is critical here!
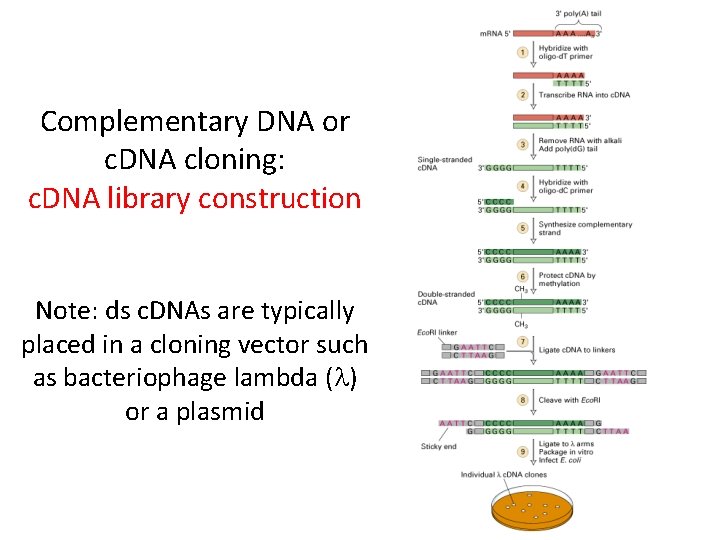
Complementary DNA or c. DNA cloning: c. DNA library construction Note: ds c. DNAs are typically placed in a cloning vector such as bacteriophage lambda (l) or a plasmid
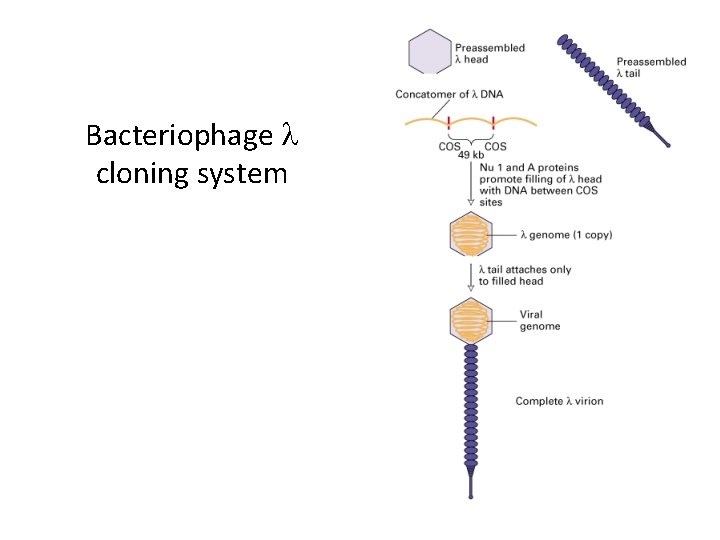
Bacteriophage l cloning system
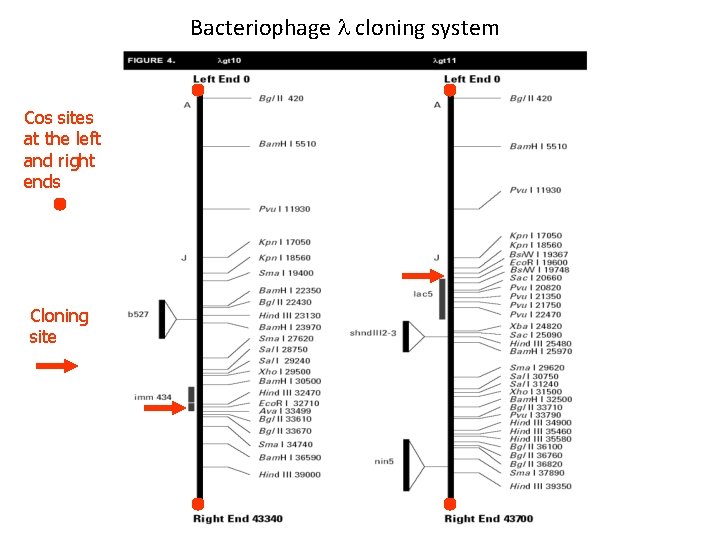
Bacteriophage l cloning system Cos sites at the left and right ends Cloning site
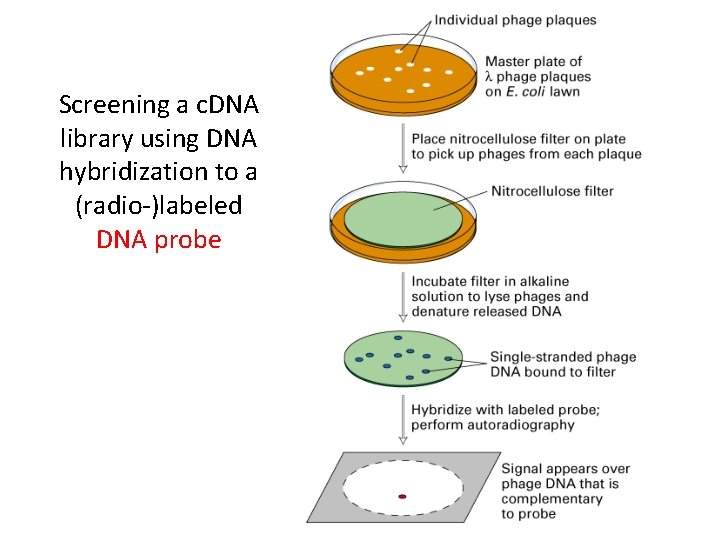
Screening a c. DNA library using DNA hybridization to a (radio-)labeled DNA probe
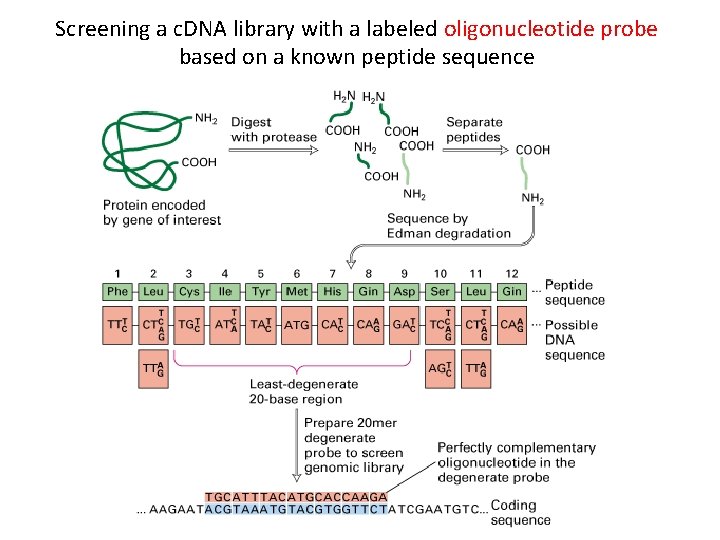
Screening a c. DNA library with a labeled oligonucleotide probe based on a known peptide sequence

Using polynucleotide kinase and g-32 P-labeled ATP to radiolabel oligonucleotide probes
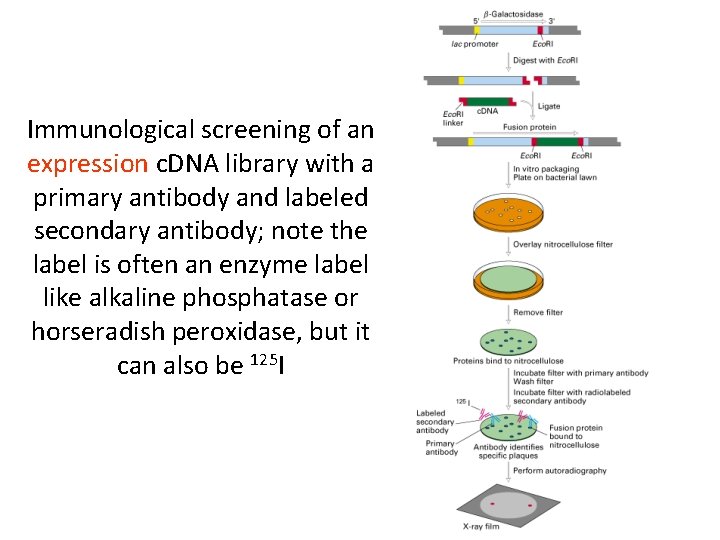
Immunological screening of an expression c. DNA library with a primary antibody and labeled secondary antibody; note the label is often an enzyme label like alkaline phosphatase or horseradish peroxidase, but it can also be 125 I

Animations for two related uses of expression vectors • Expression cloning of receptor proteins-see MCB Chapter 5 • http: //bcs. whfreeman. com/lodish 7 e/#800911__812046__ • Looking for protein-protein interactions with the yeast two hybrid system-see MCB Chapter 7 • http: //bcs. whfreeman. com/lodish 7 e/#800911__812055__
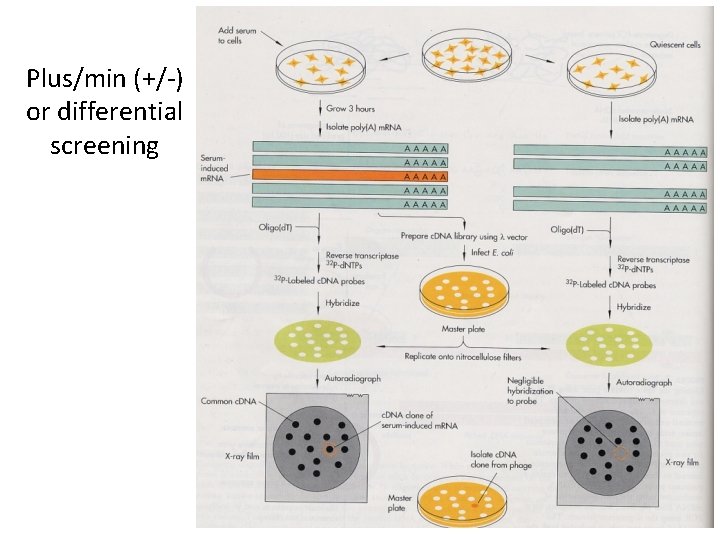
Plus/min (+/-) or differential screening
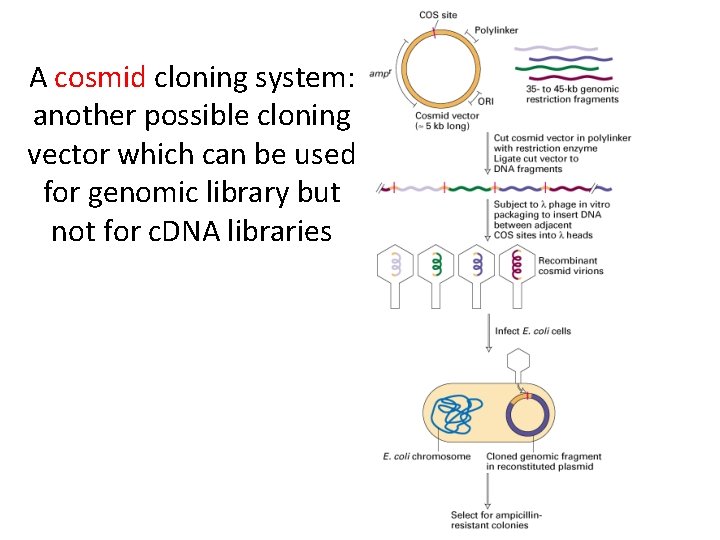
A cosmid cloning system: another possible cloning vector which can be used for genomic library but not for c. DNA libraries

In summary, you have seen: • How to make and screen gene libraries • How to make and screen c. DNA libraries • Several different cloning vectors including plasmids, bacteriophage lambda (l), and cosmids

How you will know the plasmid or DNA you chosen the correct one? • In case of DNA • In case of RNA • In case of Protein
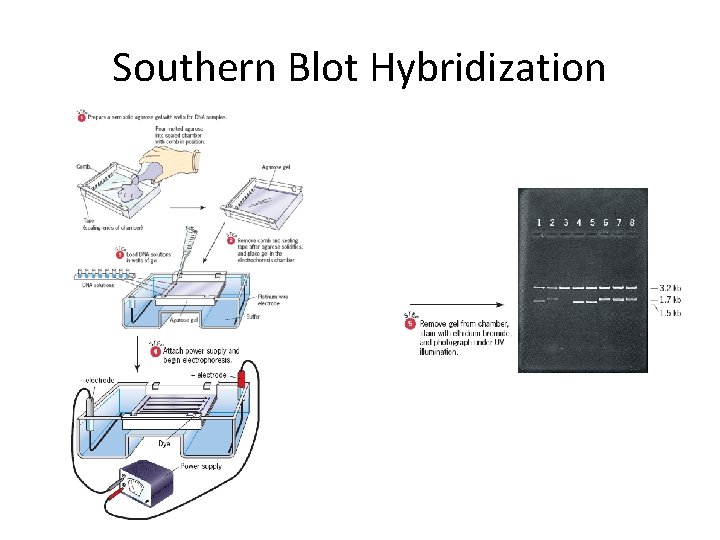
Southern Blot Hybridization
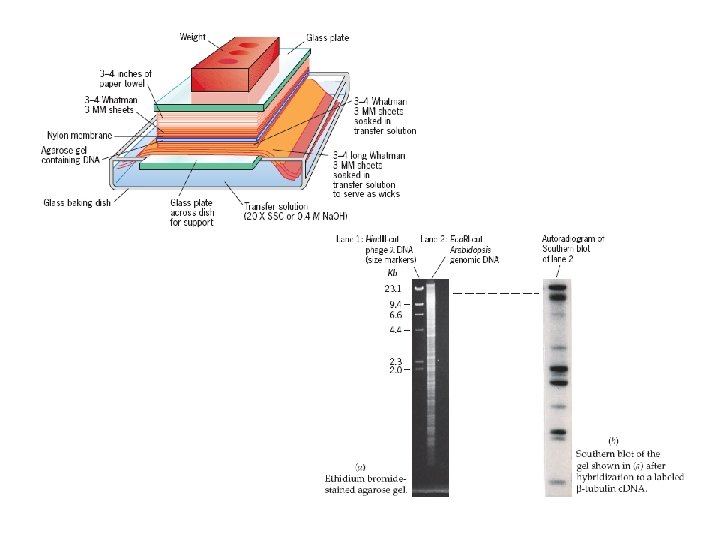

Northern Blot Hybridization • RNA blots are called northern blots in recognition of the fact that the procedure is analogous to the Southern blotting technique, but with RNA molecules being separated and transferred to a membrane.
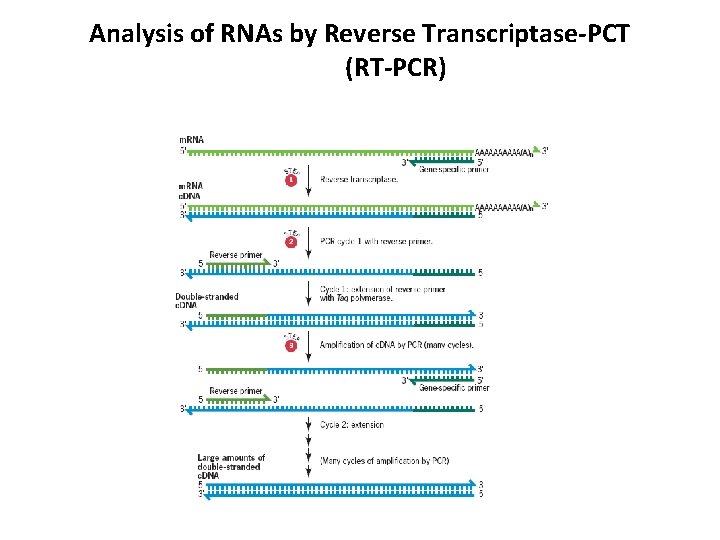
Analysis of RNAs by Reverse Transcriptase-PCT (RT-PCR)
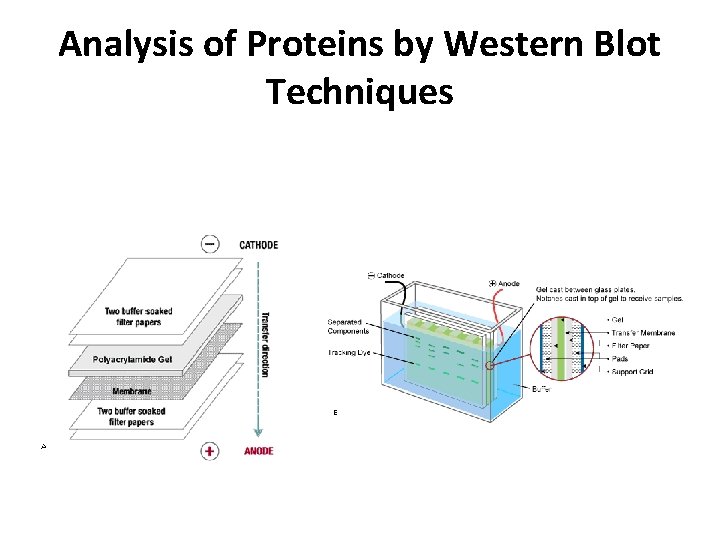
Analysis of Proteins by Western Blot Techniques

END of First Part
- Slides: 96