Molecular Biology Databases NCBI DDBL EMBL and others

Molecular Biology Databases NCBI, DDBL, EMBL and others

What is a Database? • A database can be defined as "a collection of data arranged for ease and speed of search and retrieval. “ • A DNA database contains individual records or data entries of the DNA sequences as well as information about the sequences. • A DNA database often contains flat-files. These are relatively simple database systems in which each database is contained in a single table. • In contrast, relational database systems can use multiple tables to store information, and each table can have a different record format.

Gen. Bank as a Database • Gen. Bank is the National Institute of Health (NIH) genetic sequence database, an annotated collection of all publicly available DNA sequences. • It is maintained by the National Center for Biotechnology Information (NCBI) within the National Institute of Health (NIH).

Anatomy of a Genome Info. System Information structure – Records of hierarchical, complex documents; Tables of rows and columns of numbers, letters, words – Table of contents, Reports, Indexing (as a reference book) – Browse thru available structure. – Search and retrieve according to biological questions – Bulk data selection & retrieval for other uses Information content – Primary: Literature (referenced, abstracted and curated), Sequence and feature analyses, maps, controlled vocabulary/ontologies relevant to biology, people, research methods, contacts, etc. – Metadata describing primary data, along with protocols, notes, sources Informatics / software – “Back-end” database, data collection, management, with some analyses – “Front-end” information services (hypertext web, document search/retrieval methods); ease of understanding and usage (HCI) – “Middleware” glue code, software, etc. – Specialized application for genome data: maps, BLAST searches, ontologies

History of Sequence Databases • The first bioinformatics databases were constructed a few years after the first protein sequences began to become available. • The first protein sequence reported was that of bovine insulin in 1956, consisting of 51 residues. • Nearly a decade later, the first nucleic acid sequence was reported, that of yeast alanine t. RNA with 77 bases. • Just a year later, Dayhoff gathered all the available sequence data to create the first bioinformatic database. • The Protein Data. Bank followed in 1972 with a collection of ten X-ray crystallographic protein structures, and the SWISSPROT protein sequence database began in 1987.

Gen. Bank History • DNA databases began in the early 1980 s with a database called Gen. Bank, which was originated by the U. S. Department of Energy to hold the short stretches of DNA sequence that scientists were just beginning to obtain from a range of organisms. • In the early days of Gen. Bank, rooms of technicians sat at keyboards consisting of only the four letters A, C, T and G, tediously entering the DNA-sequence information published in academic journals.

The National Center for Biotechnology Information • Created as a part of NLM in 1988 – Establish public databases • U. S. National DNA Sequence Database – Perform research in computational biology – Develop software tools for sequence analysis – Disseminate biomedical information

Gen. Bank History • Newer communication technologies enabled researchers to dial up Gen. Bank and dump in their sequence data directly. • The administration of Gen. Bank was transferred to National Institutes of Health's National Center for Biotechnology Information (NCBI). • With the advent of the World Wide Web, researchers could access the data in Gen. Bank for free from around the globe. • Once the Human Genome Project (HGP) began in 1990, DNA-sequence data in Gen. Bank began to grow exponentially. • With the introduction in the 1990 s of high-throughput sequencing additions to Gen. Bank skyrocketed.
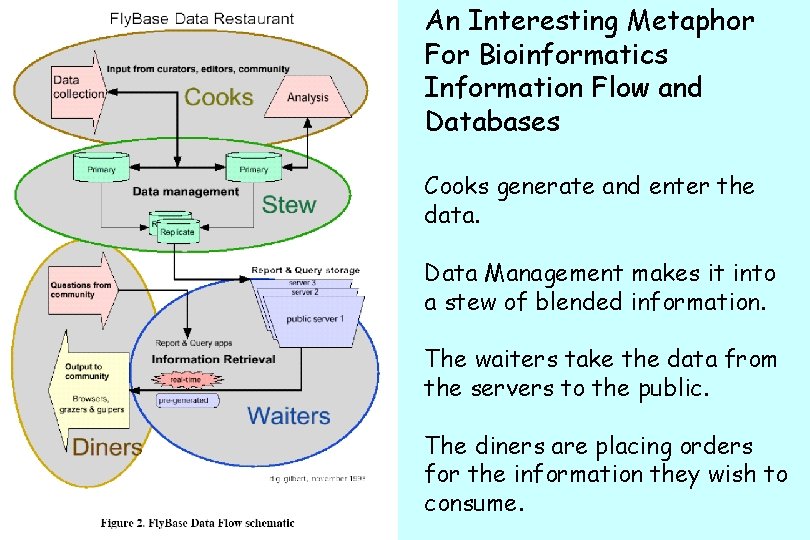
An Interesting Metaphor For Bioinformatics Information Flow and Databases Cooks generate and enter the data. Data Management makes it into a stew of blended information. The waiters take the data from the servers to the public. The diners are placing orders for the information they wish to consume.

Molecular Databases • Primary Databases – Original submissions by experimentalists – Database staff organize but don’t additional information • Example: Gen. Bank, SNP, GEO • Derivative Databases – Human curated • compilation and correction of data • Example: SWISS-PROT, NCBI Ref. Seq m. RNA – Computationally Derived • Example: Uni. Gene – Combinations • Example: NCBI Genome Assembly

What, the scientists submit their own DNA sequences? • Who checks for error? • Who makes people actually send their data to the database so all can share it? • Learn from success, failure of Gen. Bank/EMBL extensive publicly shared bio-data • Carrot/stick approach. Granting agencies and journals began requiring scientists to publish sequence data. Patented sequences must be entered in the databases too. • However, there is significant public databank error due to data ownership by scientists; no inducements to update or go back and correct errors.
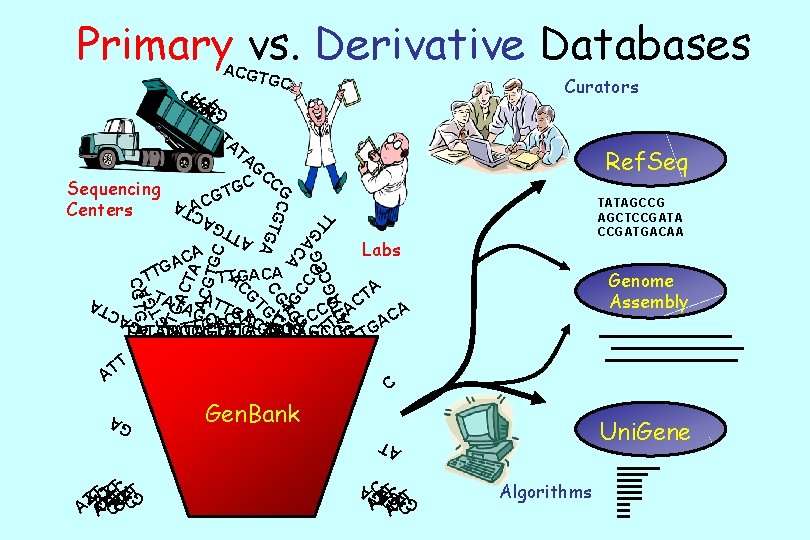
Primary vs. Derivative Databases ACGT GC C TC ATCATCT TA Curators GAG A A AT T A AC T G A AT GTG CTGA CT A ACG TGC TT G A C A AC CG G CC G ATA T G AC Ref. Seq CG TG Sequencing Centers TA G CC C G G TATAGCCG AGCTCCGATA CCGATGACAA Labs Genome Assembly T TA TA GC A TG CG GC T G C Gen. Bank Uni. Gene AT C TC ATCATCT GAG A A AT T A AC T G GA TACTTTCTT T A ATCA C A CGTGA G TT TTGACA A T TA A T C TA CGGA GC TGTAA CA C C A A G T G G TTATAGCCG A TA AT TATAGCCG ATT TATAGCCG TG T A T T AT C Algorithms GAGA GAG A

Gen. Bank is NCBI’s Primary Sequence Database • Nucleotide only sequence database • Archival in nature • Gen. Bank Data – Direct submissions (traditional records ) – Batch submissions (EST, GSS, STS) – ftp accounts (genome data) • Three collaborating databases – Gen. Bank – DNA Database of Japan (DDBJ) – European Molecular Biology Laboratory (EMBL) Database

Why use Bioinformatics Databases? • Speed of information retrieval • Increasing size of data sets • Amount of information available • Save time and money by simulating experiments prior to actual experiment (a. k. a. in silico)

How do you access Databases? • Search engines – Programs that allow you to search the database • Links from other sites to the search engines • Programs that directly link to the search engines

Boolean Logic • Why do we use Boolean operators – To narrow your search – get fewer superfluous results • What are the Boolean Operators – AND-looks for entries with both terms – OR-looks for entries with one term or the other – NOT (or BUTNOT)-looks for entries with one term but not the other – * (Wildcard) -looks for ALL entries that contain the term with the * after it
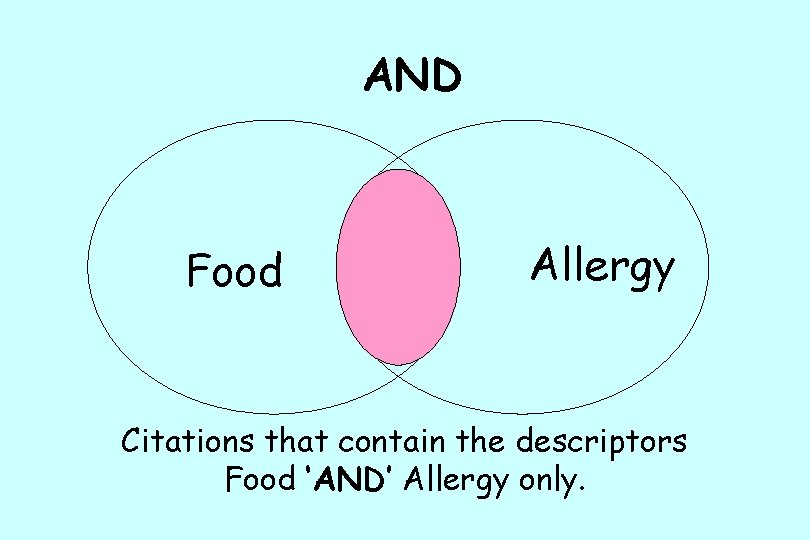
AND Food Allergy Citations that contain the descriptors Food ‘AND’ Allergy only.
OR Food Allergy Citations that contain the descriptors Food ‘OR’ Allergy. This is a bigger set.

NOT Citations that contain the descriptors Allergy ‘NOT’ Food
* (Wildcard) Food Allerg* Citations that contain the descriptors Allerg* (Allergies, Allergy, Allergen

Gen. Bank as a Database • Gen. Bank identifiers are unique combination of numbers and letters used to index Gen. Bank sequence entries. • They can be used to retrieve information about a particular gene or DNA sequence from the Gen. Bank database. • This information also includes links to similar sequence entries and other public databases, making it a relational database as well as a flat file database.

What is Gen. Bank? NCBI’s Primary Sequence Database • Nucleotide only sequence database • Archival in nature • Gen. Bank Data – Direct submissions individual records (Bank. It, Sequin) – Batch submissions via email (EST, GSS, STS) – ftp accounts sequencing centers • Data shared three collaborating databases – Gen. Bank – DNA Database of Japan (DDBJ). – European Molecular Biology Laboratory Database (EMBL) at EBI.
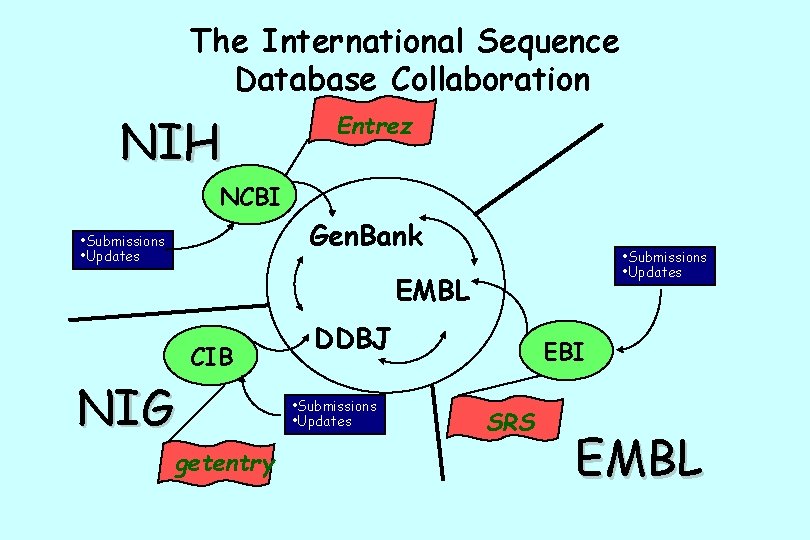
The International Sequence Database Collaboration NIH Entrez NCBI Gen. Bank • Submissions • Updates EMBL NIG CIB getentry DDBJ • Submissions • Updates EBI SRS EMBL

Gen. Bank: NCBI’s Primary Sequence Database Release 131 18, 197, 119 22, 616, 937, 182 110, 000 + August 2002 Records Nucleotides Species • full release every two months • incremental and cumulative updates daily • available only through internet ftp: //ftp. ncbi. nih. gov/genbank/ 83. 65 Gigabytes of data

Gen. Bank: NCBI’s Primary Sequence Database Release 135 24, 027, 936 31, 099, 264, 455 120, 000 + April 2003 Records Nucleotides Species • full release every two months • incremental and cumulative updates daily • available only through internet ftp: //ftp. ncbi. nih. gov/genbank/ 114 Gigabytes

Gen. Bank: NCBI’s Primary Sequence Database Release 139 December 2003 30, 968, 418 36, 553, 368, 485 >140, 000 Records Nucleotides Species 138 Gigabytes 570 files • full release every two months • incremental and cumulative updates daily • available only through internet ftp: //ftp. ncbi. nih. gov/genbank/
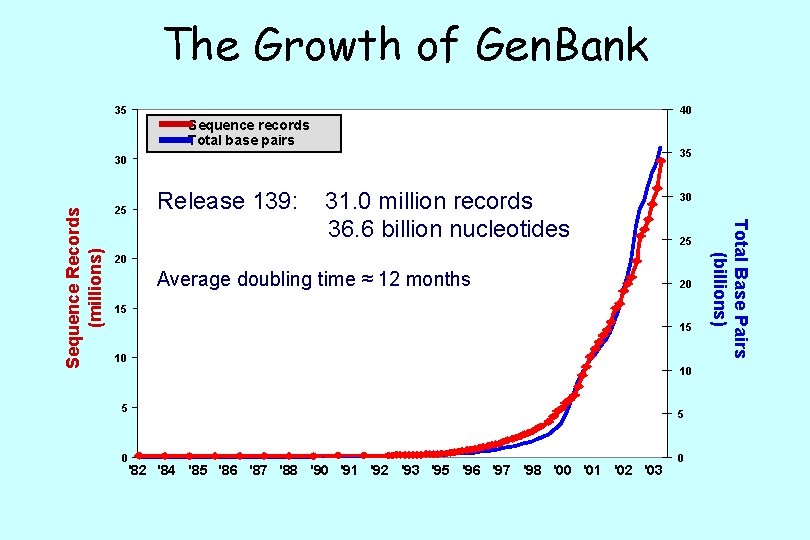
The Growth of Gen. Bank 35 40 Sequence records Total base pairs 35 Release 139: 25 31. 0 million records 36. 6 billion nucleotides 30 25 20 Average doubling time ≈ 12 months 20 15 15 10 10 5 0 '82 '84 '85 '86 '87 '88 '90 '91 '92 '93 '95 '96 '97 '98 '00 '01 '02 '03 5 0 Total Base Pairs (billions) Sequence Records (millions) 30
The Entrez System

Entrez Nucleotides Primary • Gen. Bank / EMBL / DDBJ 35, 116, 960 Derivative • Ref. Seq • Third Party Annotation • PDB Total 259, 219 3, 182 4, 703 35, 384, 248
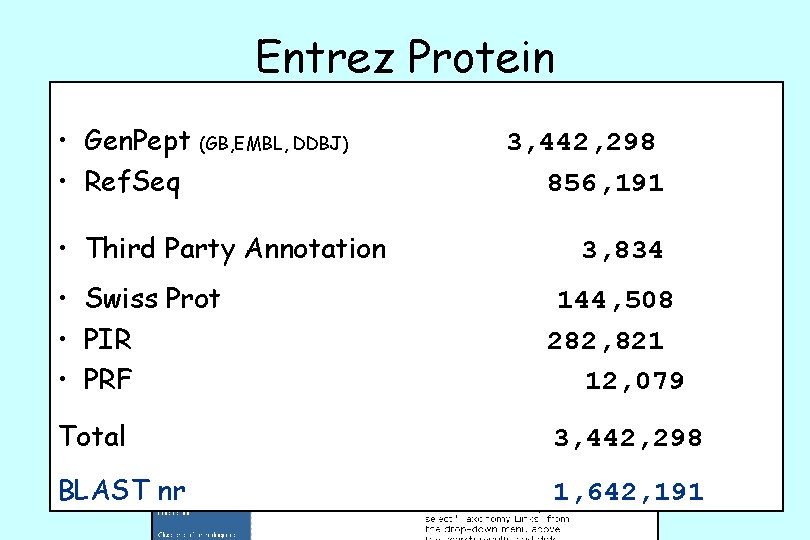
Entrez Protein • Gen. Pept (GB, EMBL, DDBJ) • Ref. Seq • Third Party Annotation 3, 442, 298 856, 191 3, 834 • Swiss Prot • PIR • PRF 144, 508 282, 821 12, 079 Total 3, 442, 298 BLAST nr 1, 642, 191

Organization of Gen. Bank: Gen. Bank Divisions Records are divided into 17 Divisions. 1 Patent (11 files) 5 High Throughput 11 Traditional Divisions: BULK Divisions: • Direct Batch Submissions Submission (Sequin and. FTP) Bank. It) (Email and • Accurate Inaccurate • Well characterized Poorly characterized EST (288) Expressed Sequence Tag PRI (27) Primate GSS (98) Genome Survey Sequence PLN (10) Plant and Fungal HTG (61) High Throughput Genomic BCT (8) Bacterial and Archeal STS (3) Sequence Tagged Site INV (6) Invertebrate HTC (3) High Throughput c. DNA ROD (11) Rodent VRL (3) Viral VRT (4) Other Vertebrate MAM (1) Mammalian (ex. ROD and PRI) PHG (1) Phage SYN (1) Synthetic (cloning vectors) UNA (1) Unannotated Entrez query: gbdiv_xxx[Properties]

Traditional Gen. Bank Divisions • Direct Submissions (Sequin and Bank. It) • Accurate • Well characterized BCT INV MAM PHG PLN PRI ROD SYN VRL VRT Bacterial and Archeal Invertebrate Mammalian (ex. ROD and PRI) Phage Plant and Fungal Primate Rodent Synthetic (cloning vectors) Viral Other Vertebrate

A Helpful Resource • This is a link to a sample annotated Gen. Bank Record. Click on any of the underlined links to learn more about the file structure. • http: //www. ncbi. nlm. nih. gov/Sitemap/sampler ecord. html

What is an Accession Number? • An accession number is label that used to identify a sequence in the various databases. It is a string of letters and/or numbers that corresponds to a molecular sequence. • Examples (all for retinol-binding protein, RBP 4): – – – – – X 02775 NT_030059 Rs 7079946 N 91759. 1 NM_006744 NP_007635 AAC 02945 Q 28369 1 KT 7 Gen. Bank genomic DNA sequence Genomic contig db. SNP (single nucleotide polymorphism) An expressed sequence tag (1 of 170) Ref. Seq DNA sequence (from a transcript) Ref. Seq protein Gen. Bank protein Swiss. Prot protein Protein Data Bank structure record
Gen. Bank Flat File Format • When you click on an entry, you have opened a Gen. Bank Flat File • Information includes: – The Name of the gene – The Accession number – Journal articles

Gen. Bank Flat File Format • Information (Cont) – Structural information of the gene (eg intron/exon boundaries, promoters, etc) – The code for the protein – The code for the DNA (RNA-if m. RNA it is the c. DNA for the m. RNA sequenced)
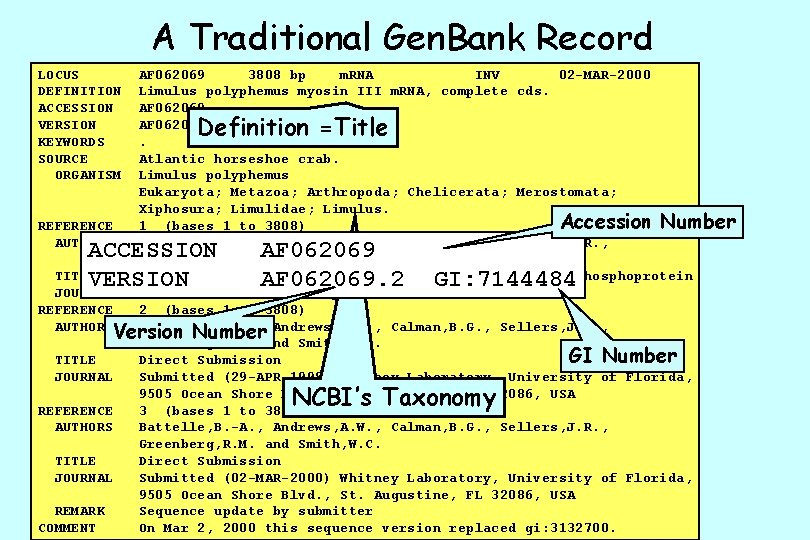
A Traditional Gen. Bank Record LOCUS DEFINITION ACCESSION VERSION KEYWORDS SOURCE ORGANISM AF 062069 3808 bp m. RNA INV 02 -MAR-2000 Limulus polyphemus myosin III m. RNA, complete cds. AF 062069. 2 GI: 7144484. Atlantic horseshoe crab. Limulus polyphemus Eukaryota; Metazoa; Arthropoda; Chelicerata; Merostomata; Xiphosura; Limulidae; Limulus. Accession Number REFERENCE 1 (bases 1 to 3808) AUTHORS Battelle, B. -A. , Andrews, A. W. , Calman, B. G. , Sellers, J. R. , Greenberg, R. M. and Smith, W. C. TITLE A myosin III from Limulus eyes is a clock-regulated phosphoprotein JOURNAL J. Neurosci. (1998) In press REFERENCE 2 (bases 1 to 3808) AUTHORS Battelle, B. -A. , Andrews, A. W. , Calman, B. G. , Sellers, J. R. , Version Number Greenberg, R. M. and Smith, W. C. GI Number TITLE Direct Submission JOURNAL Submitted (29 -APR-1998) Whitney Laboratory, University of Florida, 9505 Ocean Shore Blvd. , St. Augustine, FL 32086, USA REFERENCE 3 (bases 1 to 3808) AUTHORS Battelle, B. -A. , Andrews, A. W. , Calman, B. G. , Sellers, J. R. , Greenberg, R. M. and Smith, W. C. TITLE Direct Submission JOURNAL Submitted (02 -MAR-2000) Whitney Laboratory, University of Florida, 9505 Ocean Shore Blvd. , St. Augustine, FL 32086, USA REMARK Sequence update by submitter COMMENT On Mar 2, 2000 this sequence version replaced gi: 3132700. Definition =Title ACCESSION VERSION AF 062069. 2 GI: 7144484 NCBI’s Taxonomy

Gen. Bank Record: Feature Table FEATURES source CDS Location/Qualifiers 1. . 3808 /organism="Limulus polyphemus" /db_xref="taxon: 6850" /tissue_type="lateral eye" 258. . 3302 /note="N-terminal protein kinase domain; C-terminal myosin heavy chain head; substrate for PKA" Gen. Pept Protein IDS /codon_start=1 /product="myosin III" /protein_id="AAC 16332. 2" /db_xref="GI: 7144485" /translation="MEYKCISEHLPFETLPDPGDRFEVQELVGTGTYATVYSAIDKQA NKKVALKIIGHIAENLLDIETEYRIYKAVNGIQFFPEFRGAFFKRGERESDNEVWLGI EFLEEGTAADLLATHRRFGIHLKEDLIALIIKEVVRAVQYLHENSIIHRDIRAANIMF SKEGYVKLIDFGLSASVKNTNGKAQSSVGSPYWMAPEVISCDCLQEPYNYTCDVWSIG ITAIELADTVPSLSDIHALRAMFRINRNPPPSVKRETRWSETLKDFISECLVKNPEYR PCIQEIPQHPFLAQVEGKEDQLRSELVDILKKNPGEKLRNKPYNVTFKNGHLKTISGQ 1201 a 689 c 782 g 1136 t /protein_id="AAC 16332. 2" /db_xref="GI: 7144485" BASE COUNT ORIGIN 1 tcgacatctg tggtcgcttt ttttagtaat aaaaaattgt attatgacgt cctatctgtt 3781 aagatacagt aactagggaa aaaa //

Multiple Formats are available for Sequence Data • Historically, all the DNA and Protein software was written concurrent with the establishment of the databases. So the formats needed in the databases and the software co-evolved. • Sequence analysis software needs simpler formats than databases for speed- or else the program must be allowed to ignore most of the excess information.
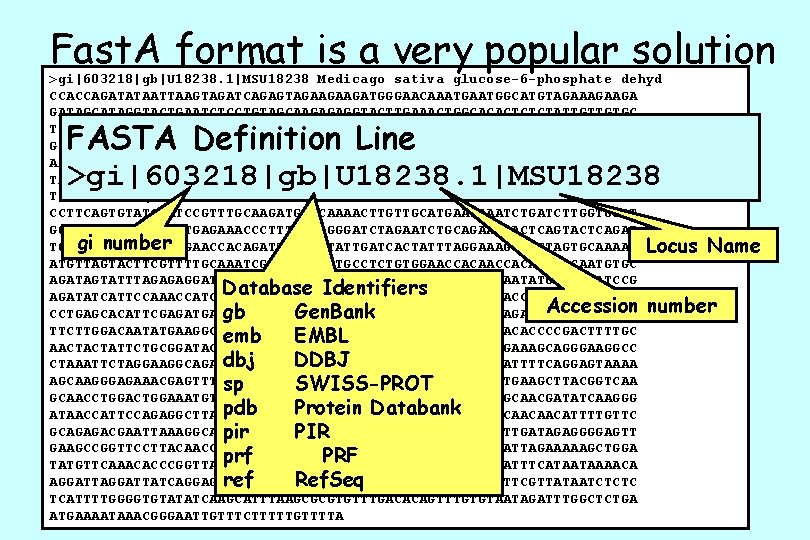
Fast. A format is a very popular solution >gi|603218|gb|U 18238. 1|MSU 18238 Medicago sativa glucose-6 -phosphate dehyd CCACCAGATATAATTAAGTAGATCAGAGTAGAAGAAGATGGGAACAAATGGCATGTAGAAGA GATAGCATAGGTACTGAATCTCCTGTAGCAAGAGAGGTACTTGAAACTGGCACACTCTCTATTGTTGTGC TTGGTGCTTCTGGTGATCTTGCCAAGAAGAAGACTTTTCCTGCACTTTTTCACTTATATAAACAGGAATT GTTGCCACCTGATGAAGTTCACATTTTTGGCTATGCAAGGTCAAAGATCTCCGATGATGAATTGAGAAAC AAATTGCGTAGCTATCTTGTTCCAGAGAAAGGTGCTTCTCCTAAACAGTTAGATGATGTATCAAAGTTTT TACAATTGGTTAAATATGTAAGTGGCCCTTATGATTCTGAAGATGGATTTCGCTTGTTGGATAAAGAGAT TTCAGAGCATGAATATTTGAAAAATAGTAAAGAGGGTTCATCTCGGAGGCTTTTCTATCTTGCACTTCCT > CCTTCAGTGTATCCGTTTGCAAGATGATCAAAACTTGTTGCATGAATAAATCTGATCTTGGTGGAT GGACACGCGTTGTTGTTGAGAAACCCTTTGGTAGGGATCTAGAATCTGCAGAAGAACTCAGTACTCAGAT gi number TGGAGAGTTATTTGAAGAACCACAGATTTATCGTATTGATCACTATTTAGGAACTAGTGCAAAAC Locus Name ATGTTAGTACTTCGTTTTGCAAATCGGTTCTTCTTGCCTCTGTGGAACCACATTGACAATGTGC AGATAGTATTTAGAGAGGATTTTGGAACTGATGGTCGTGGTGGATATTTTGACCAATATGGAATTATCCG Database Identifiers AGATATCATTCCAAACCATCTGTTGCAGGTTCTTTGCTTGATTGCTATGGAAAAACCCGTTTCTCTCAAG Accession number gb Gen. Bank CCTGAGCACATTCGAGATGAGAAAGTGAAGGTTCTTGAATCAGTACTCCCTATTAGAGATGATGAAGTTG TTCTTGGACAATATGAAGGCTATACAGATGACCCAACTGTACCGGACGATTCAAACACCCCGACTTTTGC emb EMBL AACTACTATTCTGCGGATACACAATGAAAGATGGGAAGGTGTTCCTTTCATTGTGAAAGCAGGGAAGGCC dbj DDBJ CTAAATTCTAGGAAGGCAGAGATTCGGGTTCAAGGATGTTCCTGGTGACATTTTCAGGAGTAAAA AGCAAGGGAGAAACGAGTTTGTTATCCGCCTACAACCTTCAGAAGCTATTTACATGAAGCTTACGGTCAA sp SWISS-PROT GCAACCTGGAAATGTCTGCAGTTCAAAGTGAACTAGACTTGTCATATGGGCAACGATATCAAGGG pdb Protein Databank ATAACCATTCCAGAGGCTTATGAGCGTCTAATTCTCGACACAATTAGAGGTGATCAACAACATTTTGTTC GCAGAGACGAATTAAAGGCATCATGGCAAATATTCACACCACTTTTACACAAAATTGATAGAGGGGAGTT pir PIR GAAGCCGGTTCCTTACAACCCGGGAAGTAGAGGTCCTGCAGAAGCAGATGAGTTATTAGAAAAAGCTGGA prf PRF TATGTTCAAACACCCGGTTATATATGGATTCCTCCTACCTTATAGAGTGACCAAATTTCATAATAAAACA ref Ref. Seq AGGATTATCAGGAGCTTATAAGTCTTCAATAAGCTTGTGAAATTTTCGTTATAATCTCTC TCATTTTGGGGTGTATATCAAGCATTTAAGCGCGTGTTTGACACAGTTTGTGTAATAGATTTGGCTCTGA ATGAAAATAAACGGGAATTGTTTCTTTTTGTTTTA FASTA Definition Line >gi|603218|gb|U 18238. 1|MSU 18238
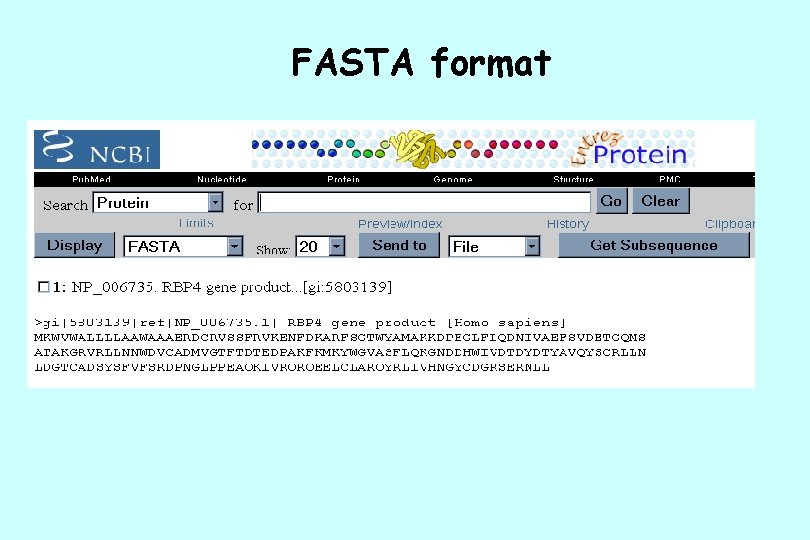
FASTA format
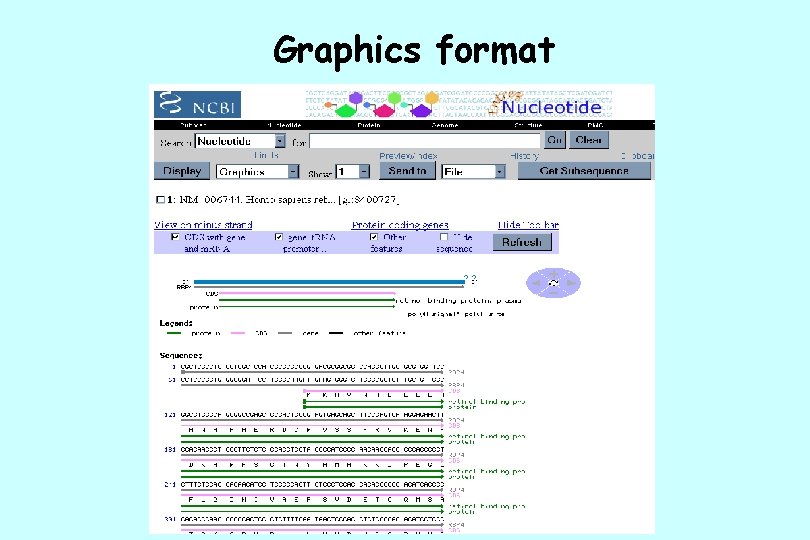
Graphics format

ASN. 1 Format • ASN. 1, or Abstract Syntax Notation One, is an International Standards Organization (ISO) data representation format used to achieve interoperability between platforms. • NCBI uses ASN. 1 for the storage and retrieval of data such as nucleotide and protein sequences, structures, genomes, and MEDLINE records. • ASN. 1 permits computers and software systems of all types to reliably exchange both the data structure and content.

NCBI Software Development Tool Kit • The "NCBI Toolbox" is a set of software and data exchange specifications used by NCBI to produce portable, modular software for molecular biology. • The software in the Toolbox is primarily designed to read ASN. 1 format records. • It is available to the public in the toolbox/ncbi_tools directory of NCBI's ftp site, and can be used in its own right or as a foundation for building tools with similar properties. • The readme files in the toolbox and toolbox/ncbi_tools directories of the FTP site contain more information about the toolbox and ASN. 1.
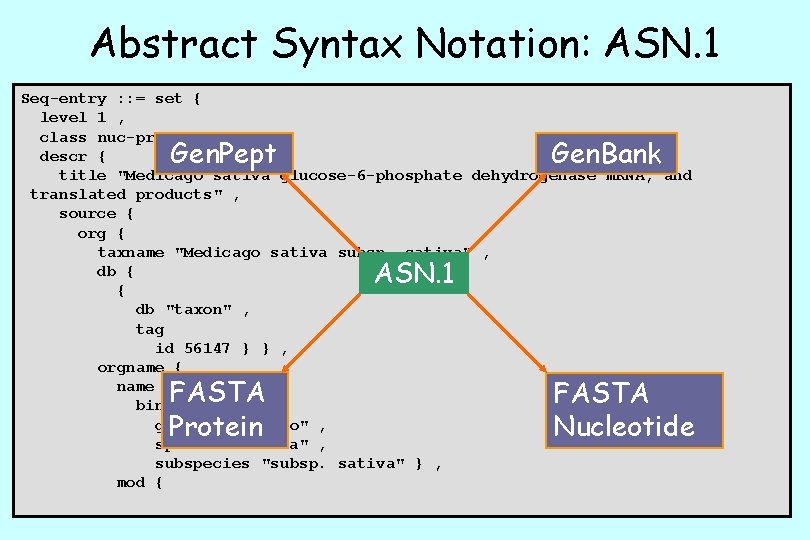
Abstract Syntax Notation: ASN. 1 Seq-entry : : = set { level 1 , class nuc-prot , descr { title "Medicago sativa glucose-6 -phosphate dehydrogenase m. RNA, and translated products" , source { org { taxname "Medicago sativa subsp. sativa" , db { { db "taxon" , tag id 56147 } } , orgname { name binomial { genus "Medicago" , species "sativa" , subspecies "subsp. sativa" } , mod { Gen. Pept Gen. Bank ASN. 1 FASTA Protein FASTA Nucleotide

NCBI Toolbox /************************************ * * asn 2 ff. c * convert an ASN. 1 entry to flat file format, using the FFPrint. Array. * *************************************/ #include <accentr. h> #include "asn 2 ff. h" #include "asn 2 ffp. h" #include "ffprint. h" #include <subutil. h> #include <objall. h> #include <objcode. h> #include <lsqfetch. h> #include <explore. h> Toolbox Sources ftp> open ftp. ncbi. nih. gov. . #ifdef ENABLE_ID 1 #include <accid 1. h>ftp> cd toolbox #endif ftp> cd ncbi_tools FILE *fpl; ftp: //ftp. ncbi. nlm. gov/toolbox/ncbi_tools Args myargs[] = { {"Filename for asn. 1 input", "stdin", NULL, TRUE, 'a', ARG_FILE_IN, 0. 0, 0, NULL}, {"Input is a Seq-entry", "F", NULL , TRUE, 'e', ARG_BOOLEAN, 0. 0, 0, NULL}, {"Input asnfile in binary mode", "F", NULL, TRUE, 'b', ARG_BOOLEAN, 0. 0, 0, NULL}, {"Output Filename", "stdout", NULL, TRUE, 'o', ARG_FILE_OUT, 0. 0, 0, NULL}, {"Show Sequence? ", "T", NULL , TRUE, 'h', ARG_BOOLEAN, 0. 0, 0, NULL},

Database Tools aren’t keeping pace • Despite the huge progress in sequencing and expression analysis technologies, and the corresponding magnitude of more data that is held in the public, private and commercial databases, the tools used for storage, retrieval, analysis and dissemination of data in bioinformatics are still very similar to the original systems gathered together by researchers 15 -20 years ago. • Many are simple extensions of the original academic systems, which have served the needs of both academic and commercial users for many years. • These systems are now beginning to fall behind as they struggle to keep up with the pace of change in the pharma industry.

Database Tools aren’t keeping pace • Databases are still gathered, organized, disseminated and searched using flat files. • Relational databases are still few and far between, and object-relational or fully object oriented systems are rarer still in mainstream applications. • Interfaces still rely on command lines, fat client interfaces, which must be installed on every desktop, or HTML/CGI forms. • Whilst they were in the hands of bioinformatics specialists, pharmas have been relatively undemanding of their tools. • Now the problems have expanded to cover the mainstream discovery process, much more flexible and scalable solutions are needed to serve pharma R&D informatics requirements.

There are more than one type of DNA sequence in Genebank • Genomic sequences made from genomic DNA- these do contain introns and LOTS of DNA that never becomes messenger RNA. m. RNA codes for proteins. • c. DNA sequences made from m. RNA- these don’t contain the introns • ESTS (short stretches of c. DNA sequences that are sort of a “rough draft” • mt. DNA from mitochondrial genomes • SNP single nucleotide polymorphisms with some DNA variation.

Quality of the Sequence is Variable • Some of the DNA is sequenced several times before it is added to the databases. • Some of the DNA is sequenced very quickly on automated equipment and is input directly from the computers. • Both are important types of information. • The “draft” is corrected by curators who assemble the pieces into the genome.
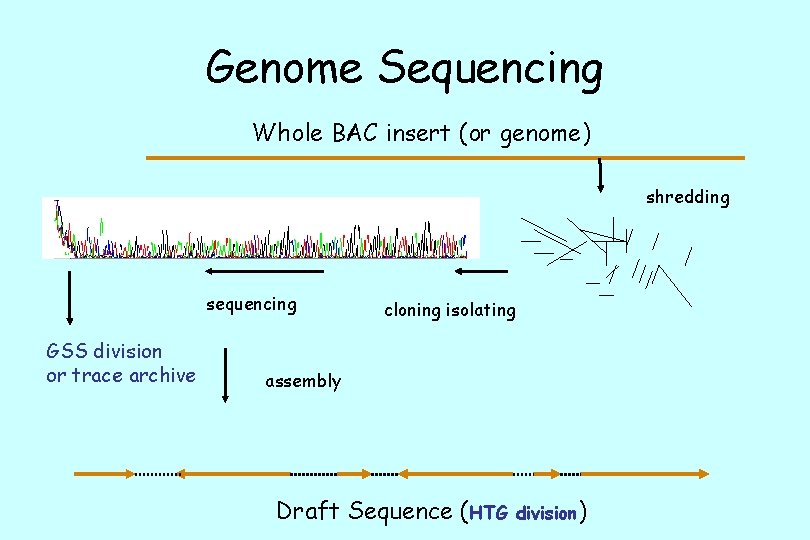
Genome Sequencing Whole BAC insert (or genome) shredding sequencing GSS division or trace archive cloning isolating assembly Draft Sequence (HTG division)
Working Draft Sequence gaps

Assembly Required. • All the data is still in the pieces used to assemble the genomes. • So, that means all the overlapping pieces are still in the databases. • So, searching comes up with many versions and shorter subclones: pieces which are used to assemble the “genomic contigs” or contiguous pieces which are assembled into whole chromosomes. • Sometimes you want to use the smaller pieces, since handling the whole chromosome is awkward in sequence analysis.
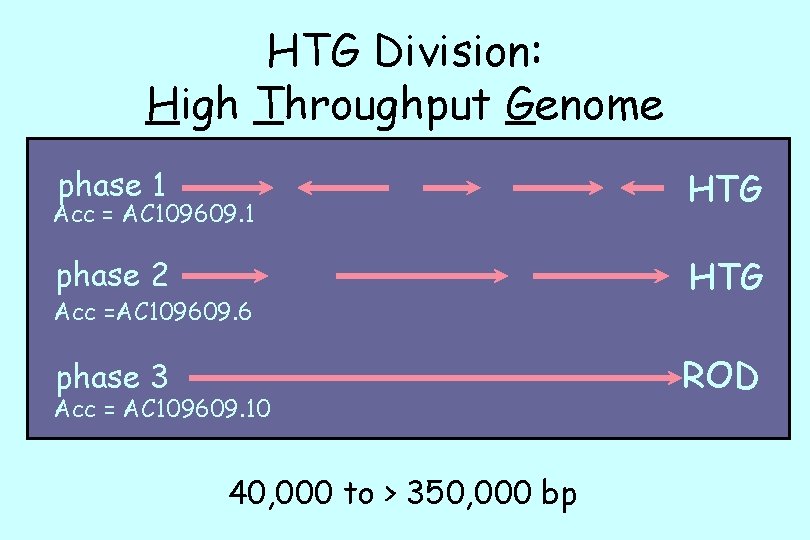
HTG Division: High Throughput Genome phase 1 HTG phase 2 HTG phase 3 ROD Acc = AC 109609. 1 Acc =AC 109609. 6 Acc = AC 109609. 10 40, 000 to > 350, 000 bp

HTG Division: High Throughput Genome 40, 000 to > 350, 000 bp

Whole Genome Shotgun

STS Division : Sequence Tagged Sites • Segment of gene, EST , m. RNA or genomic DNA of known position (microsatellite) • PCR with STS primers gives one product per genome • Basis of Radiation Hybrid Mapping – Uni. Gene – Genome Assembly • Related resource: Electronic PCR http: //www. ncbi. nlm. nih. gov/genome/sts/epcr. cgi

Be aware of errors in the databases Sequence errors: • genome projects’ error rate is 1/10, 000 nucleotides; • ESTs’ error rate is 1/100 nucleotides. Annotation errors: • Many databases annotate their sequences using automated computer programs. These programs do not always give correct annotations. • Swiss. Prot is a protein database curated annotated manually by biologists. It’s regarded as the most reliable database, but does not have the most up-to-date sequence information.

There is a Lot of Sequence in the Databases • One problem is finding what you are looking for in the database. • Try putting in the search term human beta hemoglobin into the nucleotide database. It won’t be easy to find the sequence in the 88 pages of hits! • Ref. Seq was invented to help you find some of the common sequences based on a human (or now, a computer) looking over all the similar submissions of the same sequence to the database. • Ref. Seq corrects some of those sequence errors by comparing lots of sequences.

Ref. Seq: NCBI’s Derivative Sequence Database • Curated transcripts and proteins – reviewed – human, mouse, rat, fruit fly, zebrafish, arabidopsis, C. elegans • Human model transcripts and proteins • Assembled Genomic Regions (contigs) – draft human genome – mouse genome • Chromosome records – microbial – organelle

Ref. Seq Benefits • non-redundancy • explicitly linked nucleotide and protein sequences • updates to reflect current sequence data and biology • data validation • format consistency • distinct accession series • stewardship by NCBI staff and collaborators
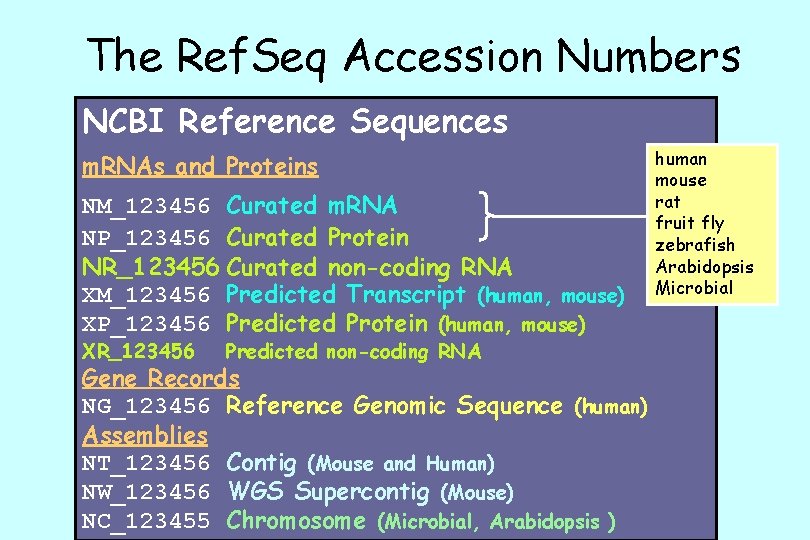
The Ref. Seq Accession Numbers NCBI Reference Sequences m. RNAs and Proteins NM_123456 Curated m. RNA NP_123456 Curated Protein NR_123456 Curated non-coding RNA XM_123456 Predicted Transcript (human, mouse) XP_123456 Predicted Protein (human, mouse) XR_123456 Predicted non-coding RNA Gene Records NG_123456 Reference Genomic Sequence (human) Assemblies NT_123456 Contig (Mouse and Human) NW_123456 WGS Supercontig (Mouse) NC_123455 Chromosome (Microbial, Arabidopsis ) human mouse rat fruit fly zebrafish Arabidopsis Microbial

Gen. Bank Sequences: Lipase Human Lipoprotein
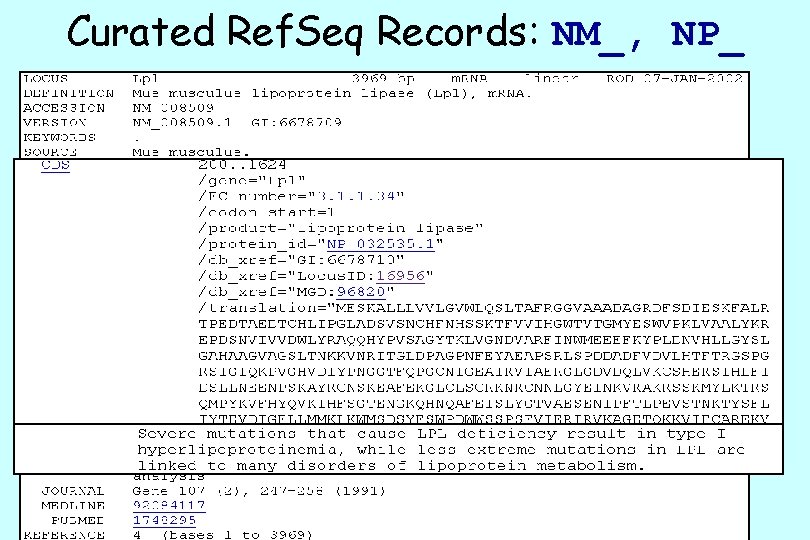
Curated Ref. Seq Records: NM_, NP_

Alignment Based Models
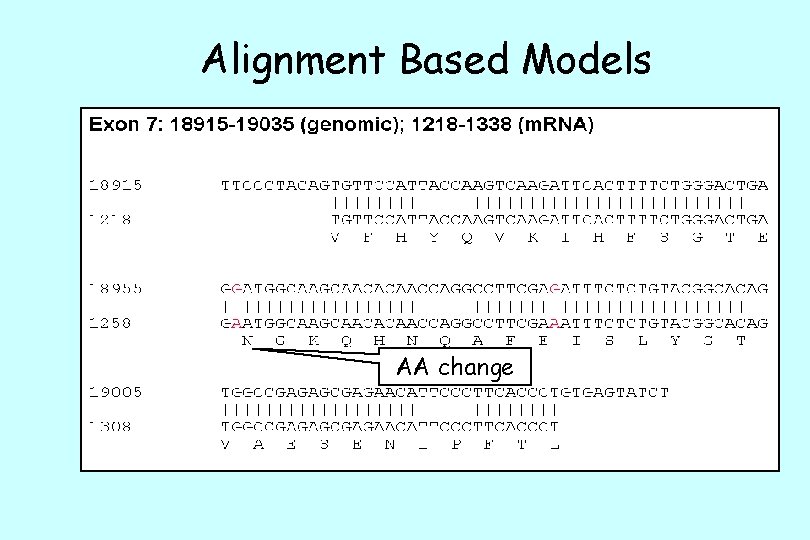
Alignment Based Models AA change
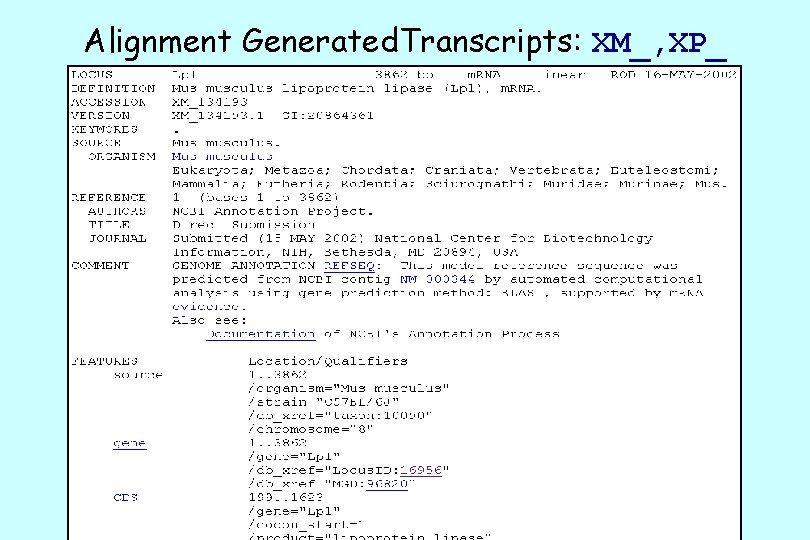
Alignment Generated. Transcripts: XM_, XP_

Ref. Seq Contig: NT_, NW_
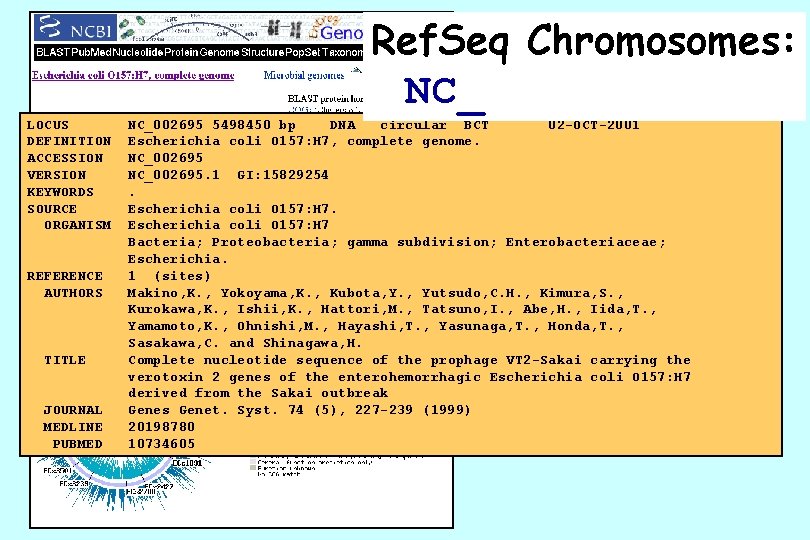
Ref. Seq Chromosomes: NC_ LOCUS DEFINITION ACCESSION VERSION KEYWORDS SOURCE ORGANISM REFERENCE AUTHORS TITLE JOURNAL MEDLINE PUBMED NC_002695 5498450 bp DNA circular BCT 02 -OCT-2001 Escherichia coli O 157: H 7, complete genome. NC_002695. 1 GI: 15829254. Escherichia coli O 157: H 7 Bacteria; Proteobacteria; gamma subdivision; Enterobacteriaceae; Escherichia. 1 (sites) Makino, K. , Yokoyama, K. , Kubota, Y. , Yutsudo, C. H. , Kimura, S. , Kurokawa, K. , Ishii, K. , Hattori, M. , Tatsuno, I. , Abe, H. , Iida, T. , Yamamoto, K. , Ohnishi, M. , Hayashi, T. , Yasunaga, T. , Honda, T. , Sasakawa, C. and Shinagawa, H. Complete nucleotide sequence of the prophage VT 2 -Sakai carrying the verotoxin 2 genes of the enterohemorrhagic Escherichia coli O 157: H 7 derived from the Sakai outbreak Genes Genet. Syst. 74 (5), 227 -239 (1999) 20198780 10734605
Integrated WWW Access: BLAST and Entrez
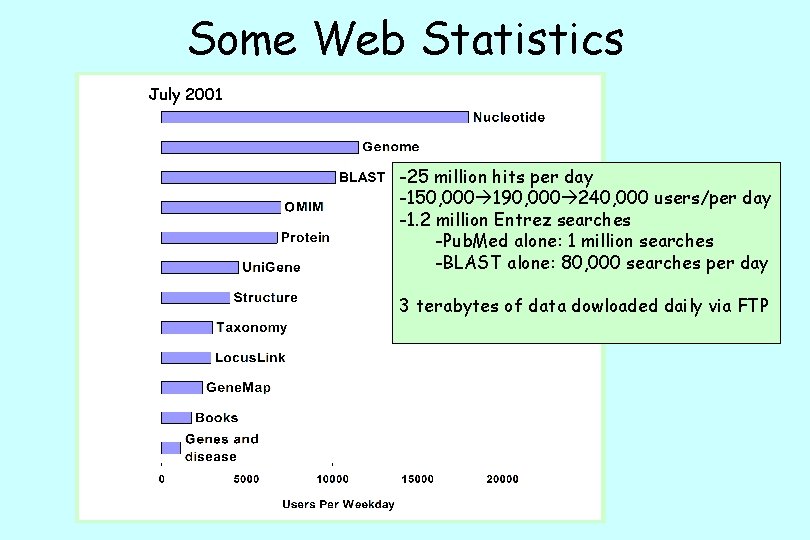
Some Web Statistics July 2001 -25 million hits per day -150, 000 190, 000 240, 000 users/per day -1. 2 million Entrez searches -Pub. Med alone: 1 million searches -BLAST alone: 80, 000 searches per day 3 terabytes of data dowloaded daily via FTP
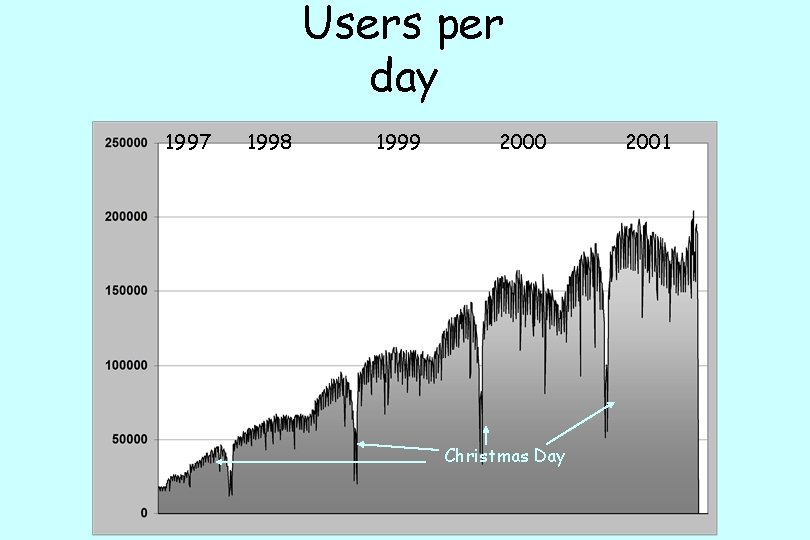
Users per day 1997 1998 1999 2000 Christmas Day 2001

Bulk Gen. Bank Divisions • Batch Submission and htg (email and ftp) • Inaccurate • Poorly Characterized EST STS GSS HTG Expressed Sequence Tagged Site Genome Survey Sequence High Throughput Genomic
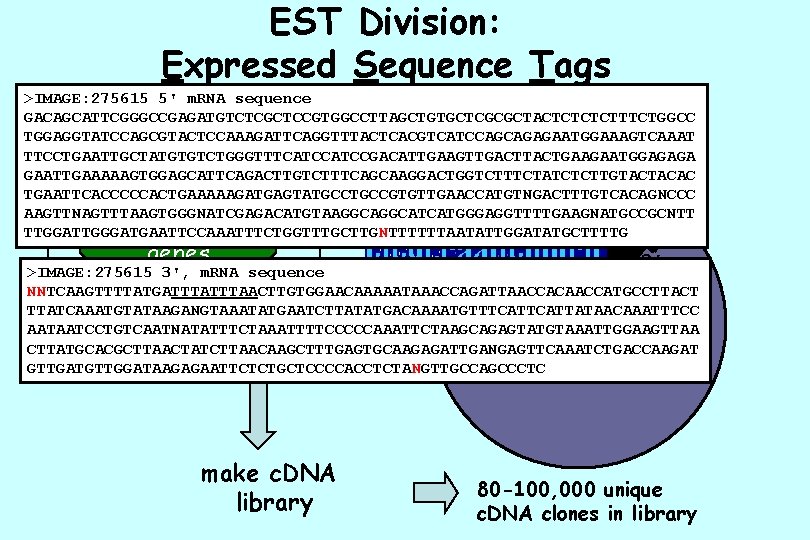
EST Division: Expressed Sequence Tags >IMAGE: 275615 5' m. RNA sequence GACAGCATTCGGGCCGAGATGTCTCGCTCCGTGGCCTTAGCTGTGCTCGCGCTACTCTTTCTGGCC TGGAGGTATCCAGCGTACTCCAAAGATTCAGGTTTACTCACGTCATCCAGCAGAGAATGGAAAGTCAAAT TTCCTGAATTGCTATGTGTCTGGGTTTCATCCGACATTGAAGTTGACTTACTGAAGAATGGAGAGA GAATTGAAAAAGTGGAGCATTCAGACTTGTCTTTCAGCAAGGACTGGTCTTTCTATCTCTTGTACTACAC TGAATTCACCCCCACTGAAAAAGATGAGTATGCCGTGTTGAACCATGTNGACTTTGTCACAGNCCC 5’ AAGTTNAGTTTAAGTGGGNATCGAGACATGTAAGGCATCATGGGAGGTTTTGAAGNATGCCGCNTT 30, 000 TTGGATTGGGATGAATTCCAAATTTCTGGTTTGCTTGNTTTTTTAATATTGGATATGCTTTTG nucleus genes 3’ >IMAGE: 275615 3', m. RNA sequence NNTCAAGTTTTATGATTTAACTTGTGGAACAAAAATAAACCAGATTAACCACAACCATGCCTTACT - isolate unique clones TTATCAAATGTATAAGANGTAAATATGAATCTTATATGACAAAATGTTTCATTATAACAAATTTCC -sequence once RNA AATAATCCTGTCAATNATATTTCTAAATTTTCCCCCAAATTCTAAGCAGAGTATGTAAATTGGAAGTTAA from each end CTTATGCACGCTTAACTATCTTAACAAGCTTTGAGTGCAAGAGATTGANGAGTTCAAATCTGACCAAGAT gene products GTTGATGTTGGATAAGAGAATTCTCTGCTCCCCACCTCTANGTTGCCAGCCCTC make c. DNA library 80 -100, 000 unique c. DNA clones in library

Unigene • A gene-oriented view of sequence entries • Uni. Gene collects expressed sequence tags (ESTs) into clusters, in an attempt to form one gene per cluster. • Use Uni. Gene to study where your gene is expressed in the body, when it is expressed, and see its abundance.

Uni. Gene • Mega. Blast based automated sequence clustering • Nonredundant set of gene oriented clusters • Each cluster a unique gene • Information on tissue types and map locations • Includes well-characterized genes and novel ESTs • Useful for gene discovery and selection of mapping reagents http: //www. ncbi. nlm. nih. gov/Uni. Gene/
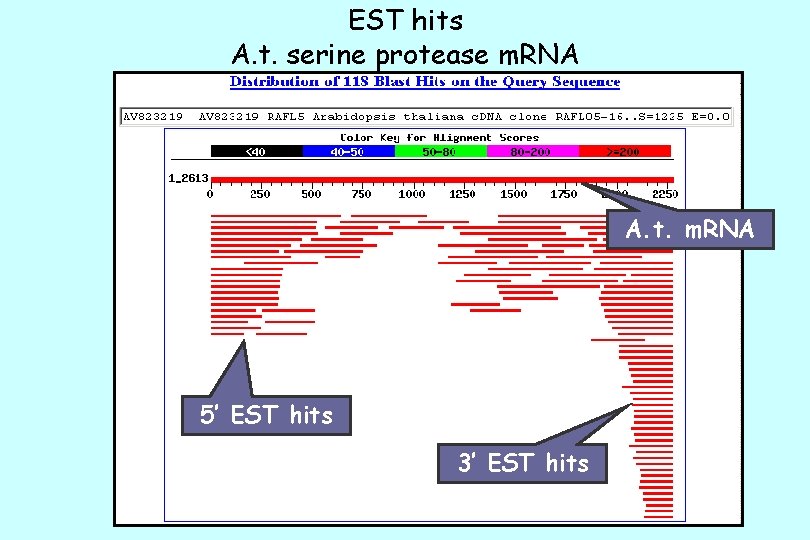
EST hits A. t. serine protease m. RNA A. t. m. RNA 5’ EST hits 3’ EST hits

Arabidopsis Uni. Gene Statistics 39, 855 87, 006 42, 137 + 32, 571 -----201, 569 m. RNAs + gene CDSs EST, 3'reads EST, 5'reads EST, other/unknown Uni. Gene Build 14 Apr. 9 th, 2002 total sequences in clusters Final Number of Clusters (sets) 26, 808 ================ 115, 000 sets total bp 25, 474 17, 654 16, 326 25, 498 expected genes 5% uncharacterized sets contain at transcripts least one known gene sets contain at least one EST sets contain both genes and ESTs

Hs Uni. Gene Statistics 73, 419 1, 181, 855 1, 461, 928 + 616, 609 -----3, 333, 811 m. RNAs + gene CDSs EST, 3'reads EST, 5'reads EST, other/unknown Uni. Gene Build 148 Apr. 8 th, 2002 total sequences in clusters Final Number of Clusters (sets) ================ 98, 816 sets total 22, 431 97, 618 21, 233 3, 000 base pairs sets at least one 30 Kcontain expected genes sets at least one 80%contain uncharacterized transcripts known gene EST sets contain both genes and ESTs
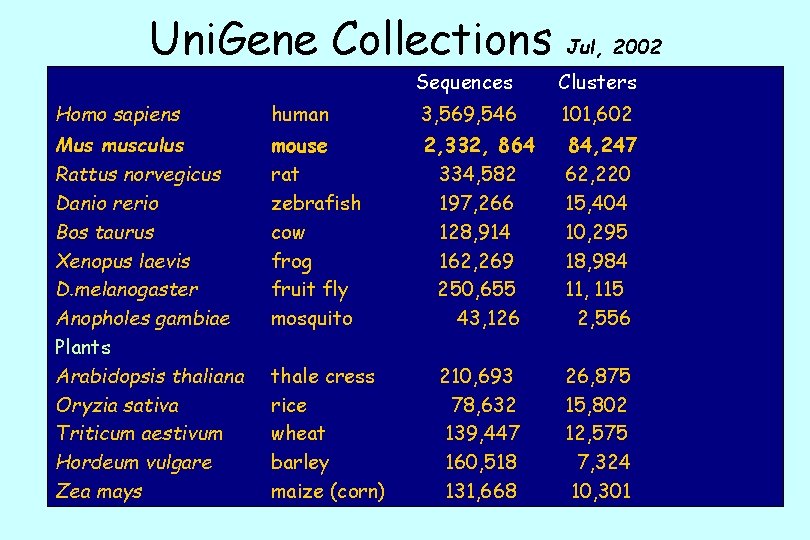
Uni. Gene Collections Jul, 2002 Sequences Clusters Homo sapiens human 3, 569, 546 101, 602 Mus musculus Rattus norvegicus Danio rerio Bos taurus Xenopus laevis D. melanogaster Anopholes gambiae Plants Arabidopsis thaliana Oryzia sativa Triticum aestivum Hordeum vulgare Zea mays mouse rat zebrafish cow frog fruit fly mosquito 2, 332, 864 334, 582 197, 266 128, 914 162, 269 250, 655 43, 126 84, 247 62, 220 15, 404 10, 295 18, 984 11, 115 2, 556 210, 693 78, 632 139, 447 160, 518 131, 668 26, 875 15, 802 12, 575 7, 324 10, 301 thale cress rice wheat barley maize (corn)
- Slides: 81