Molecular Biology Biol 480 Lecture 17 February 27

Molecular Biology Biol 480 Lecture 17 February 27 th, 2019

Announcements/Assignments Lab depends on arrival of primers. Shipment not here! Will it arrive for 10 am lab? . . . Keep reviewing lectures every day. Use the lecture summary tool, if it works for you. Exam 2 March 4 th. No lecture March 8 th. This is the ND Academy of Science meeting in Grand Forks.

Where we? TRANSCRIPTION—starting with prokaryotes • Sketch a gene—along a single line • Sketch a gene showing 2 strands---label the strands • Show the details of a promoter region • Cis and Trans elements of transcription • Initiation, elongation, termination

good animations… • http: //www. dnalc. org/resources/3 d/12 transcription-basic. html • http: //www. hhmi. org/biointeractive/dnatranscription-advanced-detail#video 7 b 9676 a 4 -5 f 24 -4837 -9 aaf-9352 eed 43 c 1 e

Transcription termination In E. coli, RNA polymerase rides along the DNA, producing RNA until it encounters a nucleotide sequence called a terminator sequence.

“Consensus” sequence of terminator • Prokaryotic terminator sequences typically have a G-C rich 5’ region and a 3’ end that includes in a string of A nucleotides. • The terminator DNA sequence is about 40 nucleotides long and RNA polymerase transcribes the sequence into RNA.

DNA and RNA play a role in termination… Near the end of the terminator, the RNA is complementary to a downstream region, and like many RNA sequences, can form intrastrand complementary base pairs.
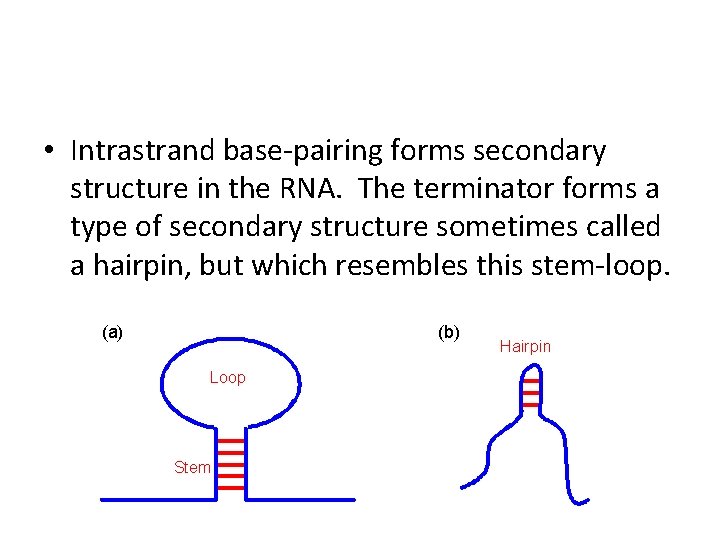
• Intrastrand base-pairing forms secondary structure in the RNA. The terminator forms a type of secondary structure sometimes called a hairpin, but which resembles this stem-loop.

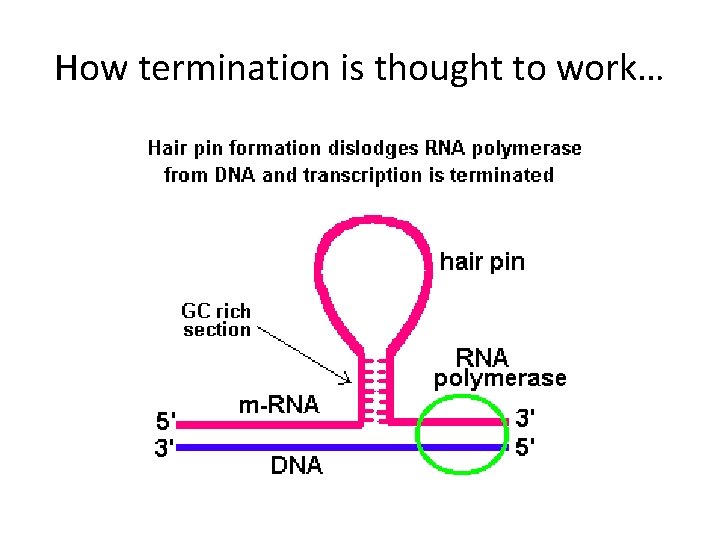
How termination is thought to work…

• Note---that the RNA itself assists in transcription termination. This is called intrinsic termination. No other factors are necessary. Termination is especially important in prokaryotes because protein encoding genes are located so near each other!

Rho-dependent termination • Some E. coli genes cannot terminate with the terminator sequence alone. They require an additional protein factor. • The factor is called Rho (r). Rho is thought to dislodge both RNApol and the new RNA from the DNA template


• Depending on the gene, many rounds of transcription will occur is succession so many molecules of RNA are formed from the same gene. • (See picture at beginning of chapter 13)
Electron microscope image Transcription trees
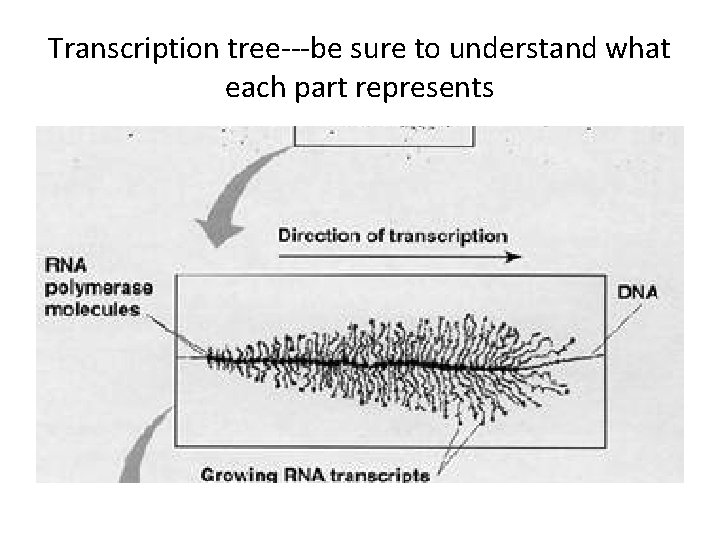
Transcription tree---be sure to understand what each part represents

• Although the synthesis of RNA is basically the same in prokaryotes and eukaryotes…There are some significant differences/additions.

In Eukaryotes… • Transcription of different types of RNA is accomplished by 3 different RNA polymerases: I, III. Table _____ , mi. RNAs

Eukaryotic gene promoters Focus on RNA polymerase II transcribed genes. (RNApol II transcribes pre-m. RNA) Transcription factors that assist RNAP II are known as TF II _____s. Core promoter elements. TATA box

• The element of the promoter common to all RNAPII genes is called the core promoter. The core promoter nearly always contains an element known as the TATA box (or Goldberg. Hogness) box at about -30. • The consensus TATA box is TATA(A or T)AA(A or G)

• The TATA box resembles the -10 consensus of the prokaryotic promoter, but does not bind RNAPII directly. – The core promoter binds a number of proteins known as transcription factors (TFs). Since any level of transcription using RNAPII requires these factors, they are called general or basal TFs.

• TFIID consists of a bunch of individual proteins. Only a few of the proteins interact with DNA. The others bind each other, in a specific way, in a specific order. • TATA-binding protein-TBP • (Recall the blobs entering the picture in the transcription video)
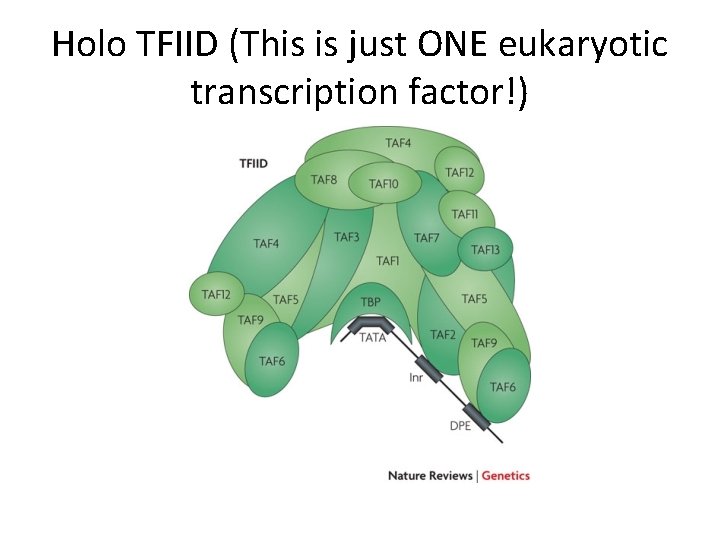
Holo TFIID (This is just ONE eukaryotic transcription factor!)
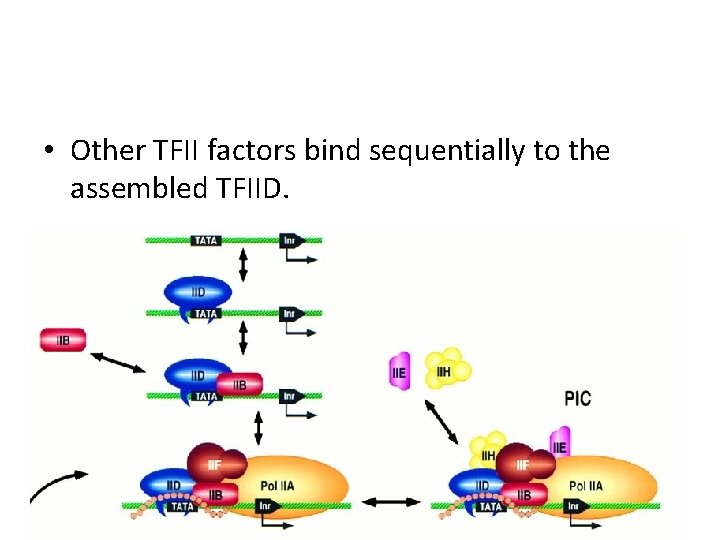
• Other TFII factors bind sequentially to the assembled TFIID.

• A second component of the core promoter is called the initiator element (INR), which is the actual DNA binding site for the RNAPII. • The INR consensus is Pu. CAPy. Py. Py, where Pu is either purine and Py is either pyrimidine.


• On either side of the TATA box and INR can be additional elements in the core promoter. • Upstream core elements are called proximal promoter elements. – While these can vary more than the TATA box consensus there a small number of distinct sequences that are used in eukaryotic promoters.
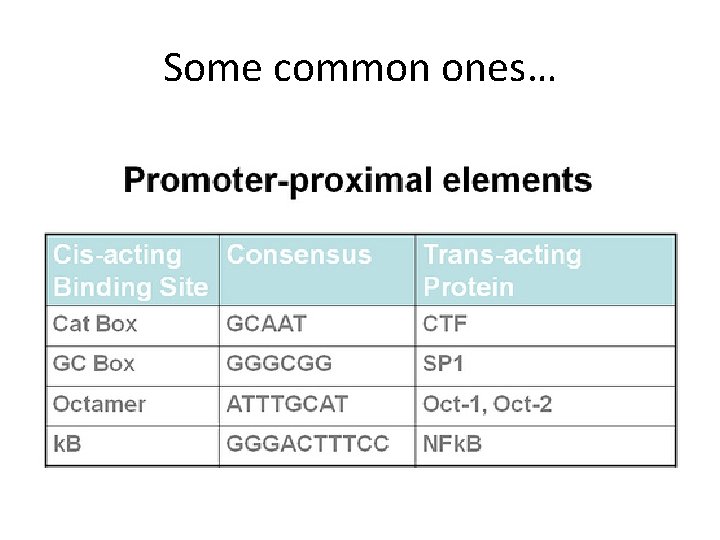
Some common ones…
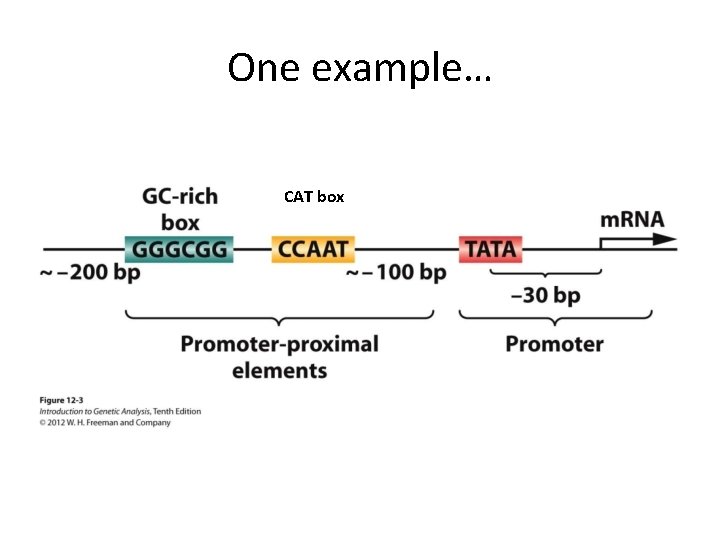
One example… CAT box

• Downstream from the TATA box can be found an additional core promoter element called the distal promoter element or DPE • The DPE seems to be essential in some eukaryotic genes for binding specific TFII basal transcription factors. It is not present in promoters of all eukaryotic genes.



Distant regulation • Finally, transcription initiation may require additional, cell-specific TFs that bind sequences outside the promoter. • A common cis element that boosts the rate of transcription initiation significantly is called an enhancer.


• The current idea on how enhancers enhance transcription is that the sequences attract and bind activator proteins. • Through a looping out of DNA between the enhancer and promoter, the activator protein(s) does make contact with the basal TFs, which somehow starts the RNAP II transcribing the gene.
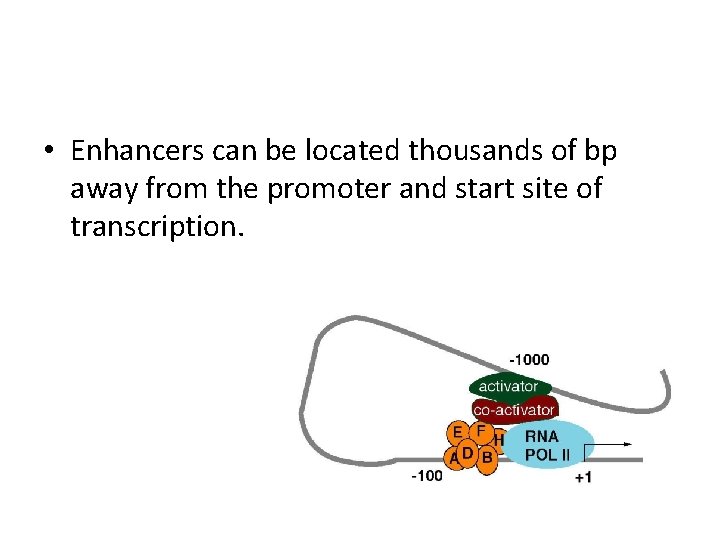
• Enhancers can be located thousands of bp away from the promoter and start site of transcription.
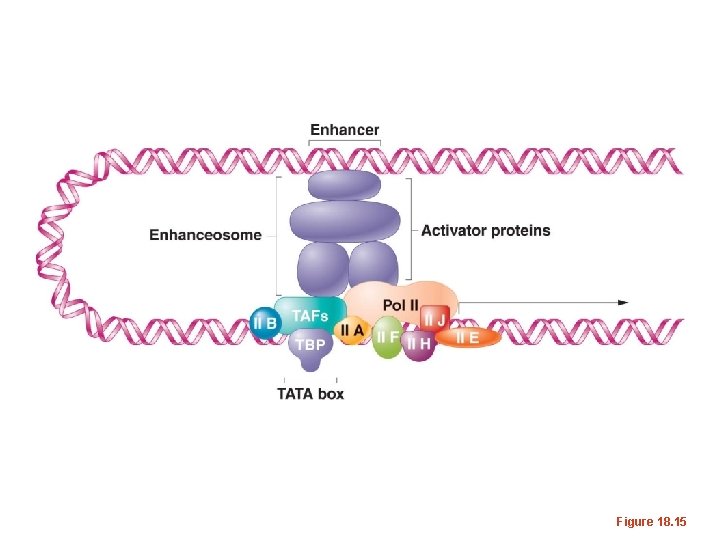
Figure 18. 15

We’ve learned that transcription initiation is a pretty complicated matter. A lot of proteins (trans elements) must meet up at the right place (the cis elements of the gene) and at the same time. • Why would such a complex system be used? – After all, our cells must have a constant supply of RNAs and proteins to carry out all the activities they use to function as cells.

• 1. Energy conservation – Transcription and translation require a constant supply of building blocks and energy. Cells can’t be wasteful, making macromolecules that aren’t needed all the time. • 2. Specialization – Cells make a specific subset of RNAs and proteins that allow them to do specific activities. Transcription initiation regulation is the primary mechanism for controlling gene expression.

• Specialization is, in large part, controlled with those elements that act at a distance—the enhancers and the activator proteins that bind them. • We will return to this soon!
- Slides: 40