Molecular Biology 1 part 2 Mamoun Ahram Ph

Molecular Biology (1) part 2 Mamoun Ahram, Ph. D Second semester, 2015 -2016

DNA replication a general mechanism

Resources This lecture Campbell and Farrell's Biochemistry, Chapter 10 3
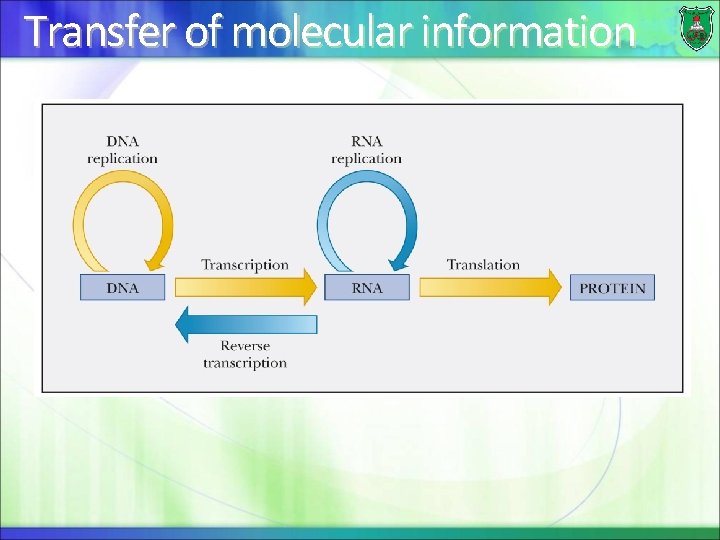
Transfer of molecular information

Some basic information The entire DNA content of the cell is known as genome DNA is organized into chromosomes Bacterial genome: usually one and circular chromosome Eukaryotic genome: multiple, linear chromosomes complexed with proteins known as histones 5
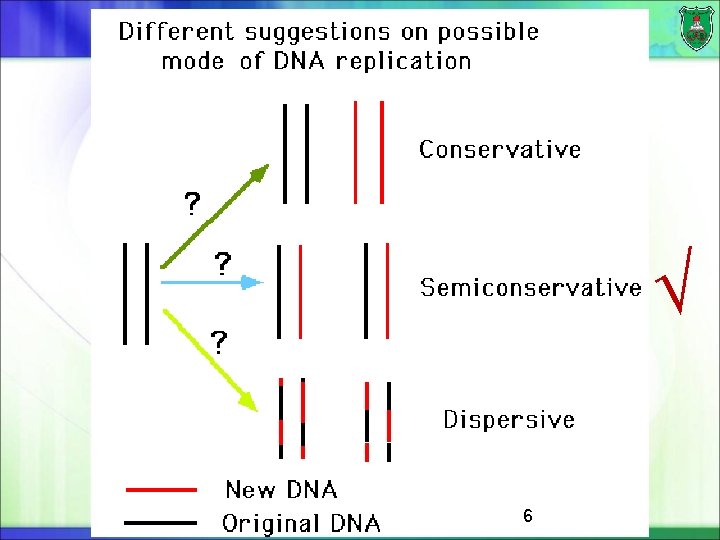
6
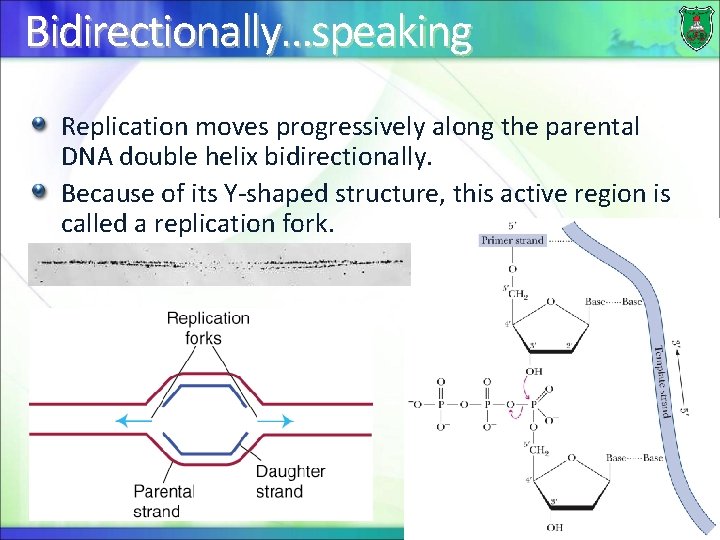
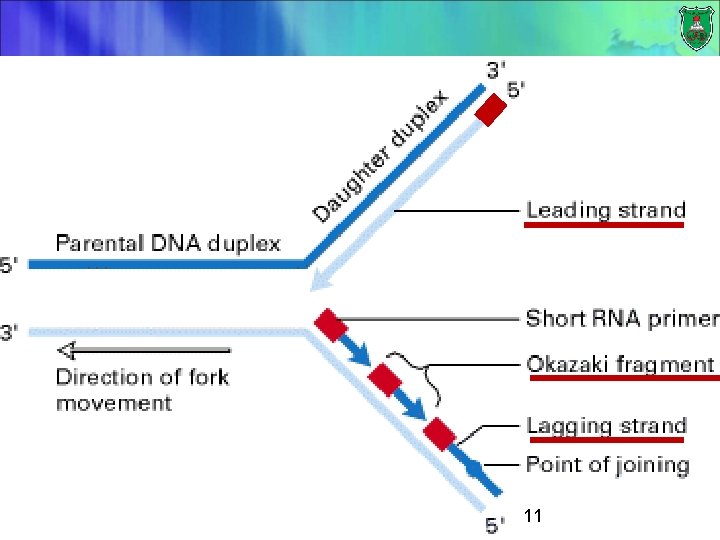
Bidirectionally…speaking Replication moves progressively along the parental DNA double helix bidirectionally. Because of its Y-shaped structure, this active region is called a replication fork. 7
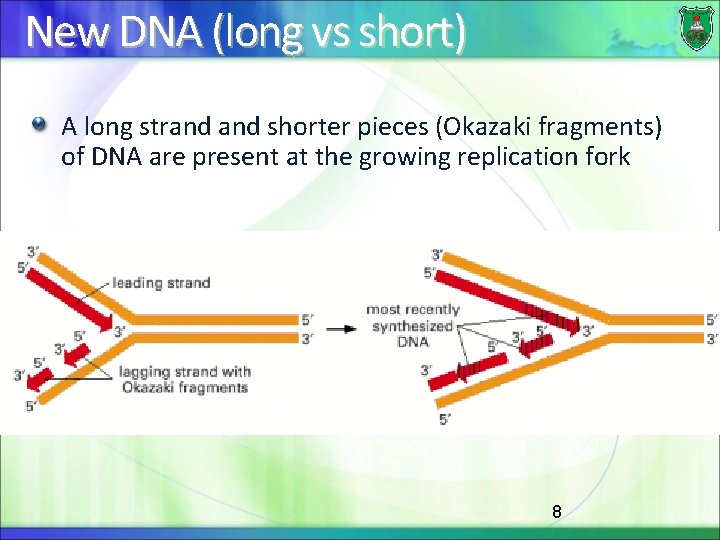
New DNA (long vs short) A long strand shorter pieces (Okazaki fragments) of DNA are present at the growing replication fork 8

Componentsof. DNAreplication 9
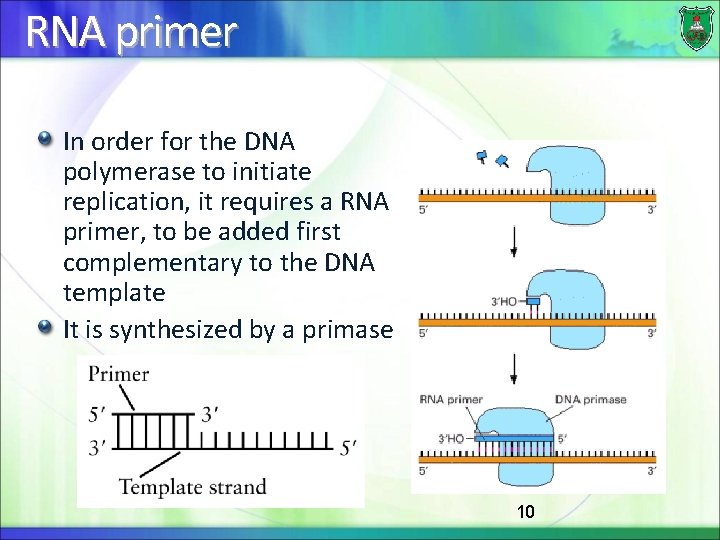
RNA primer In order for the DNA polymerase to initiate replication, it requires a RNA primer, to be added first complementary to the DNA template It is synthesized by a primase 10
11
12

DNA helicases and SSB proteins For DNA synthesis to proceed, the DNA double helix must be opened up ahead of the replication fork Opening up the DNA is done by two types of protein contribute to this process DNA helicases and single-strand DNA-binding proteins 13
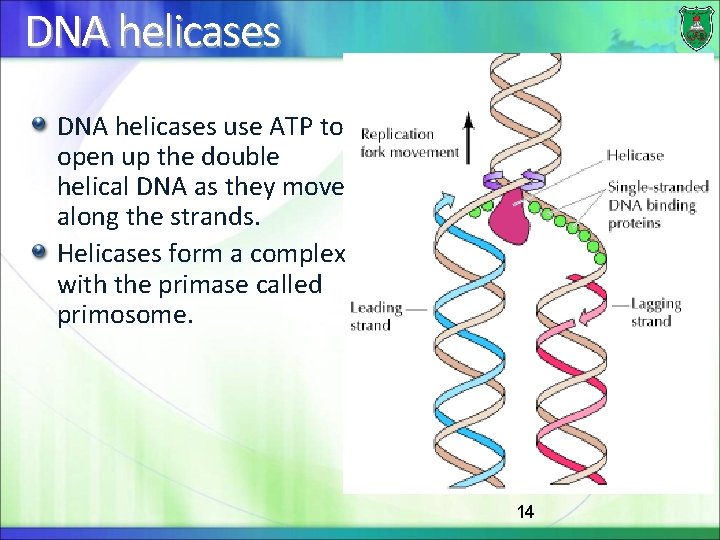
DNA helicases use ATP to open up the double helical DNA as they move along the strands. Helicases form a complex with the primase called primosome. 14

Single-strand DNA-binding (SSB) proteins bind tightly to exposed single-stranded DNA strands without covering the bases, which therefore remain available for templating These proteins: prevent the formation of the short hairpin structures protect single-stranded DNA from being degraded aid helicases by stabilizing the unwound, single-stranded conformation 15

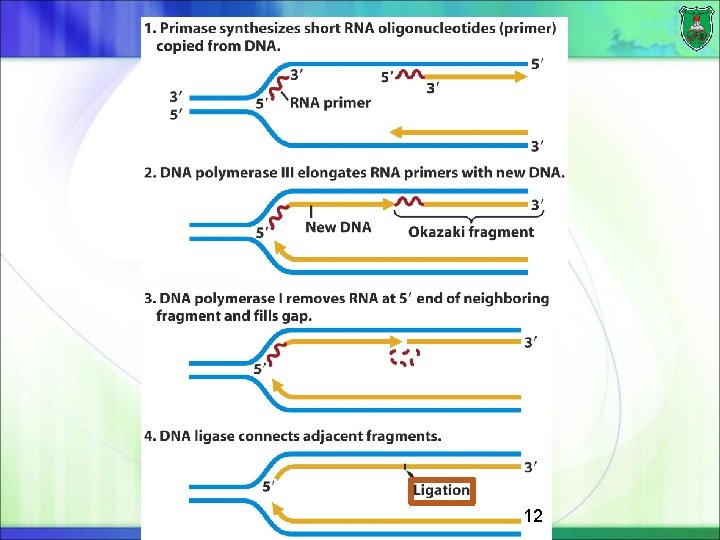
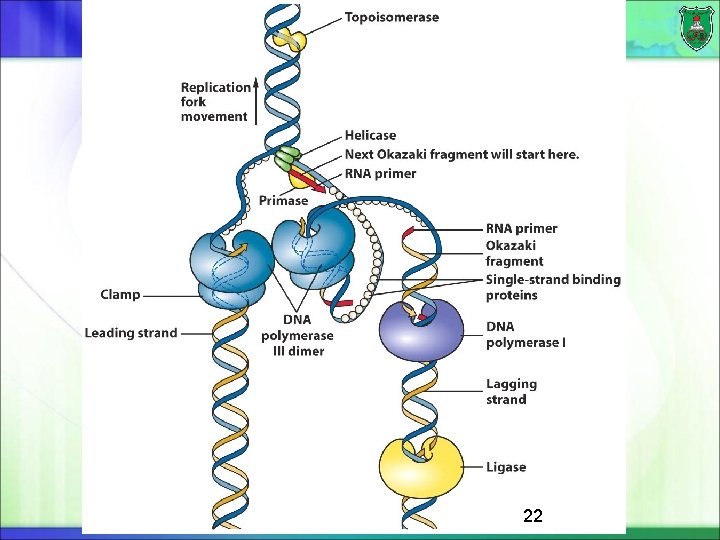
DNA polymerases in prokaryotes DNA polymerase III: DNA polymerization at the growing fork in E. coli The complex of primosome and polymerase is known as replisome DNA polymerase I: 5‘-to-3' exonuclease activity (removal of RNA primer) of each Okazaki fragment. Fills in the gaps between the lagging-strand fragments. DNA repair DNA polymerase II, IV, and V : DNA repair 16

DNA polymerase III The DNA polymerase III is a very large protein composed of 10 different polypeptides The core polymerase is composed of three subunits: α subunit contains the active site for nucleotide addition subunit is a 3 -to-5 exonuclease that removes incorrectly added (mispaired) nucleotides from the end of the growing chain subunit stimulate the exonuclease activity 17
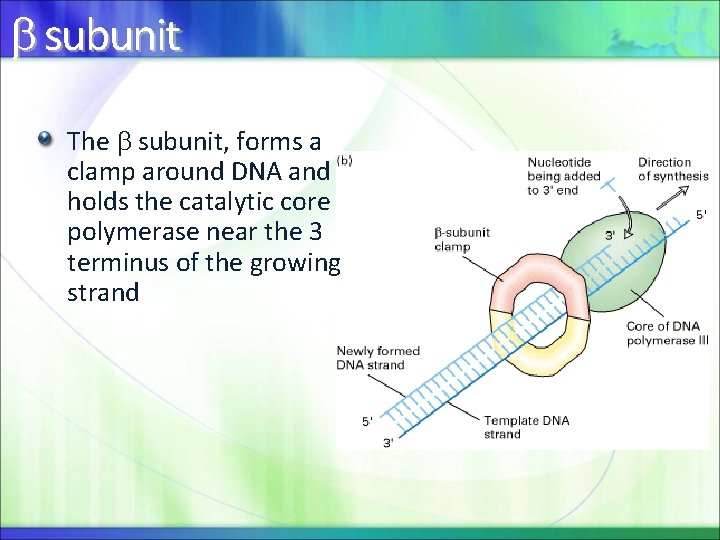
subunit The subunit, forms a clamp around DNA and holds the catalytic core polymerase near the 3 terminus of the growing strand

How accurate is DNA replication? The frequency of errors during replication is only one incorrect base per 109 to 1010 nucleotides incorporated Why is fidelity high? Hydrogen base-pairing is highly stable between G and C and between A and T. So, the DNA polymerase can catalyze the formation of phosphodiester bonds when the right hydrogen bonding takes place between the correction bases. Proofreading mechanism (a 3’ 5’ exonuclease activity) 19
20

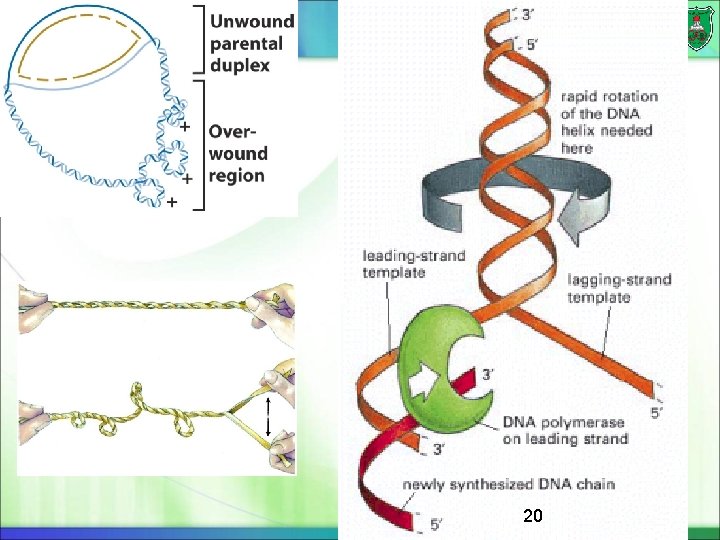
DNA topoisomerases A swivel is formed in the DNA helix by proteins known as DNA topoisomerases A DNA topoisomerase breaks then re -forms phosphodiester bonds in a DNA strand. Topoisomerase I produces an transient single-strand break (or nick) ATP-independent Topoisomerase II is responsible for untangling chromosomes by making a transient double-strand break also known as gyrase in bacteria ATP-dependent
22
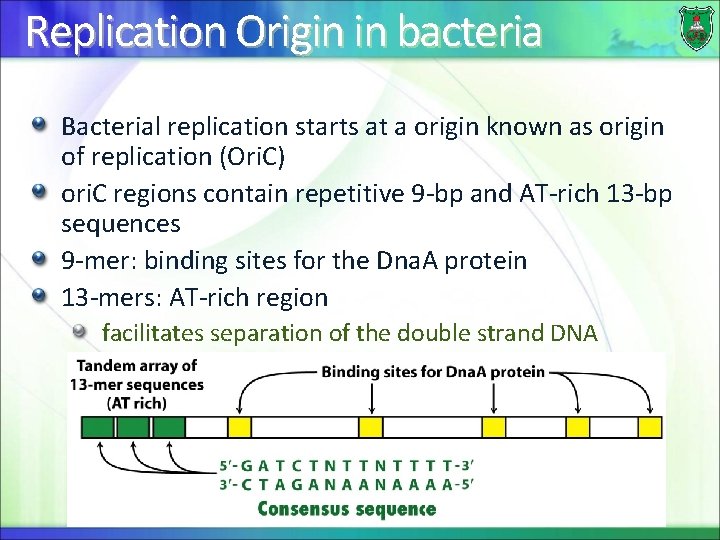
Replication Origin in bacteria Bacterial replication starts at a origin known as origin of replication (Ori. C) ori. C regions contain repetitive 9 -bp and AT-rich 13 -bp sequences 9 -mer: binding sites for the Dna. A protein 13 -mers: AT-rich region facilitates separation of the double strand DNA 23
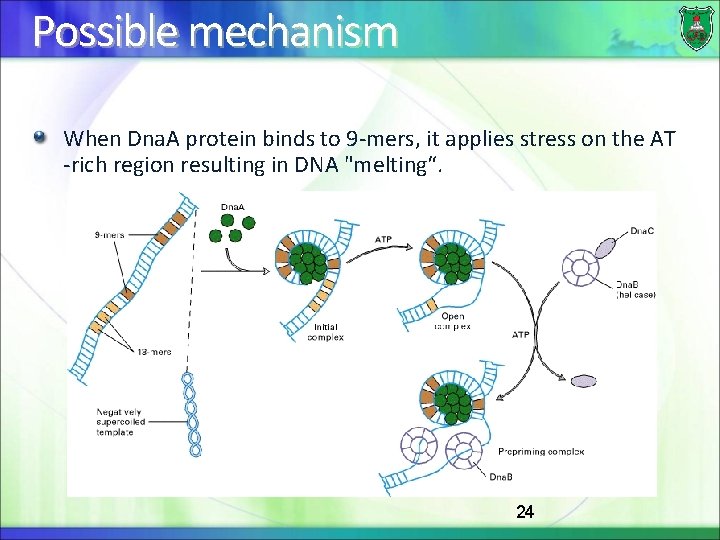
Possible mechanism When Dna. A protein binds to 9 -mers, it applies stress on the AT -rich region resulting in DNA "melting“. 24
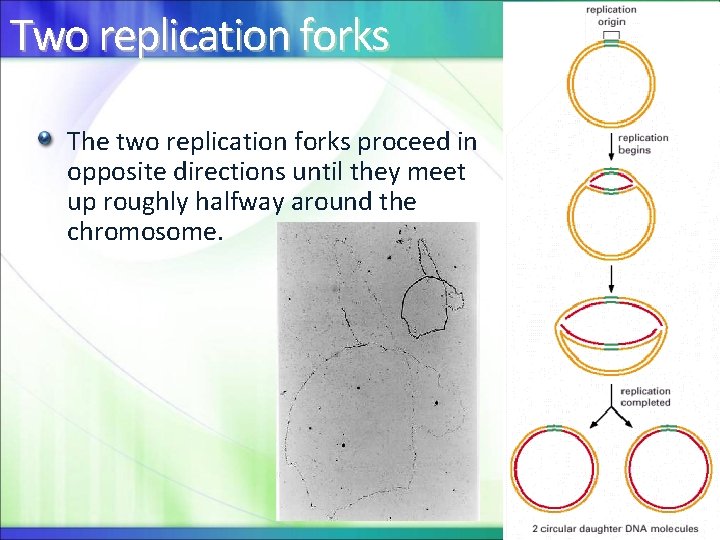
Two replication forks The two replication forks proceed in opposite directions until they meet up roughly halfway around the chromosome.
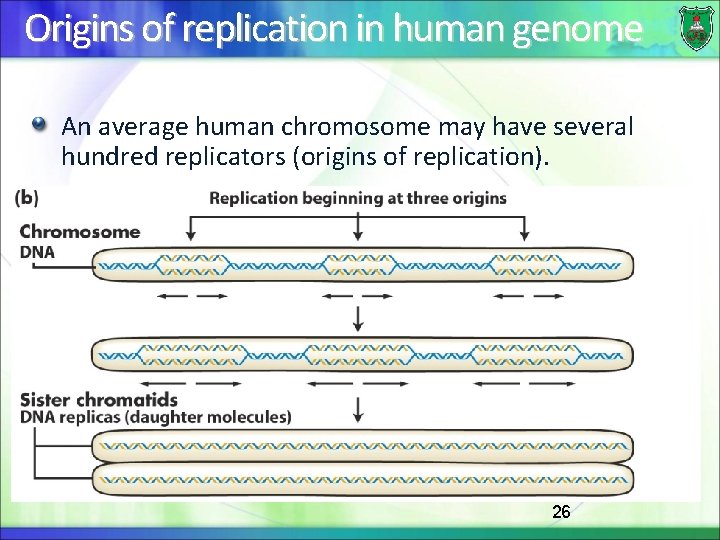
Origins of replication in human genome An average human chromosome may have several hundred replicators (origins of replication). 26

DNA polymerase in eukaryotes Eukaryotic cells contain 9 DNA polymerases; most of them for DNA repair. 27

The mechanism of replication DNA polymerase is responsible for synthesis of leading strand. The DNA primase is part of the DNA polymerase α. Polymerase α begins each Okazaki fragment on the lagging strand with RNA before passing the DNA to DNA polymerase then synthesizes the remainder of each Okazaki fragment. The polymerases do not have a 5’ 3’ exonuclease The primer is removed by two special enzymes. DNA polymerase then fills in the gap. 28

Roles of other polymerases Polymerase may function primarily in the repair of DNA damage , , , and DNA polymerases Replication and repair of damaged DNA No 3' to 5' exonuclease activity 29

Role of chromatin It seems that the timing of replication is related to the packing of the DNA in chromatin In eukaryotes, chromosome duplication requires that DNA is freed from hisotnes and that new chromosomal histones be assembled onto the DNA behind each replication fork 30
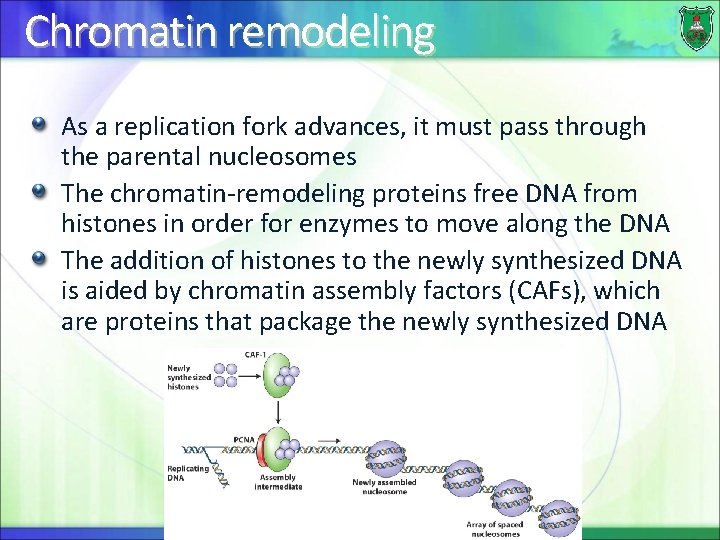
Chromatin remodeling As a replication fork advances, it must pass through the parental nucleosomes The chromatin-remodeling proteins free DNA from histones in order for enzymes to move along the DNA The addition of histones to the newly synthesized DNA is aided by chromatin assembly factors (CAFs), which are proteins that package the newly synthesized DNA 31

Role of telomeres As the growing fork approaches the end of a linear chromosome, the lagging-strand template is not completely replicated When the final RNA primer is removed, there is no place onto which DNA polymerase can build to fill the resulting gap This would lead the lagging-strand to be shortened at each cell division 32
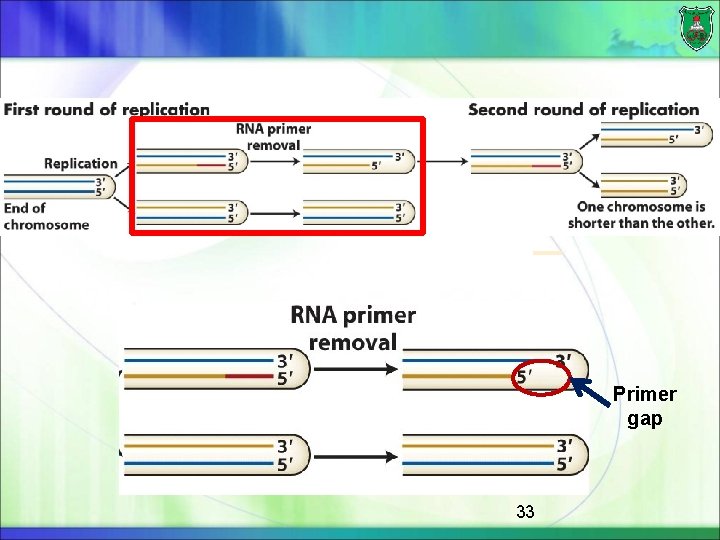
Primer gap 33

Telomerase The enzyme that prevents this progressive shortening of the lagging strand is a modified enzyme called telomerase, which can elongate the lagging-strand template 34

The mechanism Telomere DNA sequences consist of many repeats of a short sequence In humans, this sequence is GGGTTA, extending for about 10, 000 nucleotides. Telomerase recognizes the telomere DNA repeat sequence and elongates it in the 5 -to-3 direction The telomerase synthesizes a new copy of the repeat, using an RNA template/primer that is a component of the enzyme itself 35
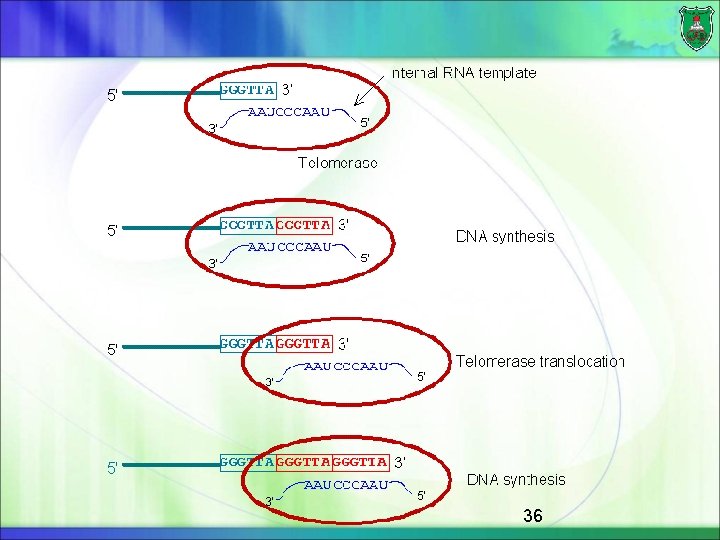
36

37

How do we age? As we grow older, the activity of telomerase is reduced An inverse relationship between age and telomeric length has been observed The gradual shortening of the chromosome ends leads to cell death, and it has even been suggested that life span is determined by the length of telomeres 38

Elixir of youth

DNA mutations http: //www. ncbi. nlm. nih. gov/books/NBK 21897/ 40

Types of mutations Micromutation that involve small regions of the DNA Macromutations that involve the chromosomes as a whole 41

DNA micromutations Single-base mutations can result in different types of mutations such as: missense: a change of an amino acid nonsense: premature termination of protein synthesis frameshift: altering the reading of amino acid sequence silent: does not lead to any change at the protein level, but contributes to genetic variability among individuals 42
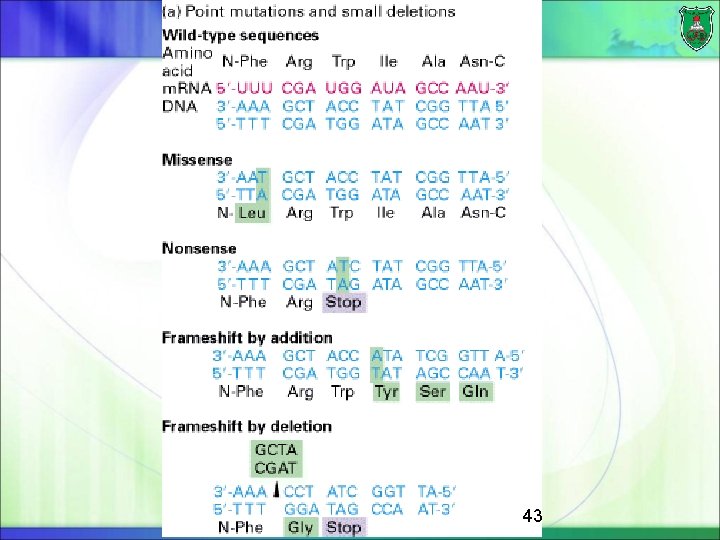
43
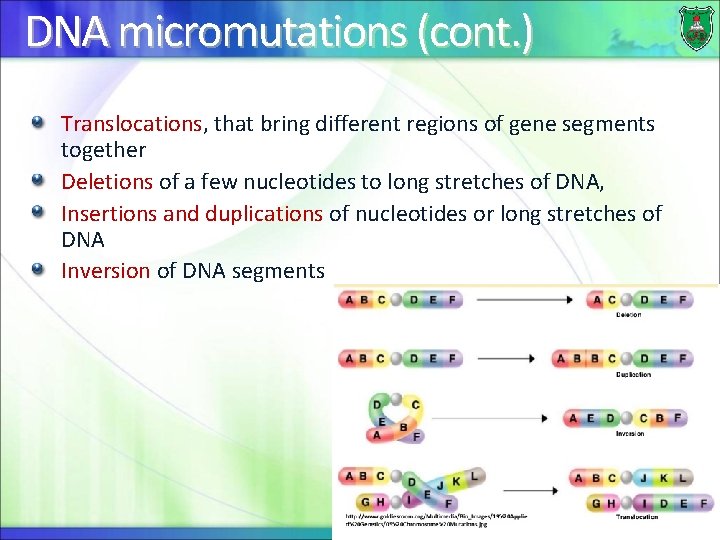
DNA micromutations (cont. ) Translocations, that bring different regions of gene segments together Deletions of a few nucleotides to long stretches of DNA, Insertions and duplications of nucleotides or long stretches of DNA Inversion of DNA segments 44

Causes of DNA mutations can arise spontaneously or induced. Spontaneous mutations are naturally occurring mutations and arise in all cells. They arise from a variety of sources, including errors in DNA replication and spontaneous lesions Induced mutations are produced when an organism is exposed to a mutagenic agent, or mutagen. 45

Errors in DNA replication An error in DNA replication can occur when an inaccurate nucleotide pair (say, A-C instead of G-C) forms in DNA synthesis, leading to a base substitution Each of the bases in DNA can appear in one of several forms, called tautomers. 46
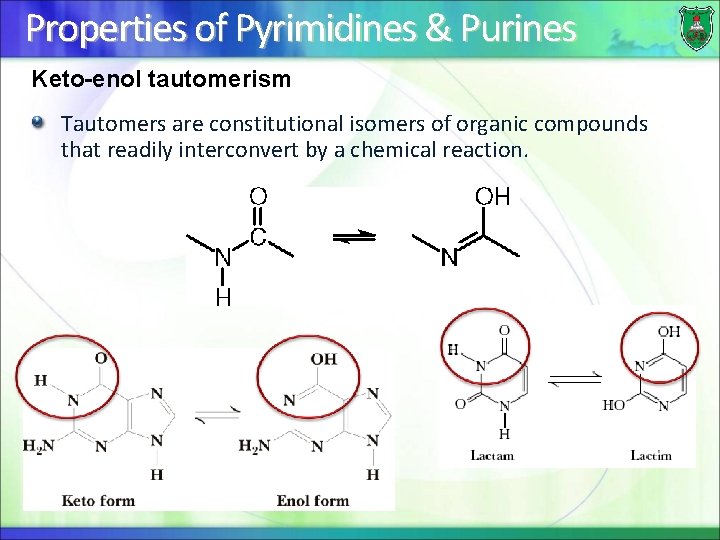
Properties of Pyrimidines & Purines Keto-enol tautomerism Tautomers are constitutional isomers of organic compounds that readily interconvert by a chemical reaction.
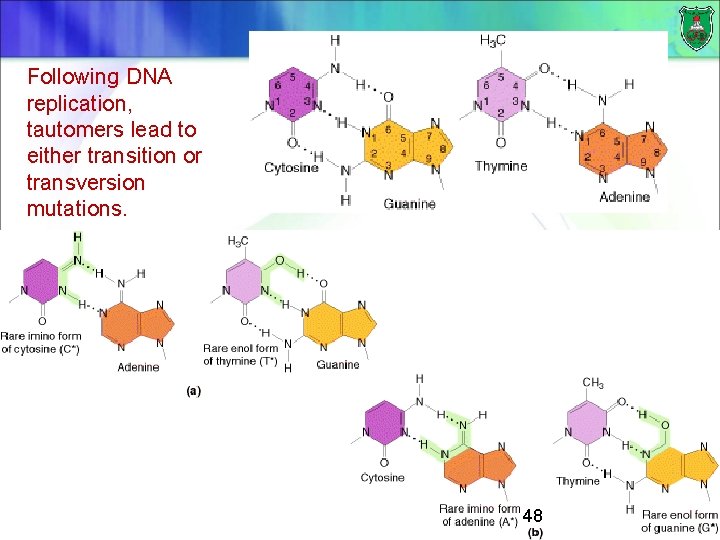
Following DNA replication, tautomers lead to either transition or transversion mutations. 48
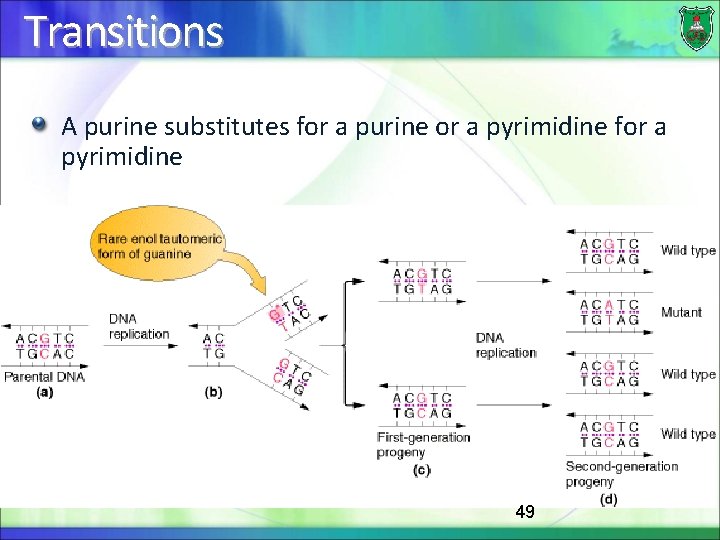
Transitions A purine substitutes for a purine or a pyrimidine for a pyrimidine 49

Transversions In transversion mutations, a pyrimidine substitutes for a purine or vice versa In this case, a replication error would require mispairing of a purine with a purine or a pyrimidine with a pyrimidine 50

Other errors of DNA replication Frameshift mutations These mutations often occurred at repeated sequences Such mutations change the reading "frame" of codons and leads to changes in the amino acid sequence of the produced protein Deletions and duplication They also often occur at sequence repeats 51

Spontaneous lesions are naturally occurring type of DNA damage that can generate mutations Depurination Deamination Oxidatively damaged bases 52
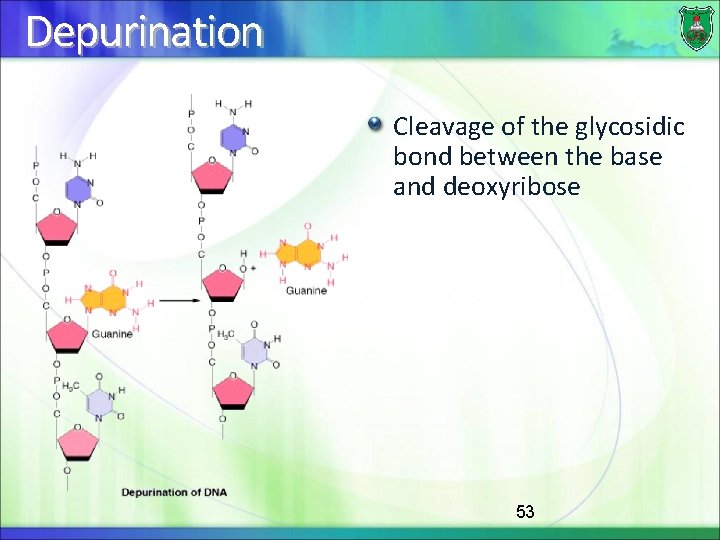
Depurination Cleavage of the glycosidic bond between the base and deoxyribose 53
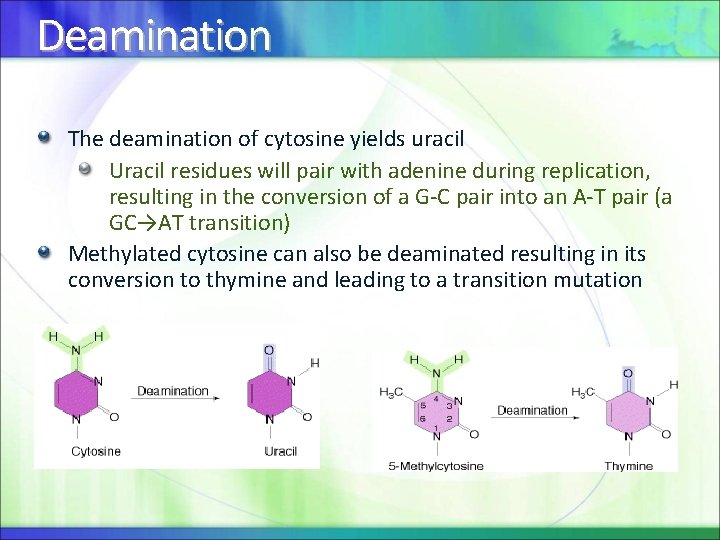
Deamination The deamination of cytosine yields uracil Uracil residues will pair with adenine during replication, resulting in the conversion of a G-C pair into an A-T pair (a GC→AT transition) Methylated cytosine can also be deaminated resulting in its conversion to thymine and leading to a transition mutation

Oxidatively damaged bases Active oxygen species such as hydrogen peroxide (H 2 O 2) are normally produced during metabolism Example: 8 -oxo-7 -hydrodeoxyguanosine (8 -oxod. G, or GO) product mispairs with A, resulting in G→T transversions. 55

Inducedmutations 56

Mechanisms of mutagenesis Mutagens induce mutations by at least three different mechanisms: replace a base in the DNA (base analog) alter a base so that it specifically mispairs with another base (alkylation) damage a base so that it can no longer pair with any base under normal conditions 57

Incorporation of base analogs Some chemical compounds are similar to the normal bases of DNA that they are incorporated into DNA in place of normal bases; such compounds are called base analogs. These analogs cause incorrect nucleotides to be inserted opposite to them during replication 58
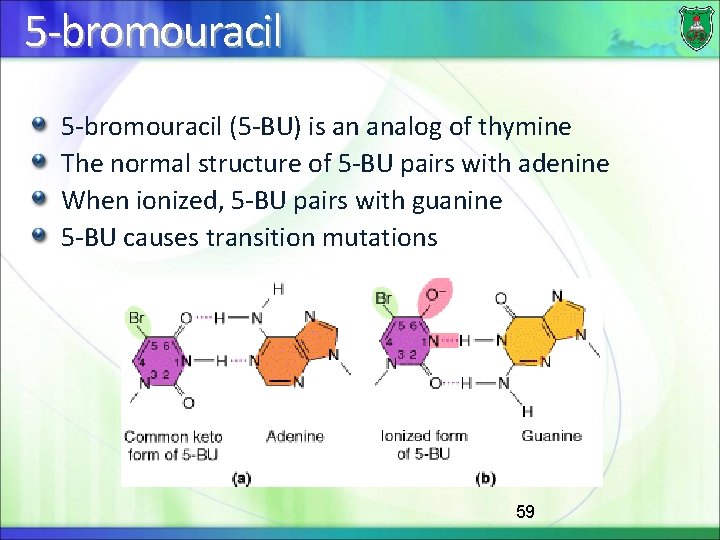
5 -bromouracil (5 -BU) is an analog of thymine The normal structure of 5 -BU pairs with adenine When ionized, 5 -BU pairs with guanine 5 -BU causes transition mutations 59
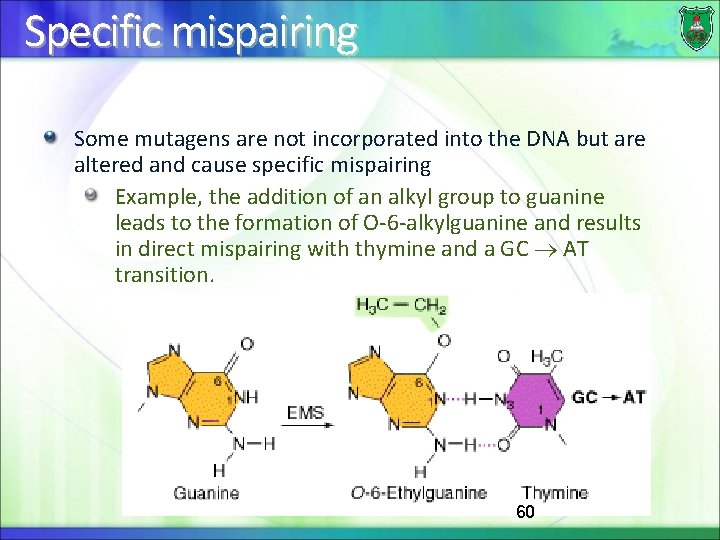
Specific mispairing Some mutagens are not incorporated into the DNA but are altered and cause specific mispairing Example, the addition of an alkyl group to guanine leads to the formation of O-6 -alkylguanine and results in direct mispairing with thymine and a GC AT transition. 60

Base damage A large number of mutagens damage one or more bases, so no specific base pairing is possible. Ionizing radiation results in the formation of ionized and excited molecules that can cause damage to cellular components including DNA Many different types of reactive oxygen species are produced that can damage bases, cause breakage of the N-glycosidic bond (AP sites), or cause strand breaks 61
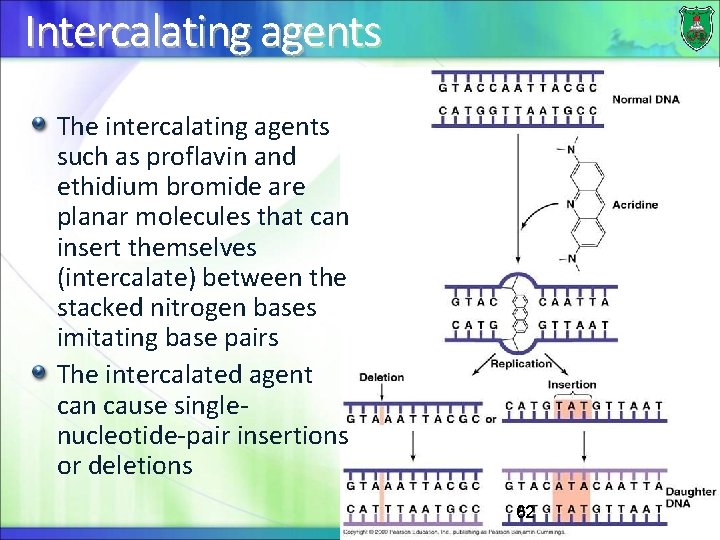
Intercalating agents The intercalating agents such as proflavin and ethidium bromide are planar molecules that can insert themselves (intercalate) between the stacked nitrogen bases imitating base pairs The intercalated agent can cause singlenucleotide-pair insertions or deletions 62

Mutagens and carcinogens Mutagenicity and carcinogenicity are clearly correlated when most mutagens are also carcinogens (approximately 90 percent). Rapid tests make use of microbes to test for mutagenicity The most widely used test was developed in the 1970 s by Bruce Ames, who worked with Salmonella typhimurium. Chemicals detected by this test can be regarded as potential carcinogens. http: //www. ncbi. nlm. nih. gov/books/NBK 21788/ 63

Features of Salmonella typhimurium The Ames test uses a mutant strain of S. typhimurium cannot grow in the absence of the amino acid histidine because a mutation has occurred in a gene that encodes one of the enzymes necessary for histidine biosynthesis They also carry a mutation that inactivates a DNA repair system They also carry a mutation that eliminates the protective lipopolysaccharide coating of wild-type Salmonella to enable the entry of many different chemicals into the cell 64

Reversion mutation In order for these cells to survive in the absence of histidine, they must have a mutation that corrects the original mutation that prevented the production of the missing enzyme This type of mutation is known as reversion, because this second mutation returns the mutant to the wildtype genotype This reversion can happen spontaneously or as the result of a mutagen 65

What is considered a mutagen? A compound must increase the number of colonies grown in the absence of histidine by double compared to cells grown in the absence of the compound Water Motor oil Alcohol Drug X ﻗﺮﻭﺵ 5 ﺷﻴﺒﺲ ﺃﺒﻮ 10 50 43 9 200 66
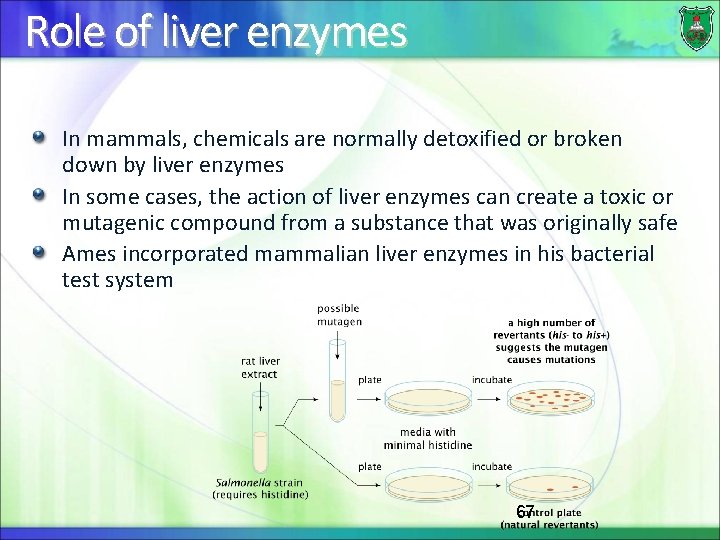
Role of liver enzymes In mammals, chemicals are normally detoxified or broken down by liver enzymes In some cases, the action of liver enzymes can create a toxic or mutagenic compound from a substance that was originally safe Ames incorporated mammalian liver enzymes in his bacterial test system 67
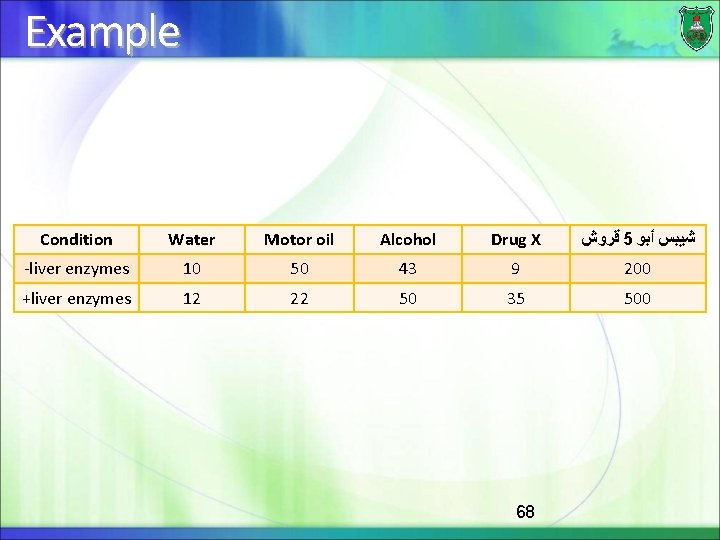
Example Condition Water Motor oil Alcohol Drug X ﻗﺮﻭﺵ 5 ﺷﻴﺒﺲ ﺃﺒﻮ -liver enzymes 10 50 43 9 200 +liver enzymes 12 22 50 35 500 68
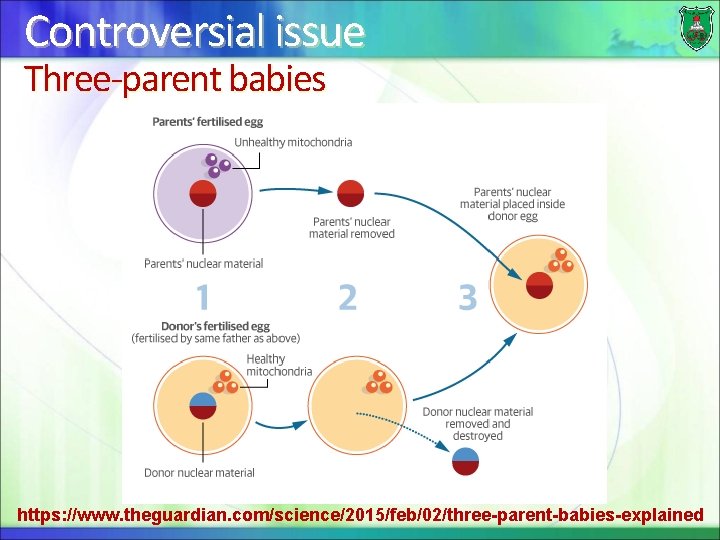
Controversial issue Three-parent babies https: //www. theguardian. com/science/2015/feb/02/three-parent-babies-explained

DNA repair mechanisms http: //www. ncbi. nlm. nih. gov/books/NBK 22004/

Resources This lecture Campbell and Farrell’s Biochemistry, pp. 263 -268 An Introduction to Genetic Analysis. 7 th edition. Griffiths AJF, Miller JH, Suzuki DT, et al. , New York: W. H. Freeman; 2000. 71

DNA repair Maintaining genetic stability requires not only an accurate mechanism of DNA replication, but also mechanisms for repairing DNA damage These mechanisms are collectively called DNA repair 72

Repair mechanisms Prevention of errors before they happen Direct reversal of damage Excision-repair pathways Postreplication repair 73

Preventionoferrorsbeforethey happen 74

Reactive oxygen species Some enzymatic systems neutralize potentially damaging compounds before they even react with DNA One example of such a system is the detoxification of reactive oxygen species and oxygen radicals, which are produced during oxidative damage to DNA 75
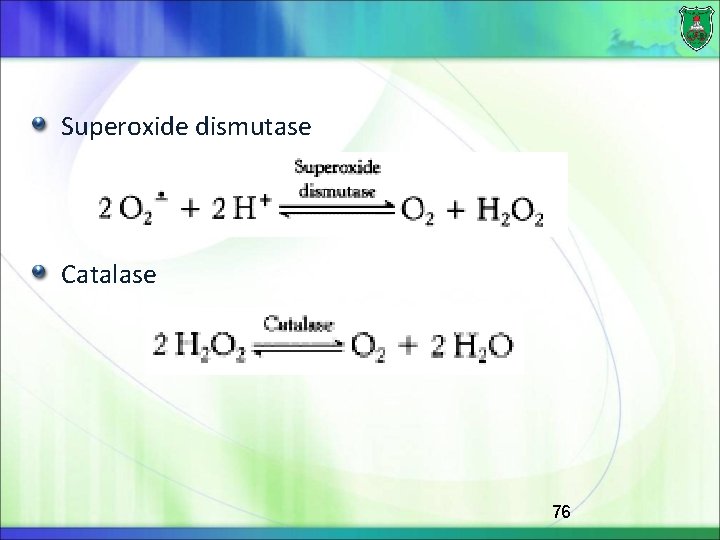
Superoxide dismutase Catalase 76
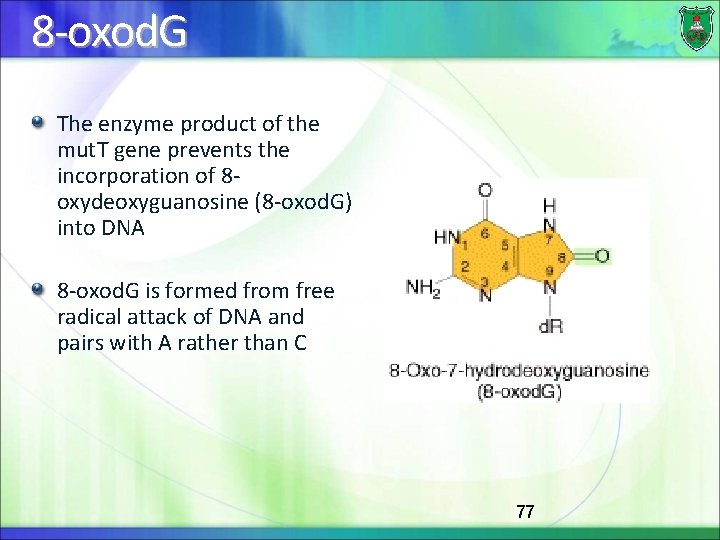
8 -oxod. G The enzyme product of the mut. T gene prevents the incorporation of 8 oxydeoxyguanosine (8 -oxod. G) into DNA 8 -oxod. G is formed from free radical attack of DNA and pairs with A rather than C 77

Directreversalofdamage 78
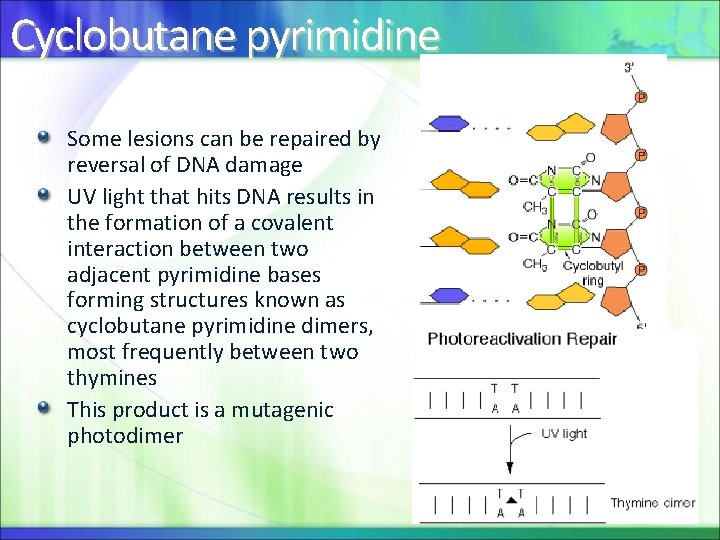
Cyclobutane pyrimidine Some lesions can be repaired by reversal of DNA damage UV light that hits DNA results in the formation of a covalent interaction between two adjacent pyrimidine bases forming structures known as cyclobutane pyrimidine dimers, most frequently between two thymines This product is a mutagenic photodimer

Photolyase Cyclobutane pyrimidine can be repaired by a photolyase that has been found in bacteria and lower eukaryotes but not in humans 80

Excision-repairpathways 81

Types General excision repair Coupling of transcription and repair Specific excision pathways 82
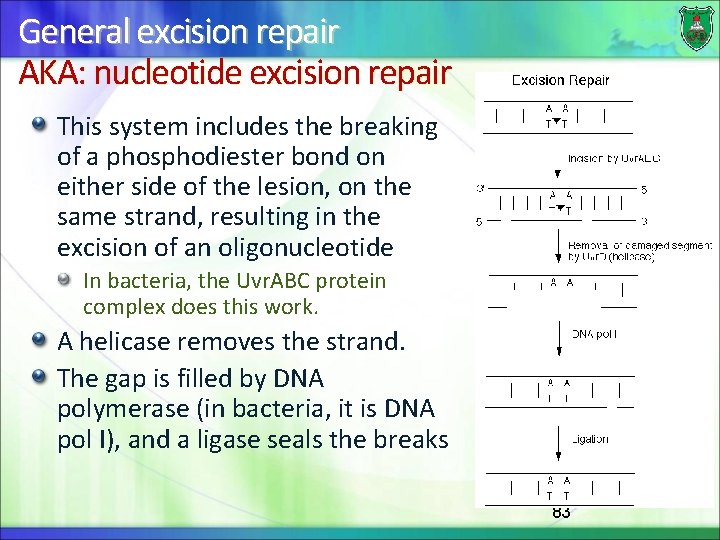
General excision repair AKA: nucleotide excision repair This system includes the breaking of a phosphodiester bond on either side of the lesion, on the same strand, resulting in the excision of an oligonucleotide In bacteria, the Uvr. ABC protein complex does this work. A helicase removes the strand. The gap is filled by DNA polymerase (in bacteria, it is DNA pol I), and a ligase seals the breaks 83

In human… In human cells, the process is more complex than its bacterial counterpart. However, the basic steps are the same as those in E. coli Defect in this mechanism causes a condition known as Xeroderma pigmentosum 84

Transcription and repair In both eukaryotes and prokaryotes, there is a preferential repair of the transcribed strand of DNA for actively expressed genes RNA polymerase pauses when encountering a lesion The general transcription factor TFIIH and other factors carry out the incision, excision, and repair reactions Then transcription can continue normally 85

Specificexcisionpathways 86

DNA glycosylase repair pathway DNA glycosylases do not cleave phosphodiester bonds, but instead cleave N-glycosidic (base-sugar) bonds of damaged bases , liberating the altered base and generating an apurinic or an apyrimidinic site, both called AP sites The resulting AP site is then repaired by an AP endonuclease repair pathway 87

DNA glycosylases Numerous DNA glycosylases exist Example: uracil-DNA glycosylase, removes uracil from DNA Uracil residues, which result from the spontaneous deamination of cytosine can lead to a C T transition if unrepaired. The AP endonucleases cleaves the phosphodiester bonds at AP sites An exonuclease removes the strand, DNA polymerase I fills in the gap, and DNA ligase and re-forms the bond. 88
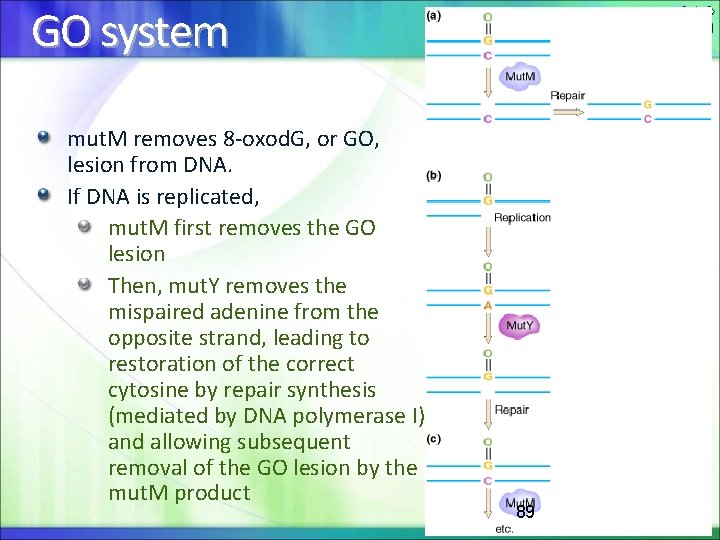
GO system mut. M removes 8 -oxod. G, or GO, lesion from DNA. If DNA is replicated, mut. M first removes the GO lesion Then, mut. Y removes the mispaired adenine from the opposite strand, leading to restoration of the correct cytosine by repair synthesis (mediated by DNA polymerase I) and allowing subsequent removal of the GO lesion by the mut. M product 89

Postreplicationrepair 90

Mismatch repair Some repair pathways are capable of recognizing errors even after DNA replication has already occurred One such system, termed the mismatch repair system, can detect mismatches that occur in DNA replication 91
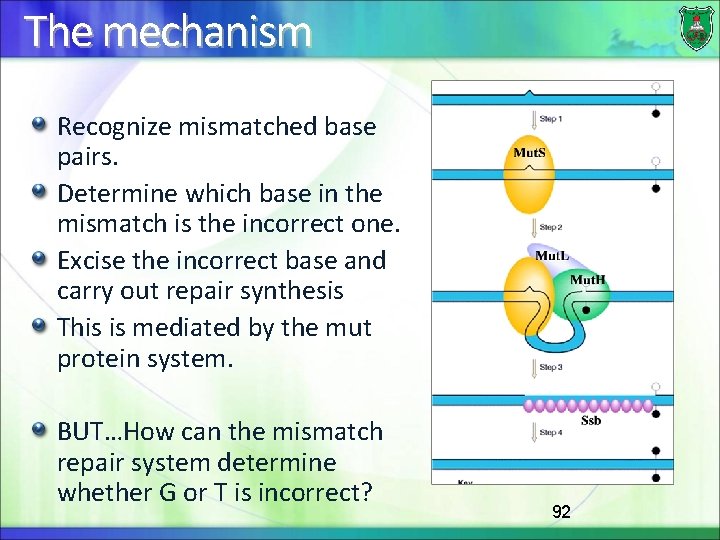
The mechanism Recognize mismatched base pairs. Determine which base in the mismatch is the incorrect one. Excise the incorrect base and carry out repair synthesis This is mediated by the mut protein system. BUT…How can the mismatch repair system determine whether G or T is incorrect? 92
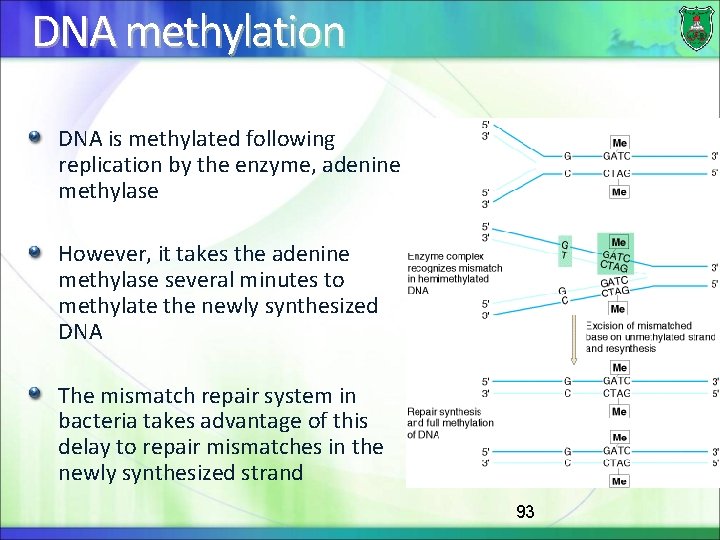
DNA methylation DNA is methylated following replication by the enzyme, adenine methylase However, it takes the adenine methylase several minutes to methylate the newly synthesized DNA The mismatch repair system in bacteria takes advantage of this delay to repair mismatches in the newly synthesized strand 93
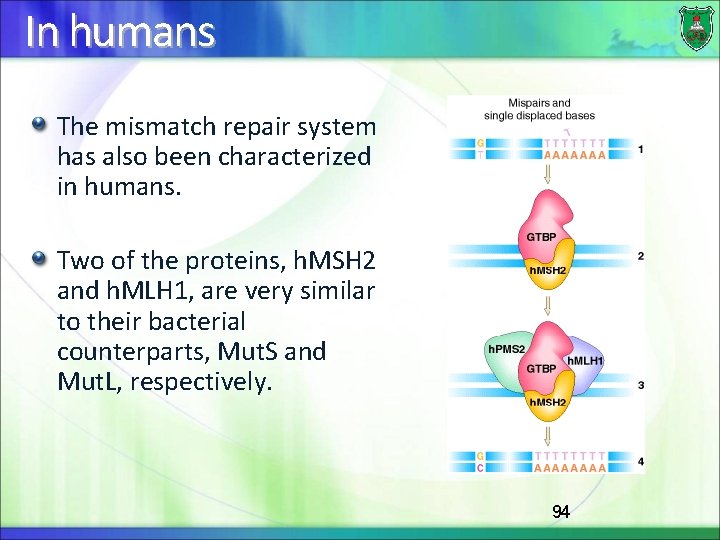
In humans The mismatch repair system has also been characterized in humans. Two of the proteins, h. MSH 2 and h. MLH 1, are very similar to their bacterial counterparts, Mut. S and Mut. L, respectively. 94

Hereditary nonpolyposis colon cancer (HNPCC) Several genes responsible a type of colon cancer have been identified, including MSH 2 and MLH 1 The protein products of the MSH 2 and MLH 1 genes appear to have critical roles in the recognition and repair of DNA mismatches They are homologous to the mut. L and mut. S genes in bacteria and yeast 95

SOS system A large number of mutagens such as ultraviolet light damage one or more bases, resulting in a replication block, because DNA synthesis will not proceed past a base that cannot specify its complementary partner by hydrogen bonding In bacterial cells, such replication blocks can be bypassed by inserting nonspecific bases. The process requires the activation of a special system, the SOS system 96
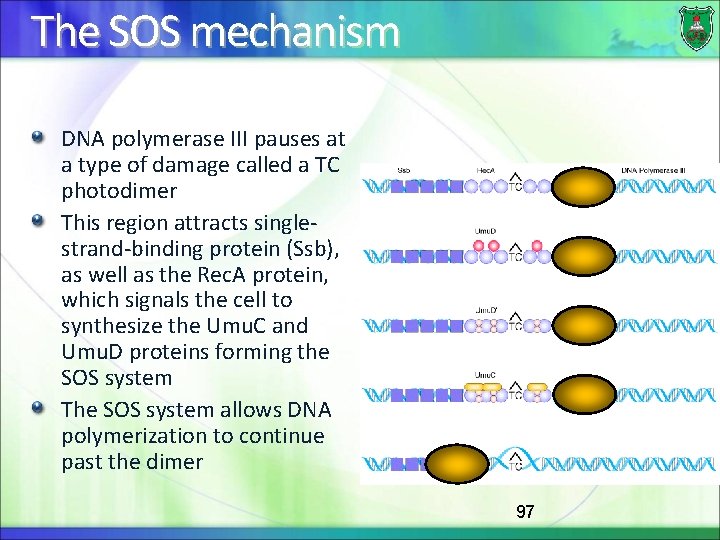
The SOS mechanism DNA polymerase III pauses at a type of damage called a TC photodimer This region attracts singlestrand-binding protein (Ssb), as well as the Rec. A protein, which signals the cell to synthesize the Umu. C and Umu. D proteins forming the SOS system The SOS system allows DNA polymerization to continue past the dimer 97

SOS system and mutations However, the SOS system inserts the necessary number of bases (often incorrect ones) directly across from the lesion and replication continues without a gap SOS system often generates mutations 98
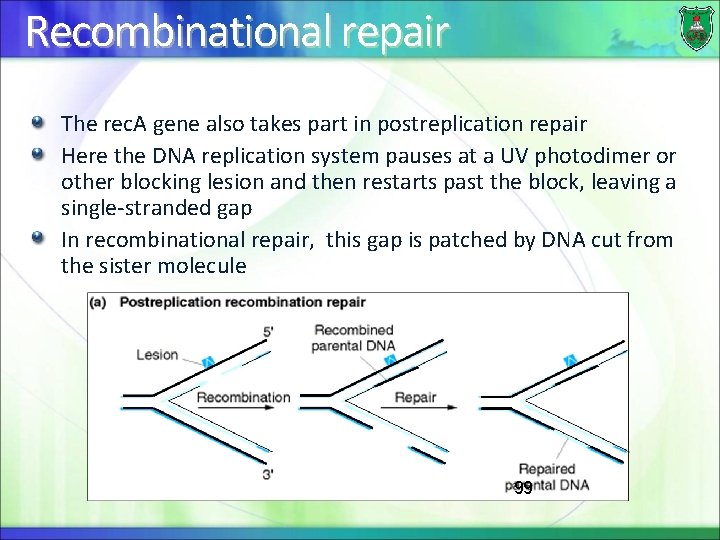
Recombinational repair The rec. A gene also takes part in postreplication repair Here the DNA replication system pauses at a UV photodimer or other blocking lesion and then restarts past the block, leaving a single-stranded gap In recombinational repair, this gap is patched by DNA cut from the sister molecule 99
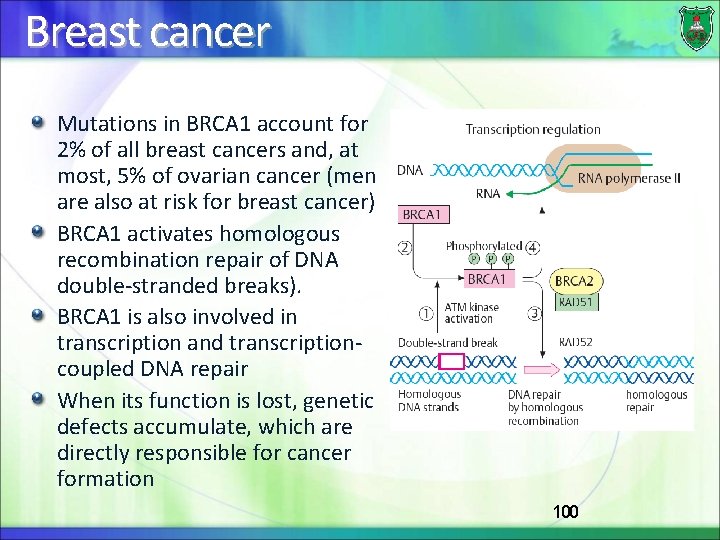
Breast cancer Mutations in BRCA 1 account for 2% of all breast cancers and, at most, 5% of ovarian cancer (men are also at risk for breast cancer) BRCA 1 activates homologous recombination repair of DNA double-stranded breaks). BRCA 1 is also involved in transcription and transcriptioncoupled DNA repair When its function is lost, genetic defects accumulate, which are directly responsible for cancer formation 100
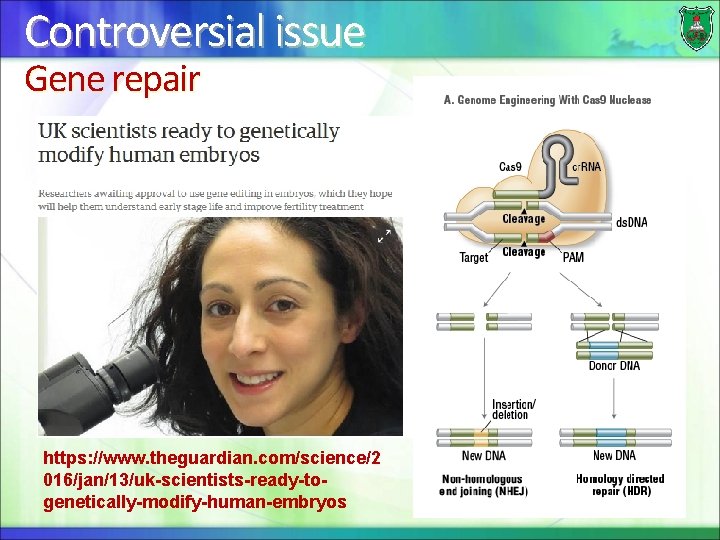
Controversial issue Gene repair https: //www. theguardian. com/science/2 016/jan/13/uk-scientists-ready-togenetically-modify-human-embryos
- Slides: 101