MOLDEN a pre and post processing program for









- Slides: 28

MOLDEN: a pre- and post- processing program for molecular and electronic structures

Interfaced via program output Gaussian GAMESS-US GAMESS-UK MOPAC, AMPAC

Interfaced via Molden Format MOLPRO ACESII MOLCAS JAGUAR ADF DALTON HONDO CADPAC

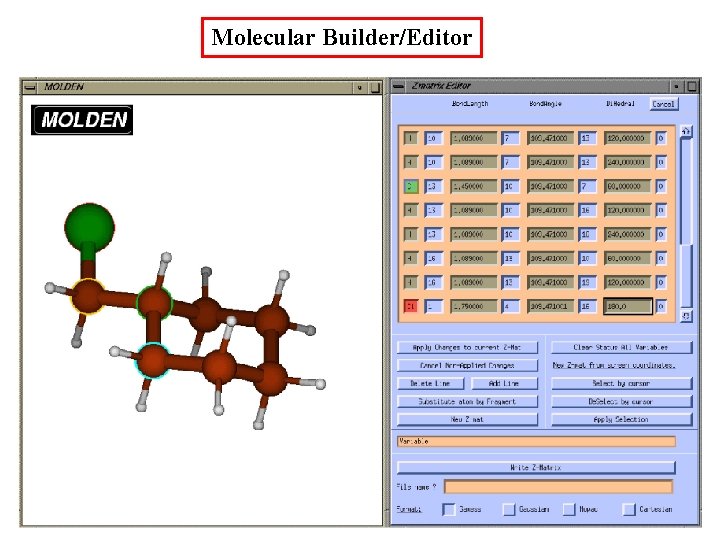
Molecular Builder/Editor

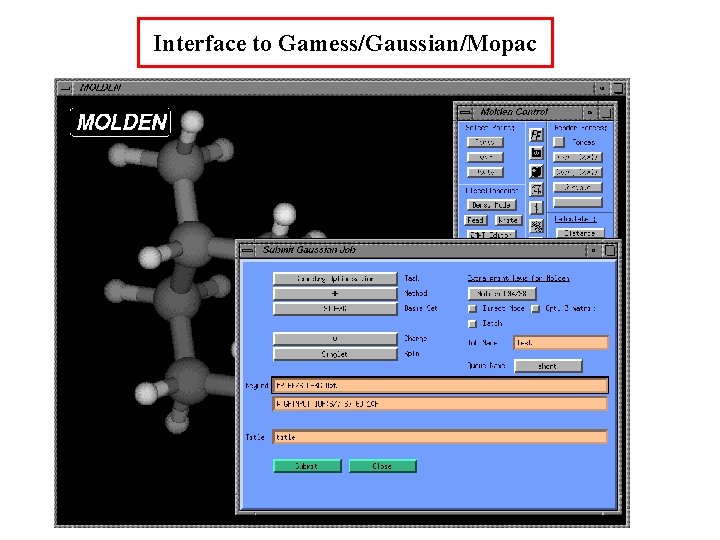
Interface to Gamess/Gaussian/Mopac

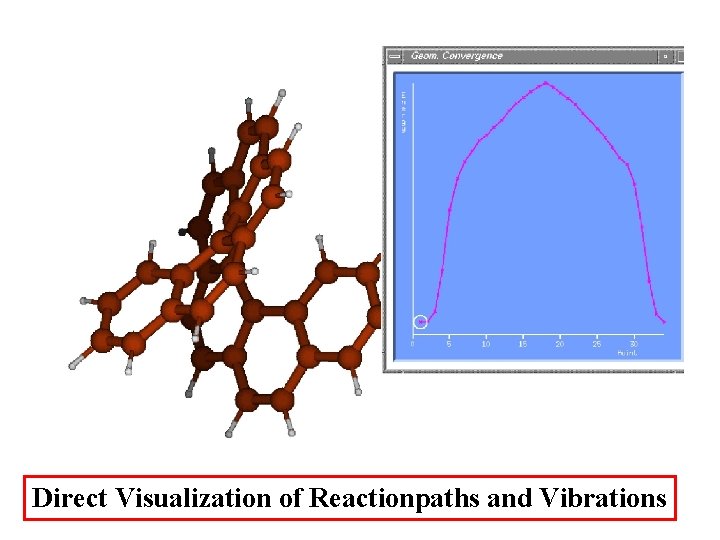
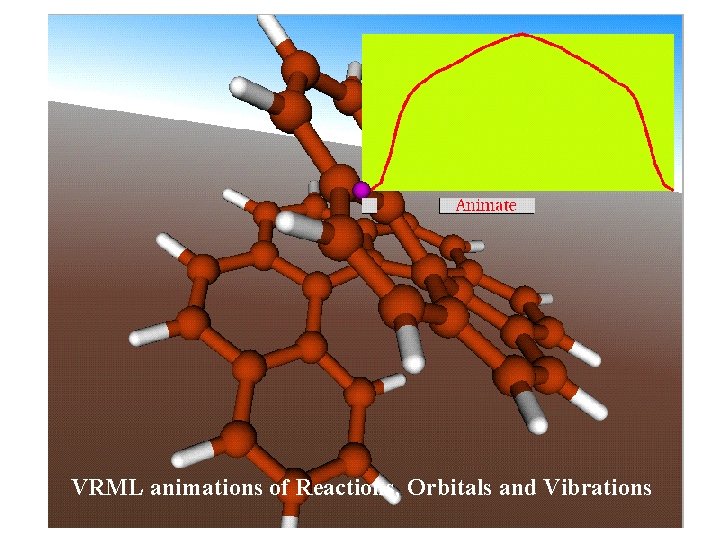
Direct Visualization of Reactionpaths and Vibrations

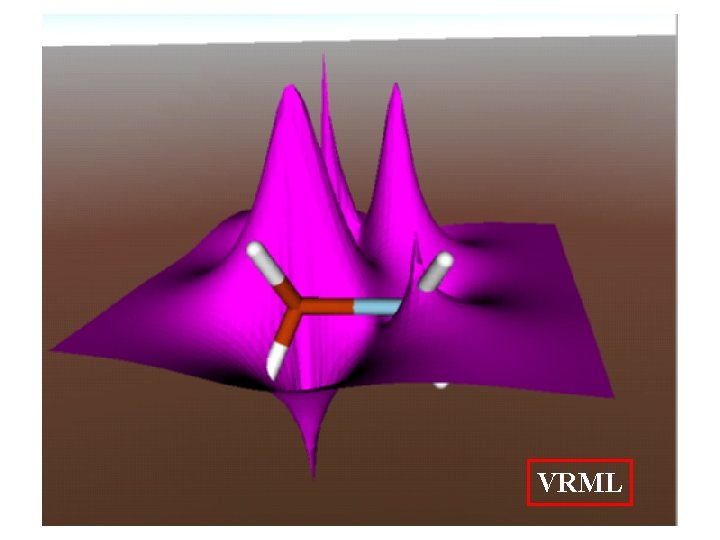
VRML animations of Reactions, Orbitals and Vibrations

Molden Calculates Electron Density: Molecular Density Molecular minus Atomic Density Laplacian of the electron density Electrostatic Potential (ESP) Multipole Moments Charges and Multipoles fit to ESP Mulliken charges, molecular dipole

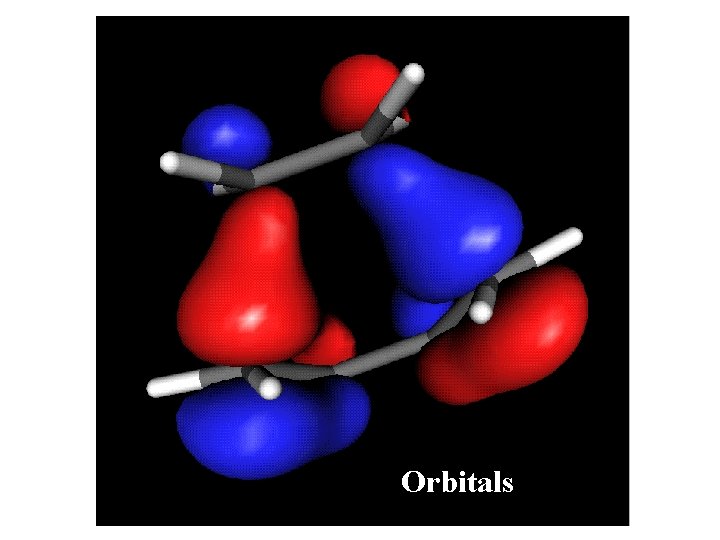
Orbitals

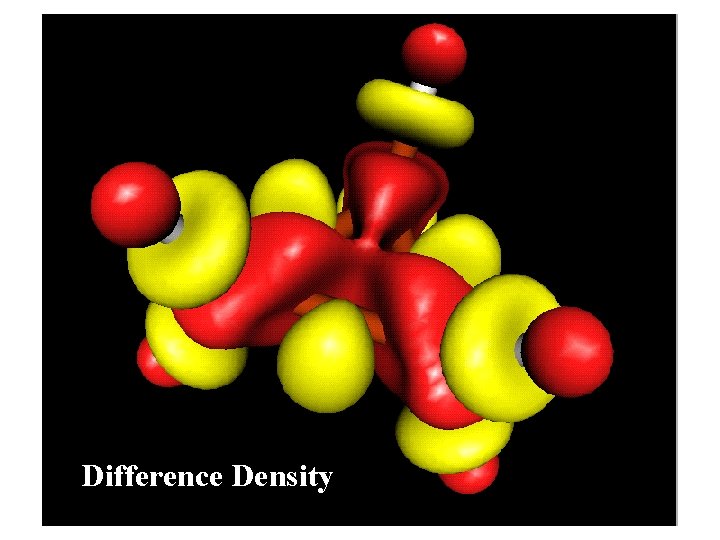
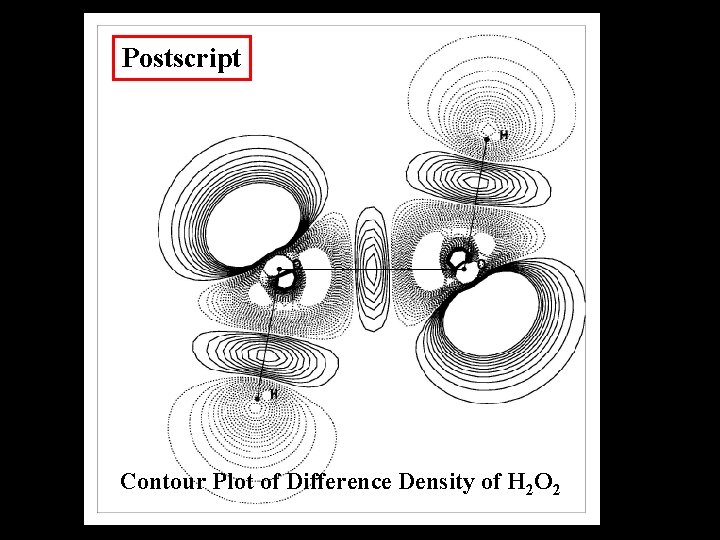
Difference Density

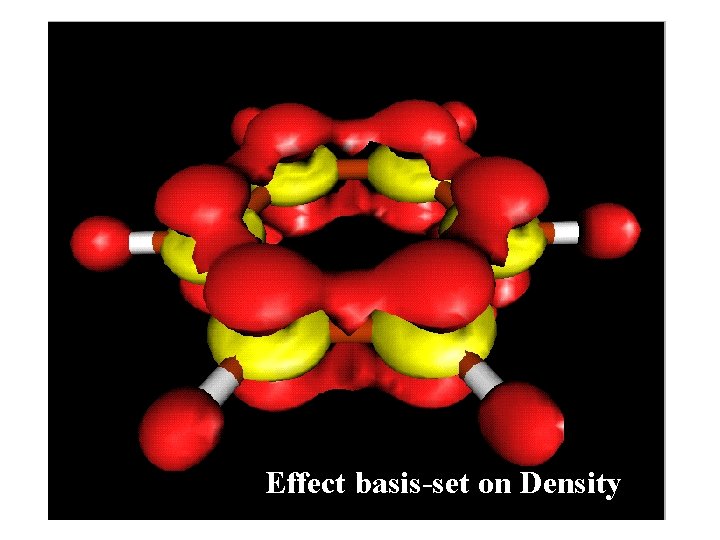
Effect basis-set on Density

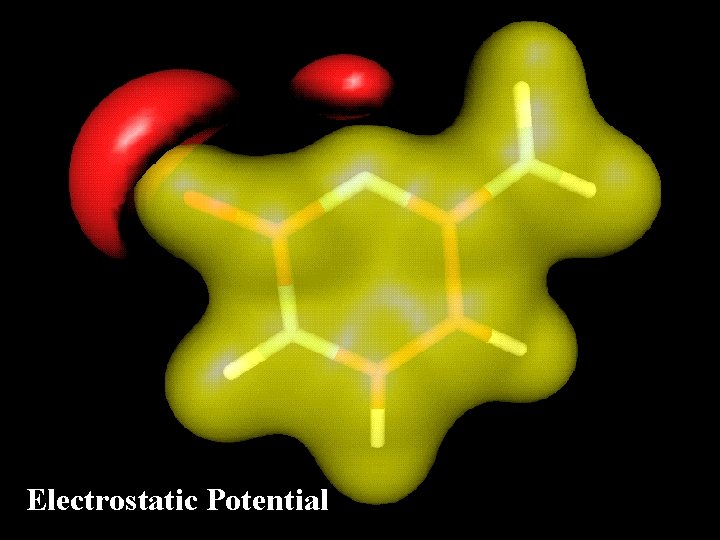
Electrostatic Potential

Electrostatic Potential mapped on an isodensity surface

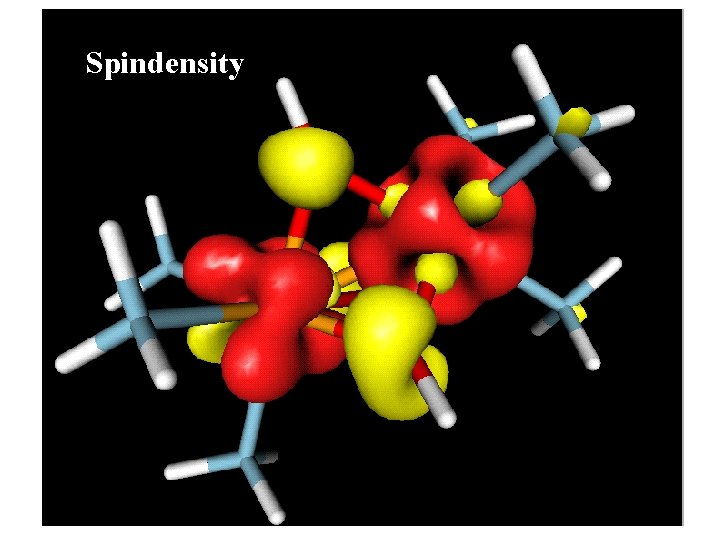
Spindensity

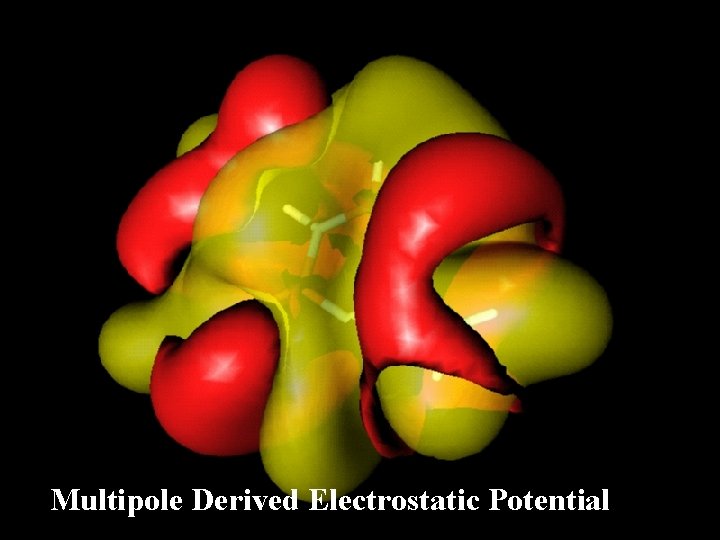
Multipole Derived Electrostatic Potential

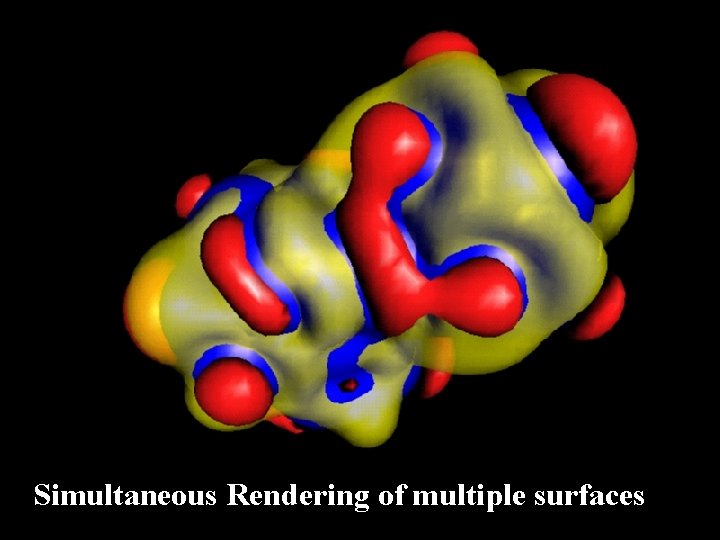
Simultaneous Rendering of multiple surfaces

Graphical Output Formats Xwindows Postscript Open. GL, VRML Pov. Ray Tek 4010, HPGL etc.

Postscript Contour Plot of Difference Density of H 2 O 2

VRML

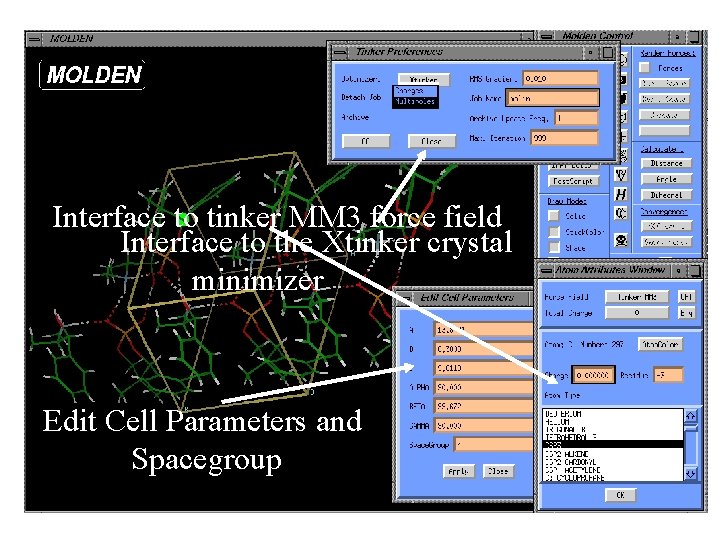
Interface to tinker MM 3 force field Interface to the Xtinker crystal minimizer Edit Cell Parameters and Spacegroup

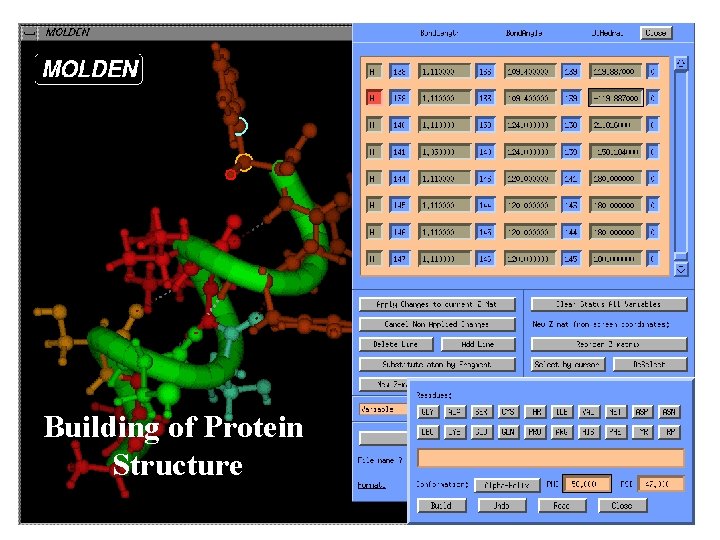
Building of Protein Structure

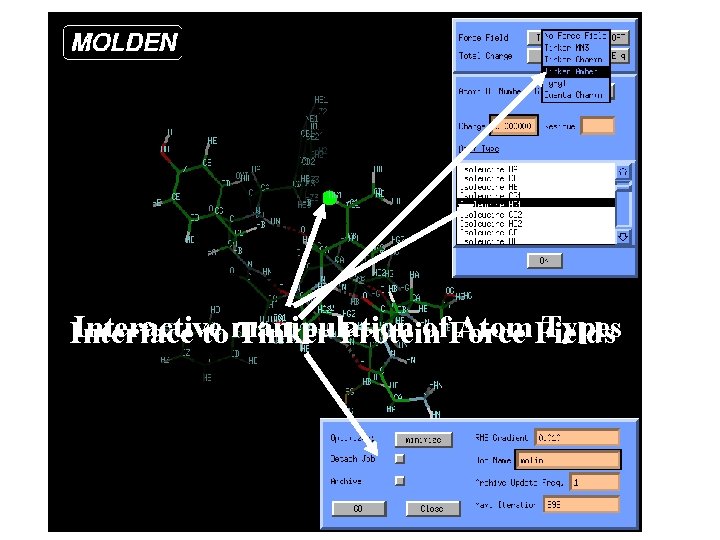
Interactive of. Force Atom Fields Types Interface to manipulation Tinker Protein

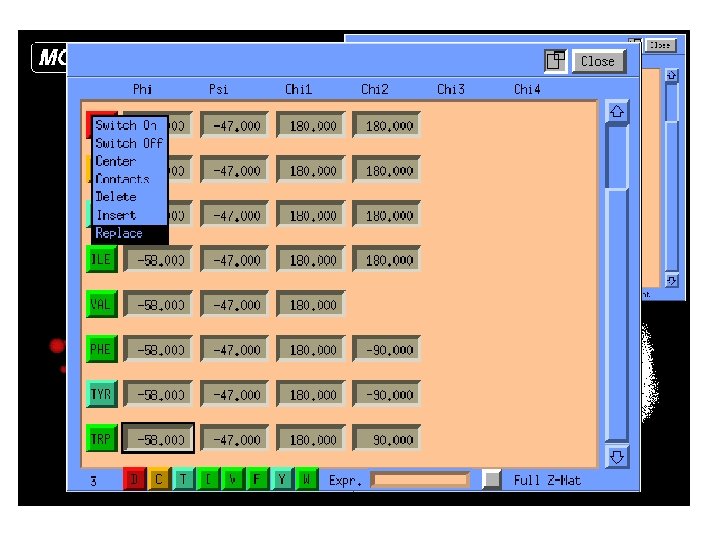
Manipulation of Protein Sidechains

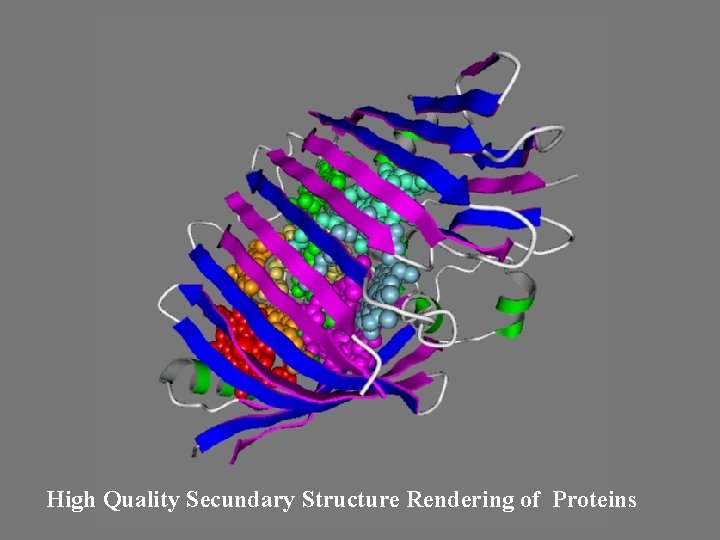
High Quality Secundary Structure Rendering of Proteins

Publications QCPE: 619 MOLDEN: A Portable Electron Density Program Published in the Journal of Computer-Aided Molecular Design: Molden: a pre- and post- processing program for molecular and electronic structures The effect of isodensity surface sampling on ESP derived charges and the effect of adding bondcenters on DMA derived charges.

Molden URL’s The Molden Home Page http: //www. cmbi. kun. nl/~schaft/molden. html Molden VRML orbital/electron density service http: //www. cmbi. kun. nl/~schaft/moldenservice. html

Molden in Web Courses/Publications Web Tutorials in Chemistry (WETCHE, CMBI) Introduction to Computational Chemistry (Frank Jensen) Practical Exercises in Quantum Chemistry (ETH) Scientific Visualization for Computational Chemistry (ACS) Computerchemische Methoden in der Physikalischen Chemie (Jena) Commodity Cluster Computing for Computational Chemistry (Adelaide)

Roundup Molden is free for the academia 3000 registered users