Modified Peptide MSMS Interpretation Karl R Clauser Broad

Modified Peptide MS/MS Interpretation Karl R. Clauser Broad Institute of MIT and Harvard Bioinformatics for Protein Identification ASMS Fall Workshop Baltimore, MD November 5 -6, 2009 Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 1

Outline • Fixed, variable, mix modifications and search space • Multiple rounds of searching • Diagnostic marker ions for modifications • Data acquisition methods specific for modifications • Ambiguity in localizing phosphorylation sites • Sample handling chemistry artifacts • Resources for masses/descriptions of known modifications Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 2

Fixed, Mix and Variable Modifications Fixed Redefine the wild type as Variable Allow 2 possibilities for an AA. Allow both in 1 spectrum if more than one location/AA. Mix Search in 2 cycles Cycle 1: all KR light Cycle 2: all KR heavy DO NOT allow both light and heavy in 1 spectrum Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 3

Variable Modifications Expand the Search Space Fixed Mods only Allow Variable Mods precursor mass filter Calculate MH+ fixed mods only tolerance filter AA composition filter Candidates passing precursor mass filter Precursor MH+ shift -256 -176 -160 -97 3 ST AA composition 1 M ? 2 ST ? 1 ST 1 M ? Karl Clauser Proteomics and Biomarker Discovery -81 -80 -32 -16 -2 -1 0 17 1 ST 1 N 1 ST 2 M 1 M 2 N 1 N * ^Q ? ? ? 1 Shift range filter Calculate MH+ Variable mod combinations tolerance filter . 05 11/30/2020 4

Methods of Constraining Allowed # of Modifications/Peptide Parent mass shift range • Spectrum Mill Max Number of mods/peptide • Sequest - all mods have same max • X!Tandem - ? • Phenyx - each mod can have different max Max Permutations of mods/peptide • Mascot - cap on permutations/peptide Candidate sequence contains sequence tags present in spectrum • Protein Pilot/Paragon Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 5

Multiple Rounds of Searching • Round 1: search all proteins • Get high confidence peptide hits • 0 -1 missed cleavages • Minimal number of variable AA modifications • Round 2: limit the search to proteins identified in round 1 • Semi-/un-specific cleavage • Increase the number of modifications • Allow for AA substitutions • Allow for undefined modifications Alternate names for similar concept • X!Tandem: refinement • Mascot: Error tolerant • Spectrum Mill: search saved hits, homology mode, unassigned single mass gap • Phenyx: 2 -rounds • Protein. Pilot/ Paragon: thorough ID, fraglet-taglet Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 6

X!Tandem - Refinement Search Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 7
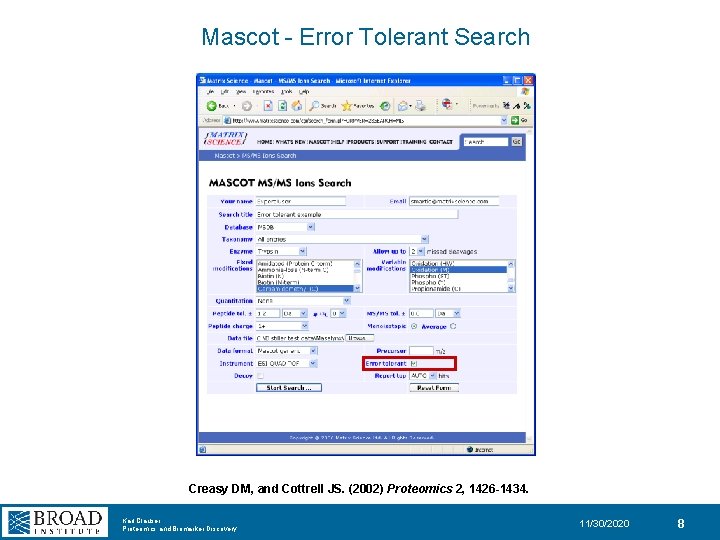
Mascot - Error Tolerant Search Creasy DM, and Cottrell JS. (2002) Proteomics 2, 1426 -1434. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 8
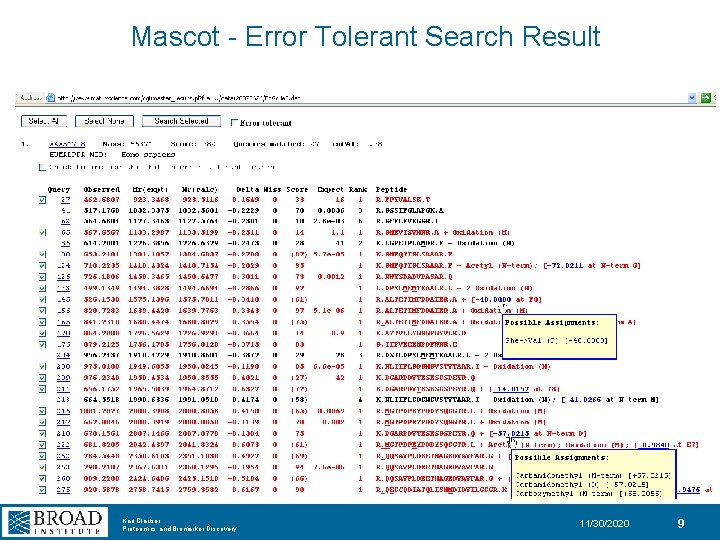
Mascot - Error Tolerant Search Result Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 9
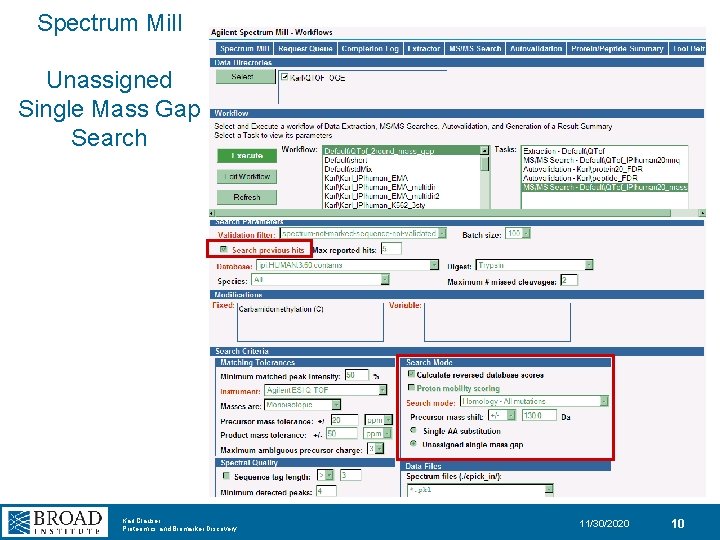
Spectrum Mill Unassigned Single Mass Gap Search Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 10
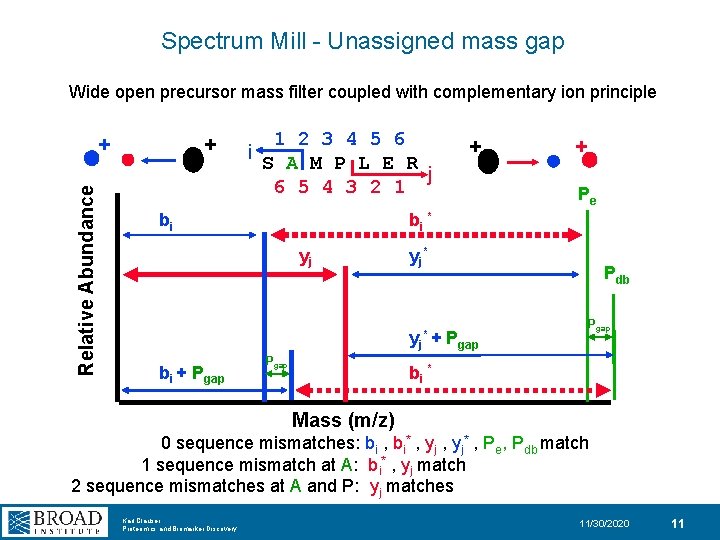
Spectrum Mill - Unassigned mass gap Wide open precursor mass filter coupled with complementary ion principle + Relative Abundance + 1 2 3 4 5 6 i S A M P L E R j 6 5 4 3 2 1 bi + Pe bi * yj yj* + bi + Pgap Pdb Pgap bi * Mass (m/z) 0 sequence mismatches: bi , bi* , yj* , Pe, Pdb match 1 sequence mismatch at A: bi* , yj match 2 sequence mismatches at A and P: yj matches Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 11
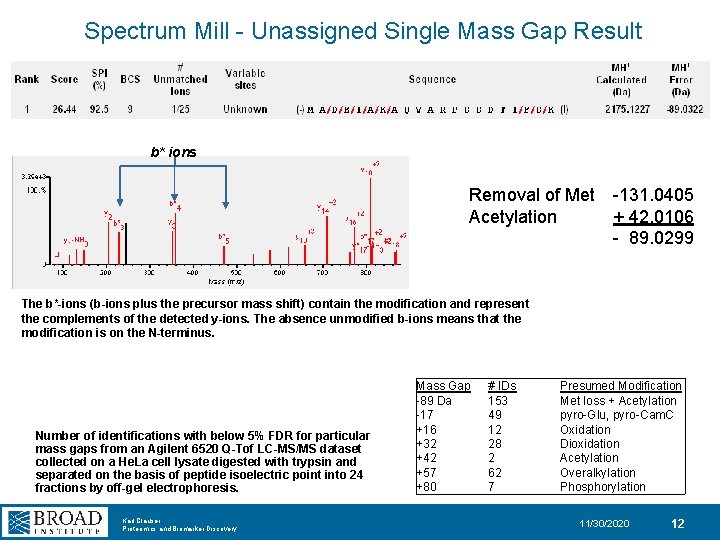
Spectrum Mill - Unassigned Single Mass Gap Result b* ions Removal of Met Acetylation -131. 0405 + 42. 0106 - 89. 0299 The b*-ions (b-ions plus the precursor mass shift) contain the modification and represent the complements of the detected y-ions. The absence unmodified b-ions means that the modification is on the N-terminus. Number of identifications with below 5% FDR for particular mass gaps from an Agilent 6520 Q-Tof LC-MS/MS dataset collected on a He. La cell lysate digested with trypsin and separated on the basis of peptide isoelectric point into 24 fractions by off-gel electrophoresis. Karl Clauser Proteomics and Biomarker Discovery Mass Gap -89 Da -17 +16 +32 +42 +57 +80 # IDs 153 49 12 28 2 62 7 Presumed Modification Met loss + Acetylation pyro-Glu, pyro-Cam. C Oxidation Dioxidation Acetylation Overalkylation Phosphorylation 11/30/2020 12
Phenyx: 2 Rounds Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 13
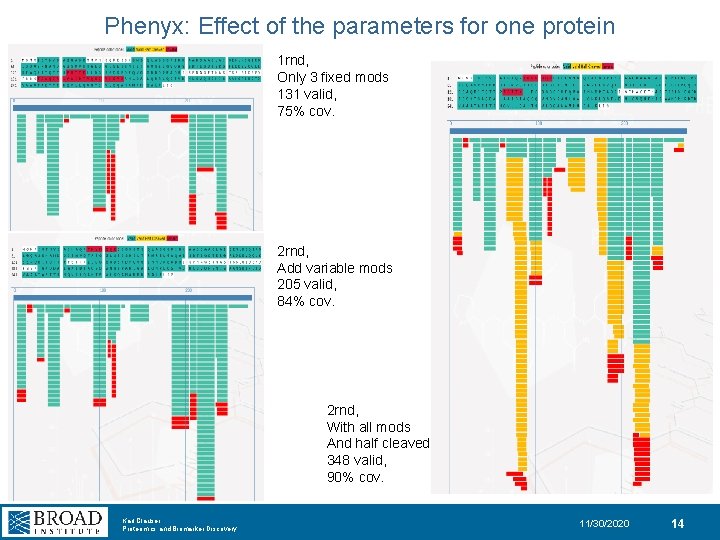
Phenyx: Effect of the parameters for one protein 1 rnd, Only 3 fixed mods 131 valid, 75% cov. 2 rnd, Add variable mods 205 valid, 84% cov. 2 rnd, With all mods And half cleaved 348 valid, 90% cov. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 14

Phenyx: Use the Annotation in Swiss. Prot, Tr. EMBL In the Feature Tables • Sequence processing annotations ◘ Removal of signal peptides ◘ Removal of transit peptides ◘ Extraction of active chains • Post-translational modifications • Sequence variants ◘ Splicing variants ◘ Sequence mutations 57292 variants / 20328 human proteins Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 15
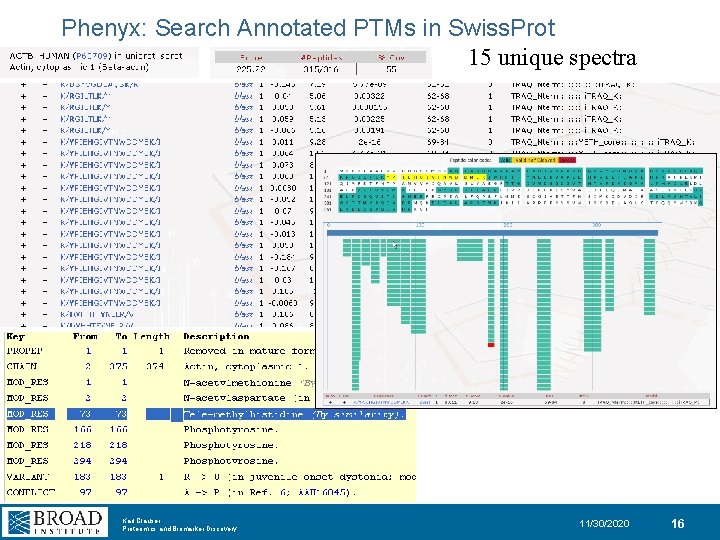
Phenyx: Search Annotated PTMs in Swiss. Prot 15 unique spectra Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 16
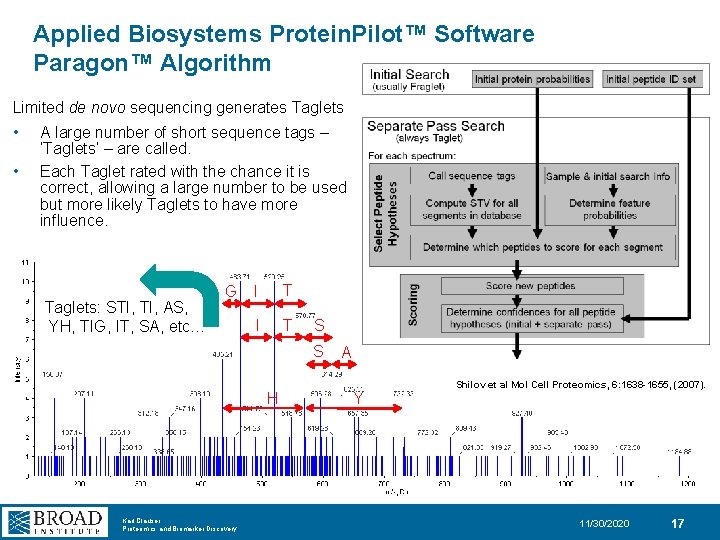
Applied Biosystems Protein. Pilot™ Software Paragon™ Algorithm Limited de novo sequencing generates Taglets • • A large number of short sequence tags – ‘Taglets’ – are called. Each Taglet rated with the chance it is correct, allowing a large number to be used but more likely Taglets to have more influence. Taglets: STI, AS, YH, TIG, IT, SA, etc… G I T S S H Karl Clauser Proteomics and Biomarker Discovery A Y Shilov et al Mol Cell Proteomics, 6: 1638 -1655, (2007). 11/30/2020 17

The Paragon™ Algorithm: Varying Search Space on a Continuum Taglets for Sequence Temperature Value (STV) Sequence Tags in Order of Decreasing Certainty: ST, TI, STI, AS, DIN, SE, EQ, NA, SEQ >DHE 3_BOVIN (P 00366) Glutamate dehydrogenase 1, mitochondrial precursor (EC 1. 4. 1. 3) (GDH) MYRYLGEALLLSRAGPAALGSASADSAALLGWARGQPAAAPQPGLVPPARRHYSEAAADREDD PNFFKMVEGFFDRGASIVEDKLVEDLKTRETEEQKRNRVRSILRIIKPCNHVLSLSFPIRRDD GSWEVIEGYRAQHSQHRTPCKGGIRYSTDVSVDEVKALASLMTYKCAVVDVPFGGAKAGVKIN PKNYTDNELEKITRRFTMELAKKGFIGPGVDVPAPDMSTGEREMSWIADTYASTIGHYDINAH ACVTGKPISQGGIHGRISATGRGVFHGIENFINEASYMSILGMTPGFGDKTFVVQGFGNVGLH A segment with cold STV SMRYLHRFGAKCITVGESDGSIWNPDGIDPKELEDFKLQHGTILGFPKAKIYEGSILEVDCDI LIPAASEKQLTKSNAPRVKAKIIAEGANGPTTPEADKIFLERNIMVIPDLYLNAGGVTVSYFE A segment with warmer STV WLNNLNHVSYGRLTFKYERDSNYHLLMSVQESLERKFGKHGGTIPIVPTAEFQDRISGASEKD The segment with the hottest STV in this protein IVHSGLAYTMERSARQIMRTAMKYNLGLDLRTAAYVNAIEKVFRVYNEAGVTFT Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 18

Controlling Search Space with the Paragon™ Algorithm • Using feature probabilities avoids include/exclude decisions and simplistic rules. • When combined with STVs, search space is dynamic by spectrum and even segment of the database. 1. 0 i. TRAQ on K, N-term Probability of Feature MMTS on C Deamidation on N, Q Oxidized M Try only most likely mods for ‘cold’ segments Try only more likely mods for ‘warm’ segments Try all mods for ‘hot’ segments in the database i. TRAQ on Y Pyroglutamic acid of E Dehydration of E, D 0 Same concept also used with digestion specificity, mass tolerances, etc. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 19

Pause for Questions Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 20

Diagnostic Marker Ions for Modifications (Immonium ions and Neutral Losses from Precursor) Mass P-98 216, P-80 P-64 P-43 204, P-203 Modification H 3 PO 4 phospho Ser, Thr phospho Tyr SOCH 4 oxidized Met carbamylated N-term N-Acetylglucosamine (Glc. NAc) m/z 98 Phosphoric Acid CID Phospho Ser Dehydroalanine Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 21

Data Acquisition Methods Specific for Modifications • ETD - Electron transfer dissociation • ECD - Electron capture dissociation • MS 3 - ion trap • Multi-stage activation - ion trap • Precursor ion scan - triple quadrupole, Q-Tof • Neutral-loss scan - triple quadrupole Review: Boersema, P; Mohammed, S; and Heck, A. Phosphopeptide fragmentation and analysis by mass spectrometry. J. Mass. Spectrom. 2009, 44, 861– 878. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 22
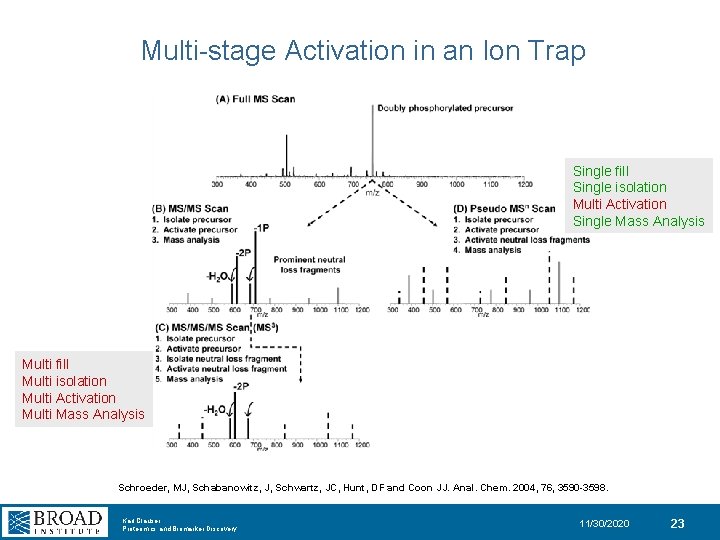
Multi-stage Activation in an Ion Trap Single fill Single isolation Multi Activation Single Mass Analysis Multi fill Multi isolation Multi Activation Multi Mass Analysis Schroeder, MJ, Schabanowitz, J, Schwartz, JC, Hunt, DF and Coon JJ. Anal. Chem. 2004, 76, 3590 -3598. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 23
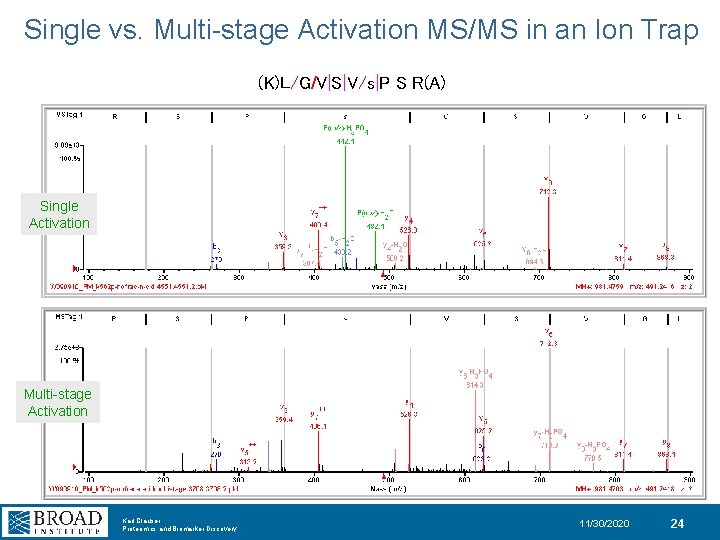
Single vs. Multi-stage Activation MS/MS in an Ion Trap (K)L/G/V|S|V/s|P S R(A) Single Activation Multi-stage Activation Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 24
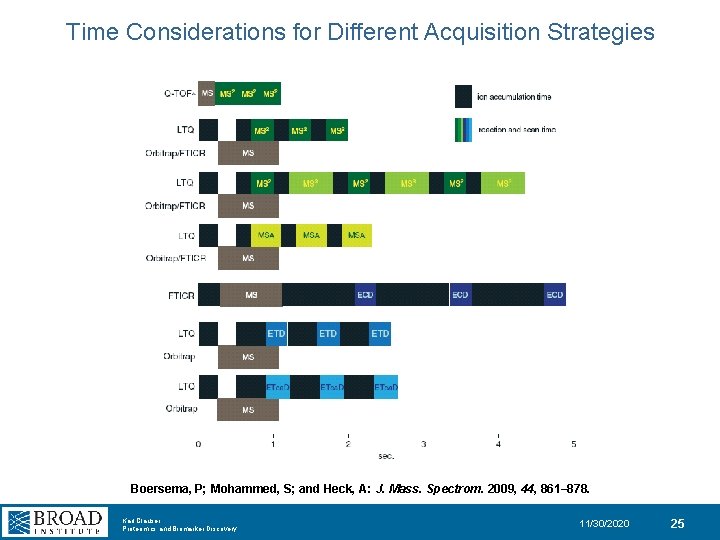
Time Considerations for Different Acquisition Strategies Boersema, P; Mohammed, S; and Heck, A: J. Mass. Spectrom. 2009, 44, 861– 878. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 25

O-Glc. NAcylation • Addition of a single sugar residue: N-Acetylglucosamine (Glc. NAc) to serine or threonine residues of nuclear and cytoplasmic proteins. • Present in all multi-cellular organisms • Different from ‘conventional’ glycosylation: • Inside the cell • Transient modification • Enzymes responsible for addition and removal of modification • i. e. analogous to phosphorylation • O-Glc. NAc modification and phosphorylation interact / affect each other • Modification is involved in cellular response to nutritional and other stresses • Clear links to Diabetes and Alzheimer Disease and elevated in cancer. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 26
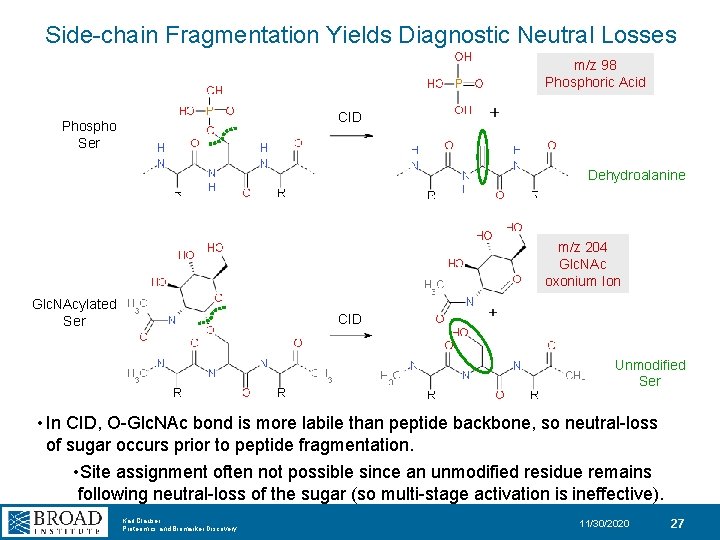
Side-chain Fragmentation Yields Diagnostic Neutral Losses m/z 98 Phosphoric Acid CID Phospho Ser Dehydroalanine m/z 204 Glc. NAc oxonium Ion Glc. NAcylated Ser CID Unmodified Ser • In CID, O-Glc. NAc bond is more labile than peptide backbone, so neutral-loss of sugar occurs prior to peptide fragmentation. • Site assignment often not possible since an unmodified residue remains following neutral-loss of the sugar (so multi-stage activation is ineffective). Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 27
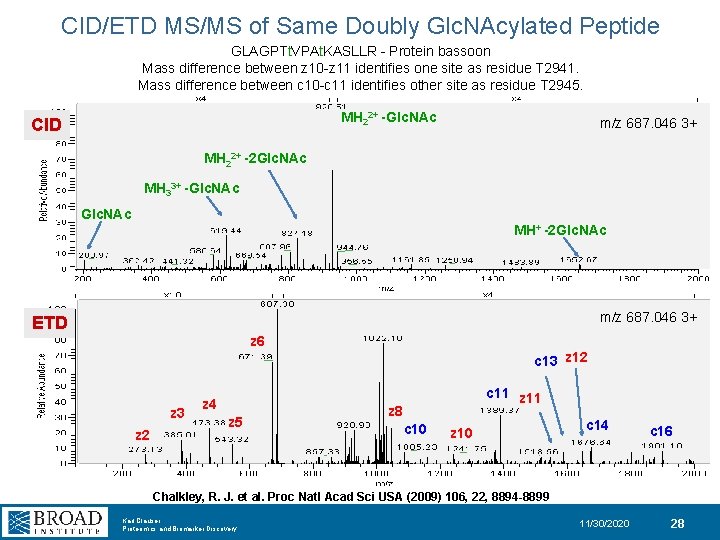
CID/ETD MS/MS of Same Doubly Glc. NAcylated Peptide GLAGPTt. VPAt. KASLLR - Protein bassoon Mass difference between z 10 -z 11 identifies one site as residue T 2941. Mass difference between c 10 -c 11 identifies other site as residue T 2945. MH 22+ -Glc. NAc CID m/z 687. 046 3+ MH 22+ -2 Glc. NAc MH 33+ -Glc. NAc MH+ -2 Glc. NAc m/z 687. 046 3+ ETD z 6 c 13 z 12 z 3 z 2 z 4 z 5 c 11 z 8 c 10 z 10 c 14 c 16 Chalkley, R. J. et al. Proc Natl Acad Sci USA (2009) 106, 22, 8894 -8899 Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 28
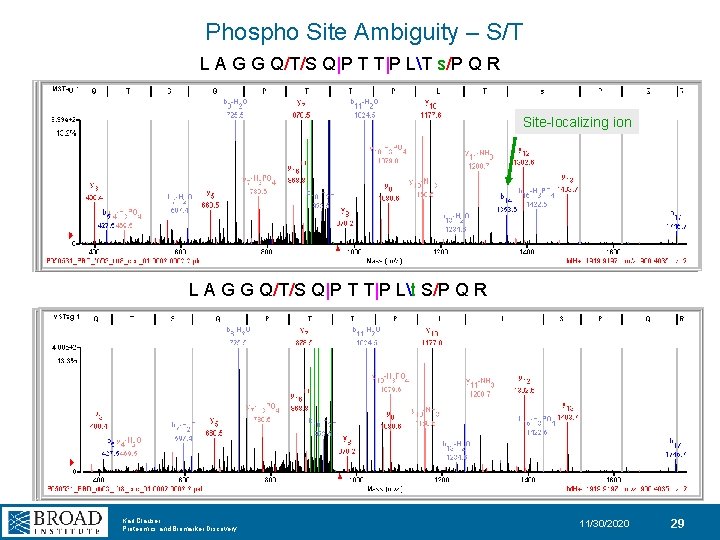
Phospho Site Ambiguity – S/T L A G G Q/T/S Q|P T T|P LT s/P Q R Site-localizing ion L A G G Q/T/S Q|P T T|P Lt S/P Q R Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 29
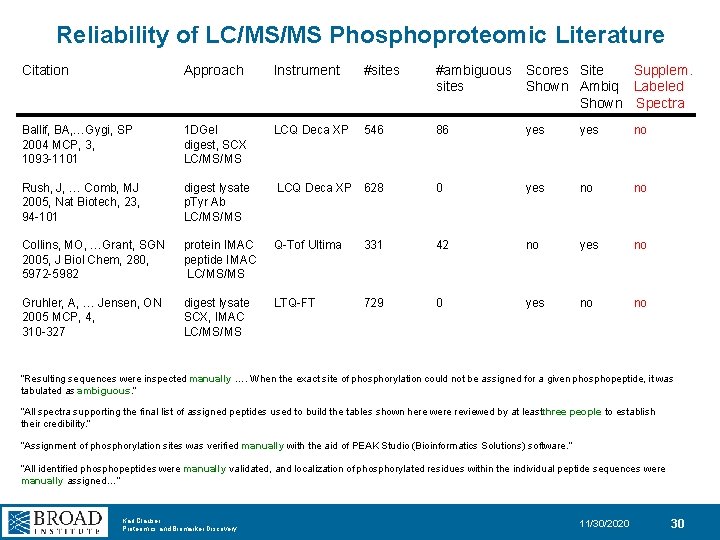
Reliability of LC/MS/MS Phosphoproteomic Literature Citation Approach Instrument #sites #ambiguous Scores Site Supplem. sites Shown Ambiq Labeled Shown Spectra Ballif, BA, …Gygi, SP 2004 MCP, 3, 1093 -1101 1 DGel digest, SCX LC/MS/MS LCQ Deca XP 546 86 yes no Rush, J, … Comb, MJ 2005, Nat Biotech, 23, 94 -101 digest lysate p. Tyr Ab LC/MS/MS LCQ Deca XP 628 0 yes no no Collins, MO, …Grant, SGN 2005, J Biol Chem, 280, 5972 -5982 protein IMAC peptide IMAC LC/MS/MS Q-Tof Ultima 331 42 no yes no Gruhler, A, … Jensen, ON 2005 MCP, 4, 310 -327 digest lysate SCX, IMAC LC/MS/MS LTQ-FT 729 0 yes no no “Resulting sequences were inspected manually …. When the exact site of phosphorylation could not be assigned for a given phosphopeptide, it was tabulated as ambiguous. ” “All spectra supporting the final list of assigned peptides used to build the tables shown here were reviewed by at leastthree people to establish their credibility. ” “Assignment of phosphorylation sites was verified manually with the aid of PEAK Studio (Bioinformatics Solutions) software. ” “All identified phosphopeptides were manually validated, and localization of phosphorylated residues within the individual peptide sequences were manually assigned…” Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 30

MCP draft Guideline for publishing PTM data http: //www. mcponline. org/ III. POST-TRANSLATIONAL MODIFICATIONS Studies focusing on posttranslational modifications require specialized methodology and documentation to assign the presence and the site(s) of modification. No current MS data analysis software is infallible in the automatic assignment of modification sites in peptides, and these analyses are particularly error prone when multiple possible sites within a peptide are being utilized. For these reasons, additional documentation supporting assignment of PTMs is required. In addition to the tabular presentation(s) of the data described in guideline II: • The site(s) of modification within each peptide sequence must be clearly presented. • An indication of the certainty of localization for each PTM: The manner in which the modification was located (by computation or manually) and a description of the software used, if any. • A justification for any localization score threshold employed. • Ambiguous assignments: Peptides containing ambiguous PTM site localizations must be listed in a separate table from those with unambiguous site localizations. In cases where there are multiple modification sites and at least one is ambiguous, then these peptides should be listed with the ambiguous assignments. Ambiguous assignments must be clearly labeled as such. Examples of ambiguities include: • Modified peptides in which one or more modification sites are ambiguous. • Instances where the peptide sequence is repeated in the same protein so the specific modification site cannot be assigned. • Instances in which the same peptide is repeated in multiple proteins, e. g. paralogs and splice variants (See also Section IV). • Isobaric modifications (e. g. , acetylation vs. trimethylation, phosphorylation vs. sulfonation etc), where the possibilities may not be distinguished. Examples of methods able to distinguish between these include mass spectrometric approaches such as accurate mass determination, observation of signature fragment ions (e. g. m/z 79 vs. m/z 80 in negative ion mode for assignment of phosphorylation over sulfation), or biological or chemical strategies. • Annotated, mass labeled spectra: Spectra for ALL modified peptides must be either submitted to a public repository or accompany the manuscript as described in guideline II. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 31
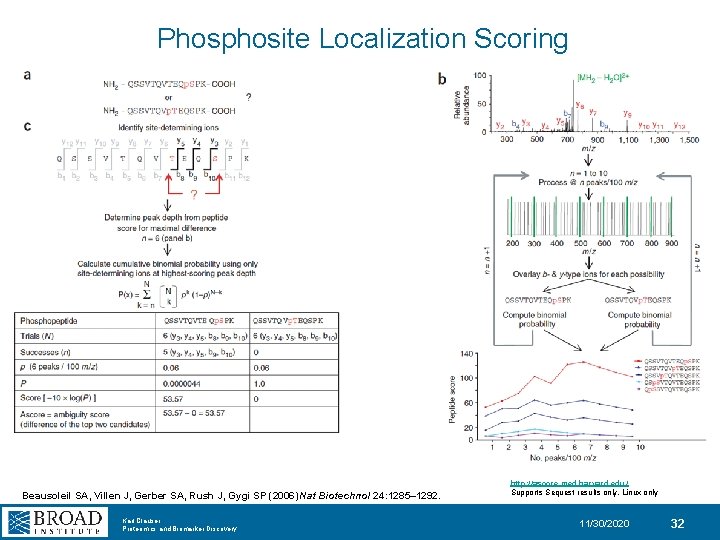
Phosphosite Localization Scoring Beausoleil SA, Villen J, Gerber SA, Rush J, Gygi SP (2006) Nat Biotechnol 24: 1285– 1292. Karl Clauser Proteomics and Biomarker Discovery http: //ascore. med. harvard. edu/ Supports Sequest results only, Linux only 11/30/2020 32
![Phosphosite Localization Scoring P = (k!/[n!(n-k)!] [pk] [(1 -p) (n-k) ]) = (k!/[n!(n-k)!] [0. Phosphosite Localization Scoring P = (k!/[n!(n-k)!] [pk] [(1 -p) (n-k) ]) = (k!/[n!(n-k)!] [0.](http://slidetodoc.com/presentation_image_h/c4b8aa678ed51d221a7b3838a5d98962/image-33.jpg)
Phosphosite Localization Scoring P = (k!/[n!(n-k)!] [pk] [(1 -p) (n-k) ]) = (k!/[n!(n-k)!] [0. 04 k] [(0. 96) (n-k) ]) PTM score = -10 x log (P) p: 0. 04 - use the 4 most intense fragment ions per 100 m/z units n: total num possible b/y ions in the observed mass range for all possible combinations of PO 4 sites in a peptide k: number of peaks matching n Olsen, J. V. ; Blagoev, B. ; Gnad, F. ; Macek, B. ; Kumar, C. ; Mortensen, P. ; Mann, M. Cell (2006), 127 (3), 635– 48. Olsen, J. V. , and Mann, M. Proc. Natl. Acad. Sci. USA. (2004) 101, 13417– 13422. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 33

True Probability or Just Effective Scores? Peak selection assumptions • All regions of spectrum equally likely • multiply charged fragements below precursor • some 100 -300 m/z values not possible dipeptide AA combinations • Tall and short peak intensities equally diagnostic Fragment ion type assumptions • All ion types equally probable • Neutral losses ignored, y-H 3 P 04, y-H 2 O Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 34
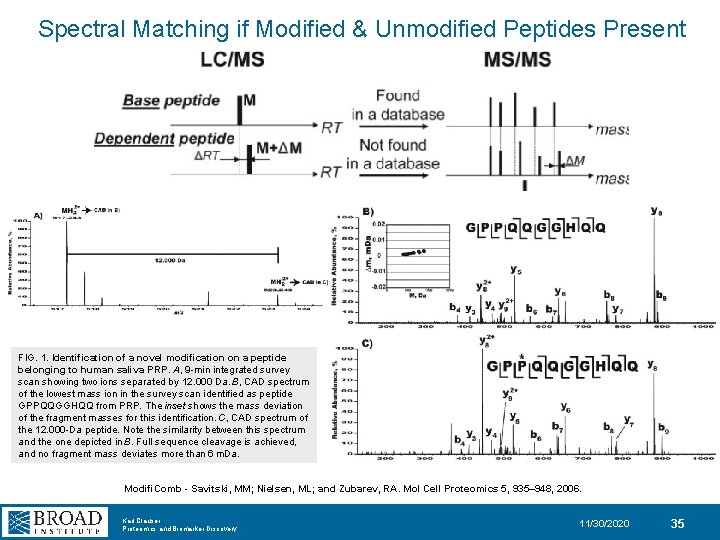
Spectral Matching if Modified & Unmodified Peptides Present FIG. 1. Identification of a novel modification on a peptide belonging to human saliva PRP. A, 9 -min integrated survey scan showing two ions separated by 12. 000 Da. B, CAD spectrum of the lowest mass ion in the survey scan identified as peptide GPPQQGGHQQ from PRP. The inset shows the mass deviation of the fragment masses for this identification. C, CAD spectrum of the 12. 000 -Da peptide. Note the similarity between this spectrum and the one depicted in B. Full sequence cleavage is achieved, and no fragment mass deviates more than 6 m. Da. Modifi. Comb - Savitski, MM; Nielsen, ML; and Zubarev, RA. Mol Cell Proteomics 5, 935– 948, 2006. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 35

Software Tools Specialized for Identifying Modifications and Localizing Sites Ascore Beausoleil SA, Villen J, Gerber SA, Rush J, Gygi SP (2006) Nat Biotechnol. 24, 1285– 1292. Max. Quant Cox J, Mann M. (2008) Nat Biotechnol. 26, 1367 - 1372. Olsen JV, Blagoev B, Gnad F, Macek B, Kumar C, Mortensen P, Mann M. (2006) Cell. 127, 635– 48. Inspect, MS-Alignment, PTMFinder Tanner S, Payne S, Dasari S, Shen Z, Wilmarth PA, David L, Loomis WF, Briggs SP, Bafna V. (2008) J Proteome Res. 7, 170– 181. Payne S, Yau M, Smolka MB, Tanner S, Zhou H, Bafna V. (2008) J Proteome Res. 7, 3373– 3381. Tsur D, Tanner S, Zandi E, Bafna V, Pevzner P. (2005) Nat Biotechnol. 23, 1562– 1567. Tanner S, Shu H, Frank A, Wang LC, Zandi E, Mumby M, Pevzner P, Bafna V. (2005) Anal Chem. 77, 4626 -4639. Phospho. Score Ruttenberg BE, Pisitkun T, Knepper MA, Hoffert JD. (2008) J Proteome Res. 7, 3054 -9. Debunker Lu B, Ruse C, Xu T, Park SK, Yates J 3 rd. (2007) Anal Chem. 79, 1301 -10. Slo. Mo - ETD/ECD Bailey CM, Sweet SM, Cunningham DL, Zeller M, Heath JK, Cooper HJ. (2009) J Proteome Res. 8, 1965 -71. Modifi. Comb Savitski MM, Nielsen ML, Zubarev RA. (2006) Mol Cell Proteomics. 5, 935– 48. Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 36

Pause for Questions Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 37

Expect Woes & Nuisances Sample Handling Chemistry • Carbamylation • Deamidation • pyro. Glutamic acid • pyro. Carbamidomethyl Cys • Oxidized Met • Cys alkylation reagent Karl Clauser Proteomics and Biomarker Discovery +43 +1 -17 +16 +x nterm, Lys N -> D nterm Q nterm C M n-term, W urea in digest buffer sample in acid gels side reaction 11/30/2020 38
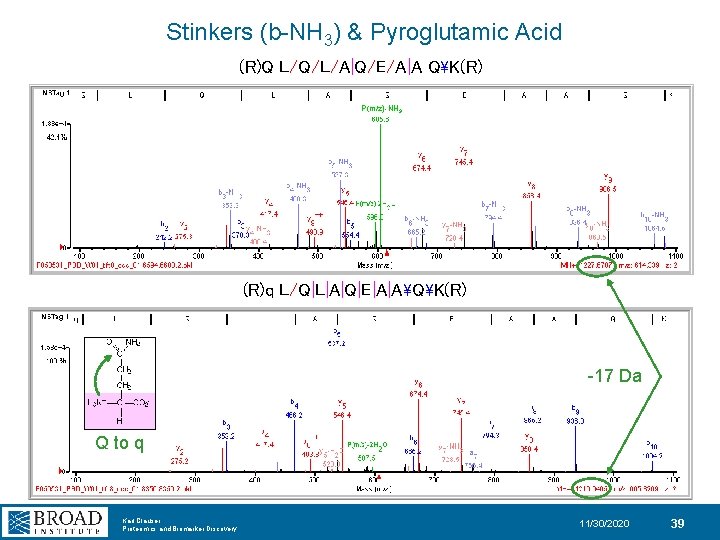
Stinkers (b-NH 3) & Pyroglutamic Acid (R)Q L/Q/L/A|Q/E/A|A QK(R) P(m/z)-NH 3 (R)q L/Q|L|A|Q|E|A|AQK(R) -17 Da Q to q Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 39

Deamidation of Asn +1 Da Asn –NH + O = Asp ionsource. com Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 40
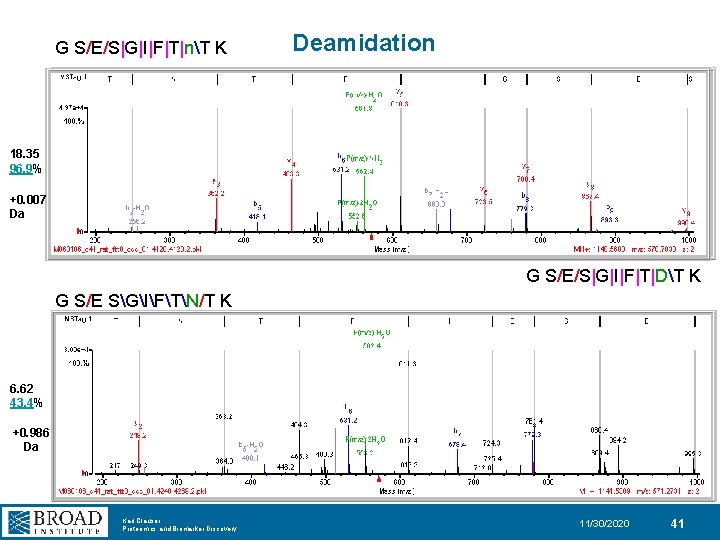
G S/E/S|G|I|F|T|nT K Deamidation 18. 35 96. 9% +0. 007 Da G S/E/S|G|I|F|T|DT K G S/E SGIFTN/T K 6. 62 43. 4% +0. 986 Da Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 41

Carbamylation from Urea in Digest Buffer +43 Da CNHO +43 Da Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 42
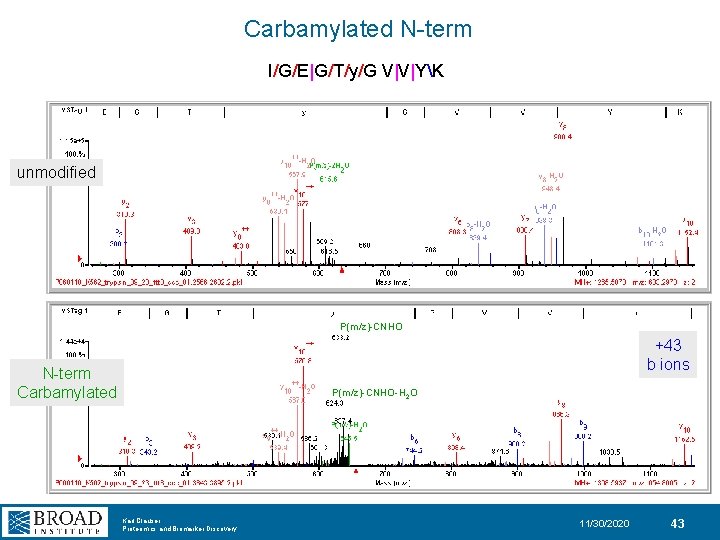
Carbamylated N-term I/G/E|G/T/y/G V|V|YK unmodified P(m/z)-CNHO +43 b ions N-term Carbamylated P(m/z)-CNHO-H 2 O Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 43

Unimod Resource for Masses of Modifications http: //www. unimod. org/modifications_list. php Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 44
Delta Mass Resource for Masses of Modifications http: //www. abrf. org/index. cfm/dm. home Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 45
RESID Resource for Masses of Modifications http: //www. ebi. ac. uk/RESID/ Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 46

Acknowledgements Broad Institute of MIT and Harvard Steven Carr Philipp Mertins Pierre-Alain Binz Robert Chalkley John Cottrell Chris Miller Sean Seymour Gene. Bio University of California San Francisco Matrix Science Agilent Technologies Applied Biosystems Karl Clauser Proteomics and Biomarker Discovery Phenyx O-Glc. NAc Mascot Spectrum Mill Protein Pilot, Paragon 11/30/2020 47

i. PRG-2010: Proteome Informatics Research Group Study Phosphopeptide Identification In this study, an LC-MS/MS dataset from a lysate digested with trypsin and enriched for phosphopeptides using strong cation exchange fractionation followed by immobilized metal affinity chromatography (SCX/IMAC) will be provided. Participants are asked to return a list of identified peptides and localized phosphorylation sites Requests to participate must be submitted by e-mail to i. PRG 2010@gmail. com prior to Monday, November 30, 2009. Please include the words “i. PRG Study 2010 request” in the subject line and provide contact name and affiliation in the body of the message. http: //www. abrf. org Karl Clauser Proteomics and Biomarker Discovery 11/30/2020 48
- Slides: 48