Modeling regulatory networks in cells using Bayesian networks

Modeling regulatory networks in cells using Bayesian networks Golan Yona Department of Computer Science Cornell University cs 726

Outline • Regulatory networks • Expression data • Bayesian Networks • What • Why • How • Learning networks from expression data • Using Bayesian networks to analyze expression data (Friedman et al) cs 726
Regulatory networks KEGG Regulatory Pathways cs 726
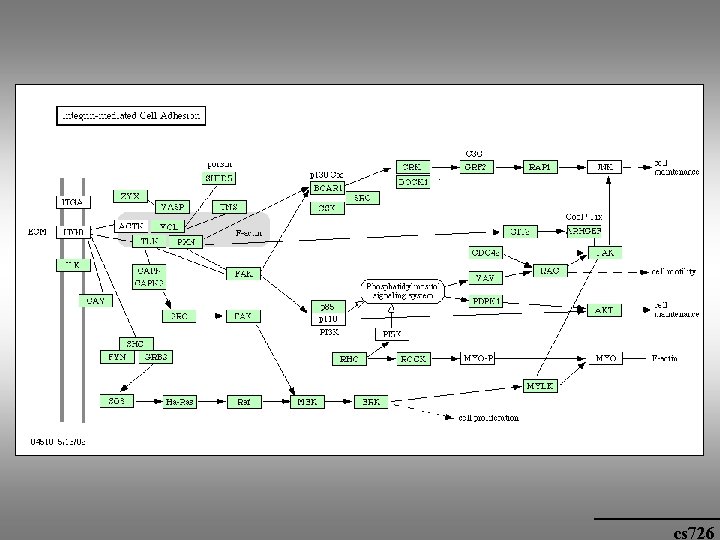
cs 726
Metabolic pathways KEGG Metabolic Pathways cs 726
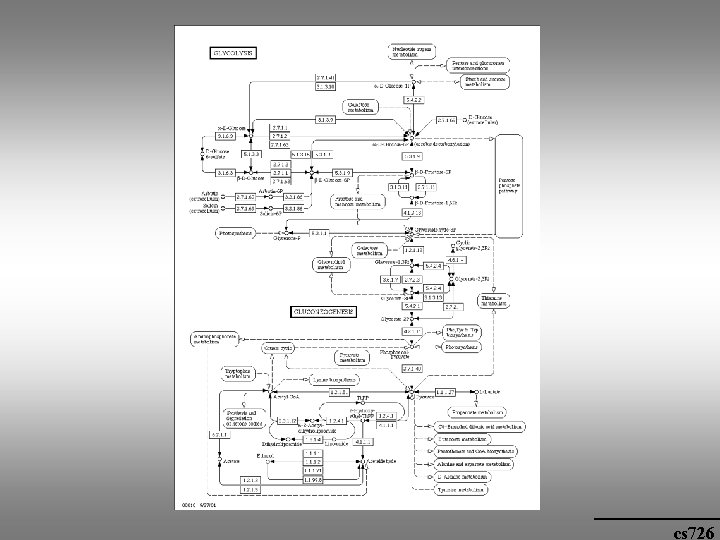
cs 726

Expression Arrays • Measure the expression levels of thousands of genes in a cell under specific conditions (e. g. cell cycle) simultaneously • Each cell has the same genomic data but different subsets of proteins are being expressed in different cells and at the same cell under different conditions. • Protein level is controlled by controlling – – – transcription initiation m. RNA transcription m. RNA transport splicing post-translational modifications degradation of m. RNA and proteins. • Microarray measure the level of m. RNA, thus providing an indirect evidence for the control of protein levels cs 726

Micro Spotting pin cs 726

cs 726

• Some are over-expressed (red), some under-expressed (green) measured with respect to a control group of genes (“fixed” genes) • Different pathways are activated under different conditions cs 726

Goals • Recover protein interactions and sub-networks that correspond to regulatory networks in the cell. • Basic assumption: some genes are dependent on others while others exhibit independence or conditional independence • The means: Bayesian networks. Capable of modeling the statistical dependencies between different variables (genes) • Different from clustering analysis. . • Applicable when the dependency between genes is “local” • Problems: data is noisy, partial, sometimes misleading (translation, activation), not enough to ensure statistically significant models, time scale cs 726

Bayesian Networks • A compromise between the assumption of complete dependency and complete conditional independence (Naïve Bayes) • Less constraining yet still tractable • We know something about the statistical dependencies between features but not necessarily about the type of the underlying distributions cs 726
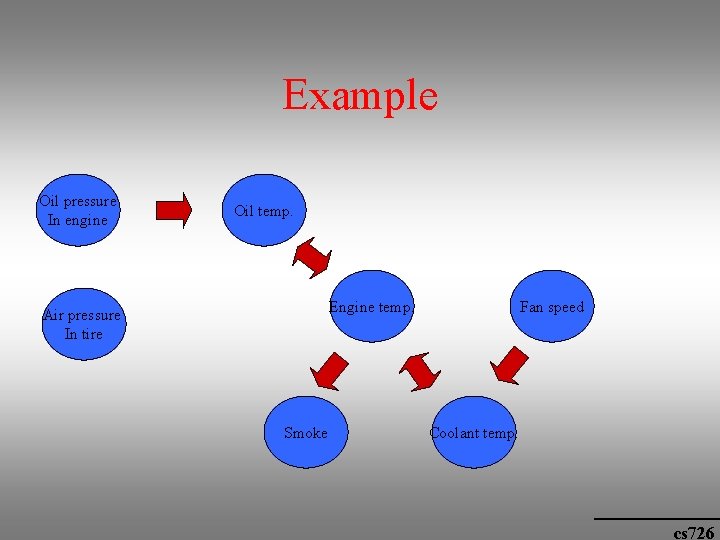
Example Oil pressure In engine Oil temp. Engine temp. Air pressure In tire Smoke Fan speed Coolant temp. cs 726

Bayesian Networks • Also called belief nets • A graph description of dependencies and independencies between variables. Each node corresponds to a variable (gene). • The graph is directed and acyclic • The variables are discrete • A variable can take on a value from a set of values {a 1, a 2, …} e. g. on/off • The probability of a specific value P(ai) and i P(ai) = 1 • A link joining node A to node C is directional and represents the set of conditional probabilities P(cj/ai) – causality (the probability that C is on when A is off) • The network is described in terms of its nodes and edges and the conditional probability distributions associated with each node. cs 726
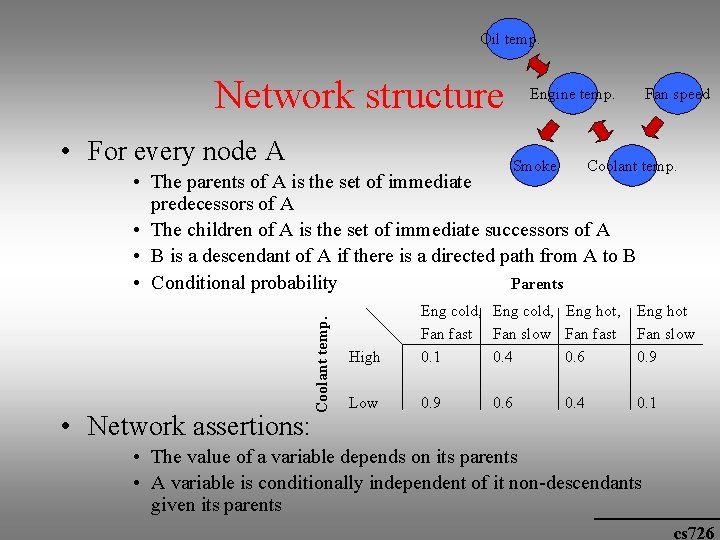
Oil temp. Network structure • For every node A Engine temp. Smoke Fan speed Coolant temp. • Network assertions: Coolant temp. • The parents of A is the set of immediate predecessors of A • The children of A is the set of immediate successors of A • B is a descendant of A if there is a directed path from A to B • Conditional probability Parents High Eng cold, Eng hot, Fan fast Fan slow Fan fast 0. 1 0. 4 0. 6 Eng hot Fan slow 0. 9 Low 0. 9 0. 1 0. 6 0. 4 • The value of a variable depends on its parents • A variable is conditionally independent of it non-descendants given its parents cs 726

Calculating the probability of an assignment • The network describes the joint probability distribution of all variables (some conditionally independent and some are not) • Depends on the structure! • The probability of a specific assignment of values y 1, y 2, …yn for the variables Y 1, Y 2, …, Yn This is the likelihood of the data given the model. All you need to know is. . cs 726

Learning Bayesian network from data • Given the data set with specific assignments for variables (on/off for each gene), how can we find the most probable network structure that explains the data (the best “match” to the data)? • How to quantify a match? • Note that there are two aspects of the network that we need to learn – Structure (nodes, edges) – Conditional probability distributions • Common strategy: assign a score to each network G cs 726

• Common strategy: assign a score to each network G Likelihood prior • Pick the network that maximizes the score cs 726

Learning • The likelihood of the data given the model is estimated by averaging over all possible assignments of parameters (conditional probabilities) to G • Summation over all possible assignments for conditional probabilities. The major contribution is from the set estimated from the data • Given a specific structure, for every node we lookup its parents and calculate the empirical conditional probability distribution cs 726

Model selection • The second term (log prior) is a measure for the complexity of the model (through uncertainty) • Occam razor: entia non sunt multiplicanda praeter necessitatem (thou shall not multiply entities) MDL principle • In the papers discussed here it is being ignored cs 726

In search for the best network • In theory: test different structures, calculate the probability of assignment to variables for each network structure, and output the network that maximizes the likelihood of the data given the network. • Impossible in practice – the number of possible networks over n genes is • For the yeast genome with 6000 genes this is > 105, 000 cs 726

Possible solution • Apply a heuristic local greedy search: Start with a random network and locally improve it, by testing perturbations over the original structure. • Test one edge at a time, by adding, removing or reversing the edge, and testing its affect on the score. If the score improves - accept cs 726

How to learn from expression data • Two types of features learned from multiple networks • First - a gene Y is in the Markov blanket of X (two genes are involved in the same biological process. No other gene mediates the dependence) • Problem of unobserved variables that can intermediate the interaction • Second type – a gene X is ancestor of Y (based on all networks that are learned) cs 726

Application to the Yeast Cell cycle data • Expression level measurements for 6177 genes along different time points in six cell cycles – altogether 76 measurements for each gene • Only 800 genes vary during cell cycle and 250 cluster into 8 fairly distinct classes. • Networks are learned for the 800 genes • Confidence values based on the set of networks learned from different bootstrap sets cs 726
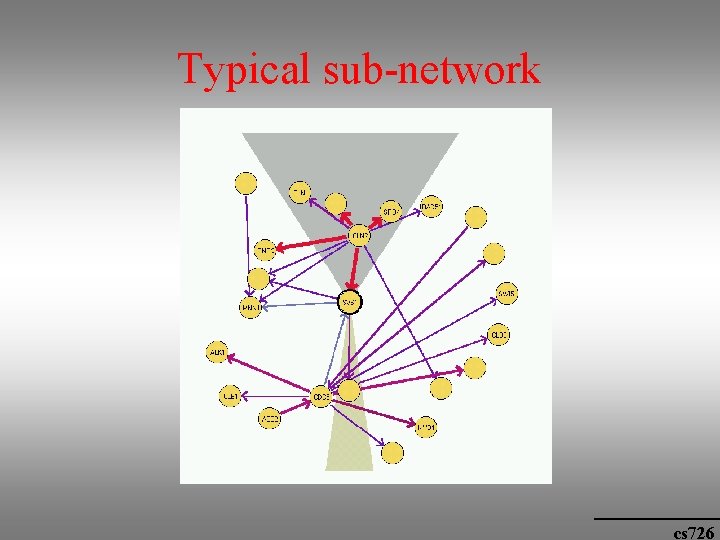
Typical sub-network cs 726

Biological significance • Order relations: there a few dominant genes that appear before many others, e. g. genes that are involved in cell cycle control and initiation. cs 726
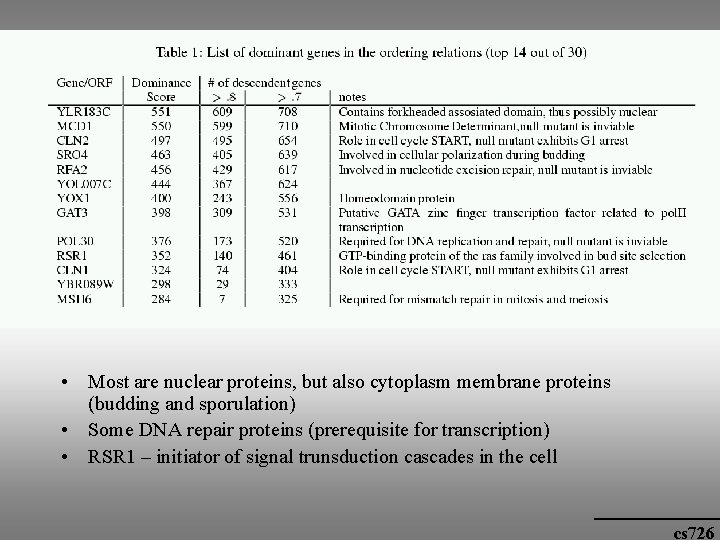
• Most are nuclear proteins, but also cytoplasm membrane proteins (budding and sporulation) • Some DNA repair proteins (prerequisite for transcription) • RSR 1 – initiator of signal trunsduction cascades in the cell cs 726

Biological significance • Markov connection: functionally related cs 726
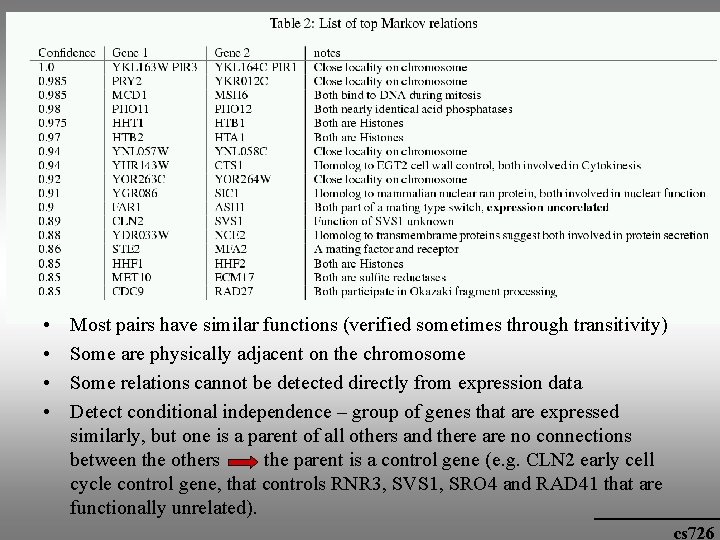
• • Most pairs have similar functions (verified sometimes through transitivity) Some are physically adjacent on the chromosome Some relations cannot be detected directly from expression data Detect conditional independence – group of genes that are expressed similarly, but one is a parent of all others and there are no connections between the others the parent is a control gene (e. g. CLN 2 early cell cycle control gene, that controls RNR 3, SVS 1, SRO 4 and RAD 41 that are functionally unrelated). cs 726

Conclusions • A powerful tool, but – not enough data – Computational problems – Learning algorithms – Authors decompose networks into basic elements again • Many possible extensions cs 726
- Slides: 30