Mining Semantic Descriptions of Bioinformatics Web Resources from

Mining Semantic Descriptions of Bioinformatics Web Resources from the Literature Hammad Afzal, Robert Stevens, Goran Nenadic School of Computer Science University of Manchester G. Nenadic@manchester. ac. uk

Motivation o A number of bioinformatics tools and resources available for service use and composition n guessimate is 3000+ Web Services publically available n how to find a service, what is out there to use? n provenance? o Semantic annotation of bioinformatics services n annotate functional capabilities n e. g. Taverna, my. Grid, my. Experiment, EBI, Bio. MOBY o Not only services and tools n databases, repositories, corpora

Motivation o Manual curation n e. g. my. Grid, Bio. Catalogue etc. n e. g. Taverna/Feta: only ~15 -20% functionally described n backlog – and the number of services is growing o Annotations combine n textual descriptions n ontological mappings
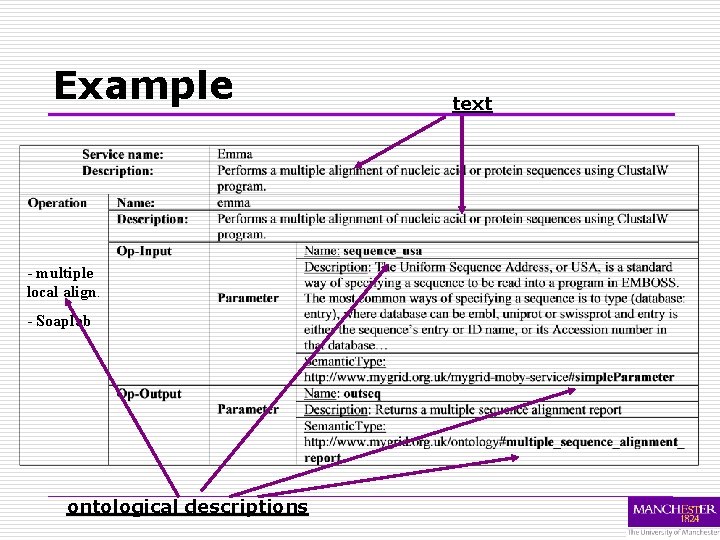
Example - multiple local align. - Soaplab ontological descriptions text

Bio. Catalogue o Single registration point for Web Service providers o Single search site for scientists and developers o Place where the community can find contacts and meet the experts and maintainers of these services o Community-sourced annotation, expert oversee o Mixed annotations: free text, tags, controlled vocabularies, community ontologies
Bio. Catalogue Beta version at http: //beta. biocatalogue. org/ Launch June 2009 at ISMB

Our approach o Collect service semantic descriptions by extracting and integrating information from text resources n full text bioinformatics journal publications o Approach: n identify descriptors that are used for service and resource annotations n locate them in text n infer the annotations o textual evidence and mappings to an ontology

The rest of the talk o Methodology n mining bioinformatics terminology n extraction of service description profiles o Experiments and results n semi-automated curation o What next?
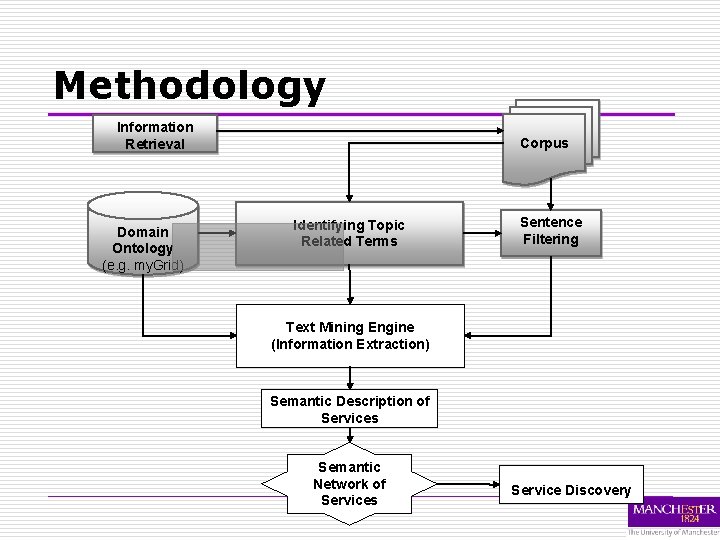
Methodology Information Retrieval Domain Ontology (e. g. my. Grid) Corpus Identifying Topic Related Terms Sentence Filtering Text Mining Engine (Information Extraction) Semantic Description of Services Semantic Network of Services Service Discovery
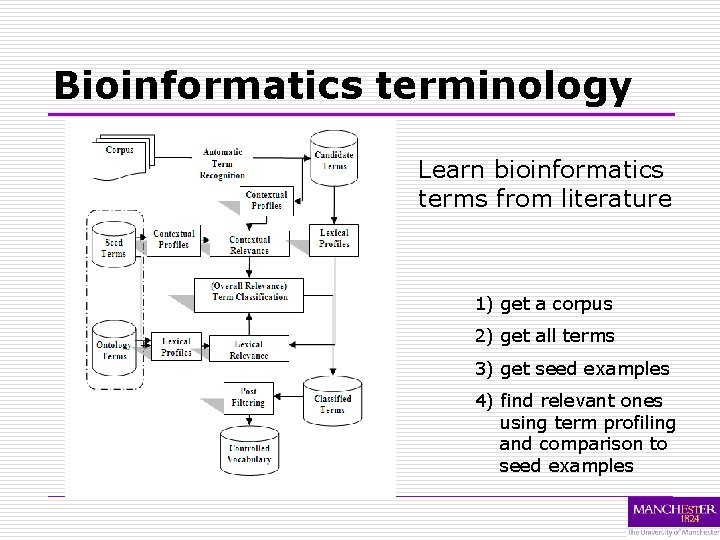
Bioinformatics terminology Learn bioinformatics terms from literature 1) get a corpus 2) get all terms 3) get seed examples 4) find relevant ones using term profiling and comparison to seed examples

Bioinformatics terminology o Use seed terms to bootstrap n e. g. known descriptors used in existing service descriptions, either in literature or service repositories o 250 terms identified, manual pruning after automatic term recognition n examples of lexical constituents and textual behaviour (pragmatics) o lexical profiling o contextual profiling

Bioinformatics terminology o Lexical profiling n what is in the name o Contextual profiling n characterise sentences in which terms appear (nouns, verbs and context-patterns) o Comparing candidate term profiles to n average seed term n best-match
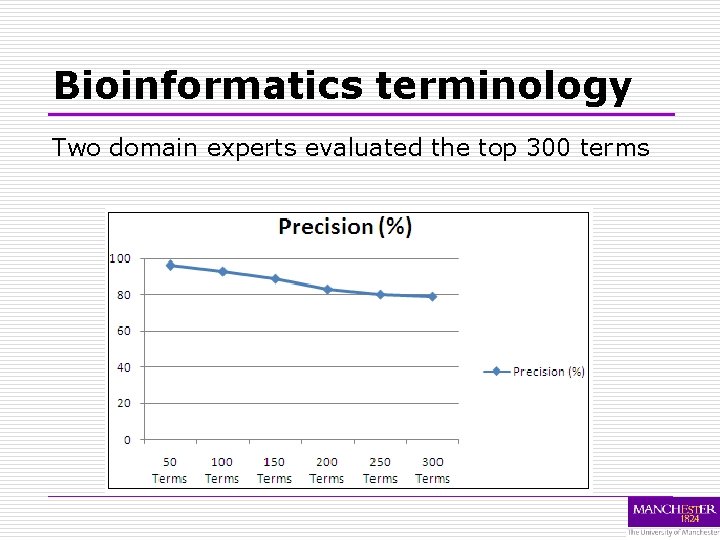
Bioinformatics terminology Two domain experts evaluated the top 300 terms

Semantic classes – my. Grid o Informatics concepts n general concepts of data, data structures, databases, metadata o Bioinformatics concepts n domain-specific data sources and algorithms for searching and analysing data n e. g. Smith-Waterman algorithm
Semantic classes – my. Grid o Molecular biology concepts n higher level concepts used to describe bioinformatics data types, used as inputs and outputs in services n e. g. protein sequence, nucleic acid sequence o Task concepts n generic tasks a service operation can perform n e. g. retrieving, displaying, aligning
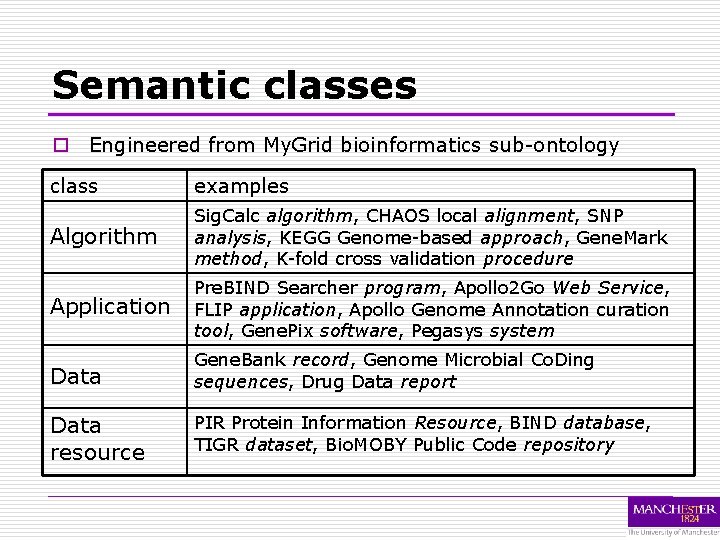
Semantic classes o Engineered from My. Grid bioinformatics sub-ontology class examples Algorithm Sig. Calc algorithm, CHAOS local alignment, SNP analysis, KEGG Genome-based approach, Gene. Mark method, K-fold cross validation procedure Application Pre. BIND Searcher program, Apollo 2 Go Web Service, FLIP application, Apollo Genome Annotation curation tool, Gene. Pix software, Pegasys system Data Gene. Bank record, Genome Microbial Co. Ding sequences, Drug Data report Data resource PIR Protein Information Resource, BIND database, TIGR dataset, Bio. MOBY Public Code repository
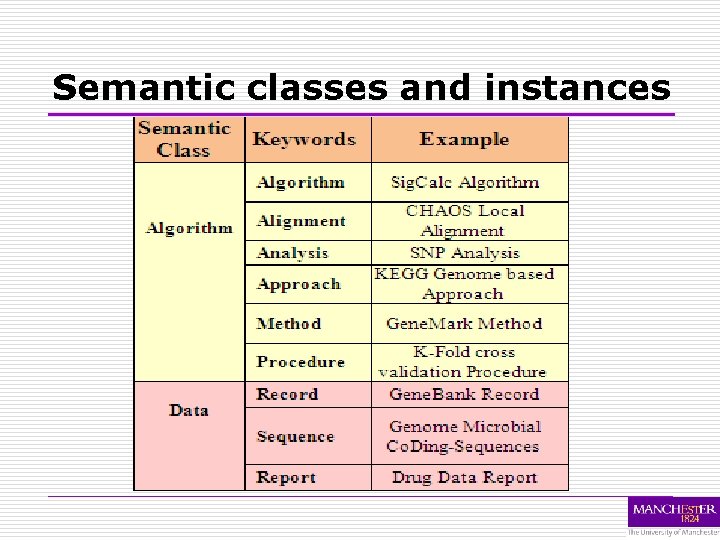
Semantic classes and instances
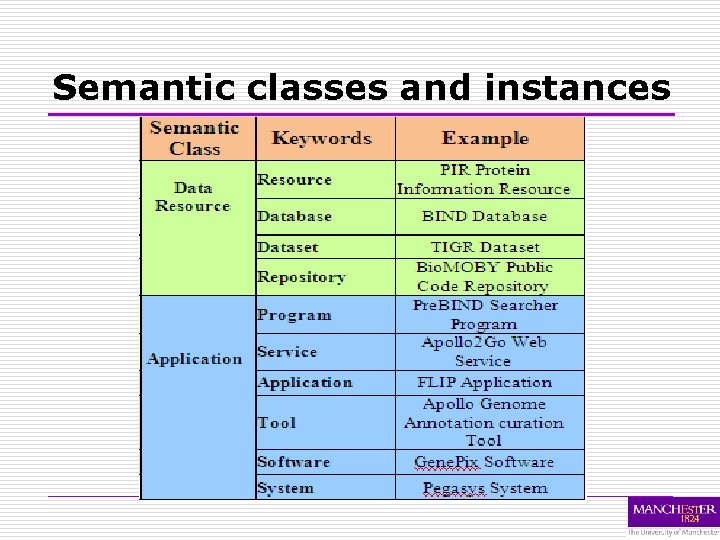
Semantic classes and instances

Service mentions o Named-entity recognition (NER) task o Recognition of service mentions using n terminological (semantic) heads of automatically recognised terms o Apollo 2 Go Web Service is an Application o BIND database is a Data source o assign the corresponding semantic class n Hearst patterns (co-ordinations, appositions, enumerations, etc. )

Semantic descriptors o Recognition of phrases depicting semantic roles n used to describe services o Flexible dictionary look-up n terms from my. Grid ontology n terms/noun phrases from existing descriptions of bioinformatics resources (collected from Taverna and other Web service providers).
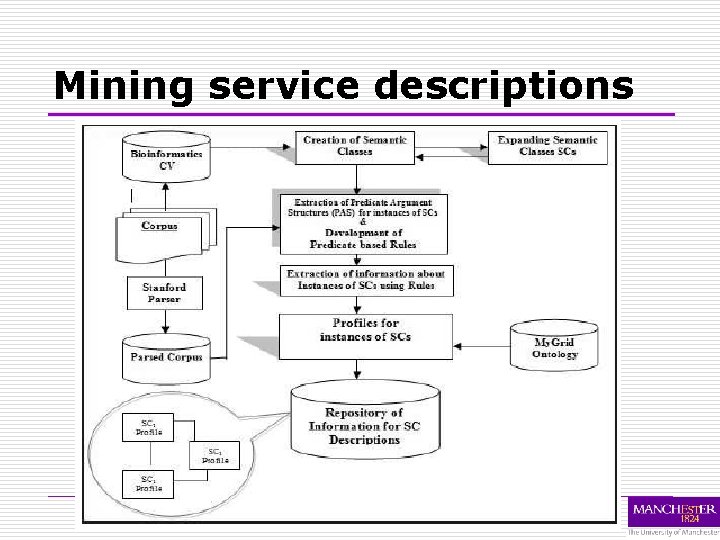
Mining service descriptions
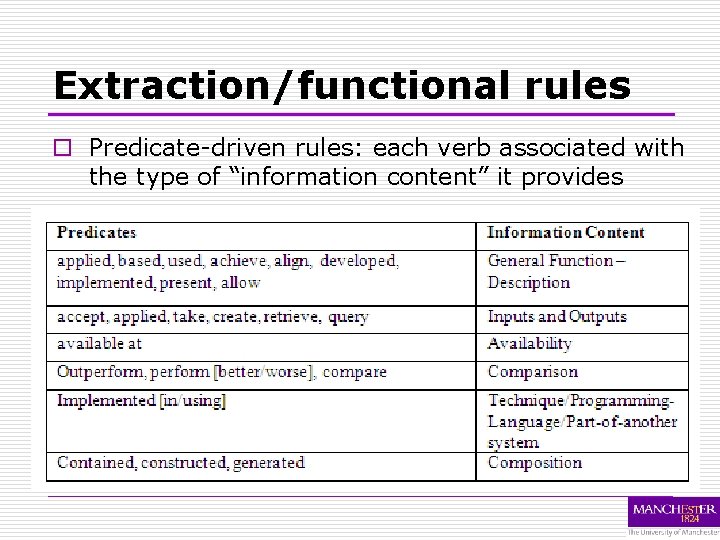
Extraction/functional rules o Predicate-driven rules: each verb associated with the type of “information content” it provides

Extraction/functional rules o Manually designed predicate-driven rules: Subject (Arg) – Verb (Predicate) – Object (Arg) o Applied on dependency parsed sentences n n Stanford parser no phrase structures complex sentences information in sub-clause

Extraction/functional rules o Phrase structures identified and integrate with the dependency o Predicate-dependent rules applied to extract specific ‘content’ and profile the services o Profiles collated for all mentions n service name variation
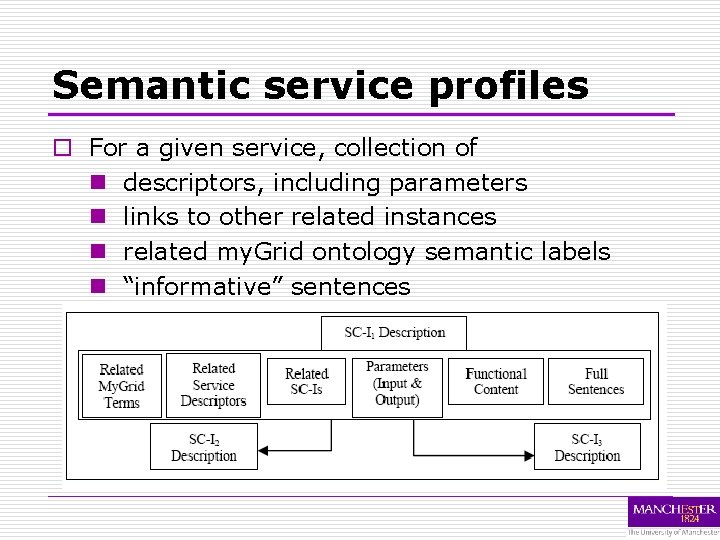
Semantic service profiles o For a given service, collection of n descriptors, including parameters n links to other related instances n related my. Grid ontology semantic labels n “informative” sentences

Example – Gene. Class o Descriptors
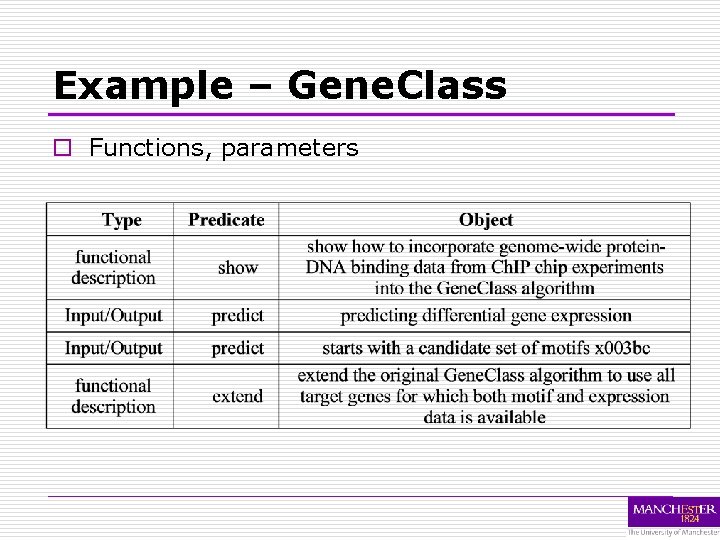
Example – Gene. Class o Functions, parameters

Example – Gene. Class o Sentences We extend the original Gene. Class algorithm to use all target genes for which both motif and expression data is available. In order to study different aspects of target gene regulation we use different sets of motifs and parents with the Gene. Class algorithm. The Gene. Class algorithm for predicting differential gene expression starts with a candidate set of motifs; representing known or putative regulatory element sequence patterns and a candidate set of regulators or parent. SS.

Experiments o 2120 BMC Bioinformatics articles n full-text articles before March 2008 o Service descriptors dictionary n 471 descriptors from my. Grid/Feta n 450 descriptors collected from other bioinformatics service/tools providers o 108 predicates used
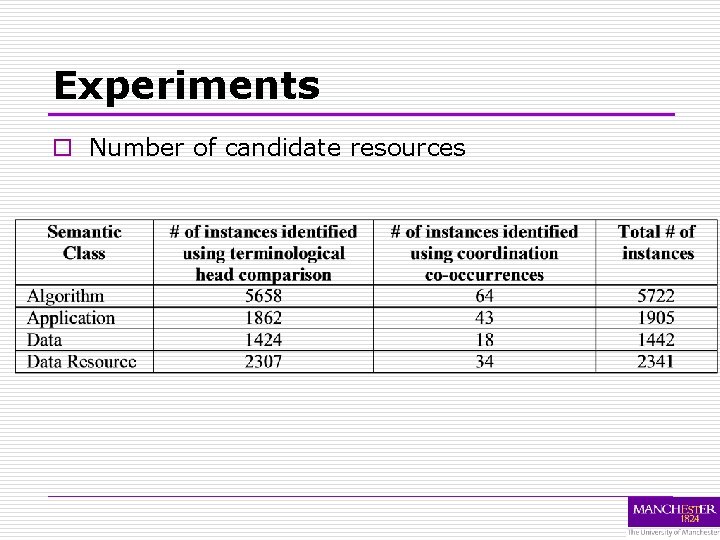
Experiments o Number of candidate resources
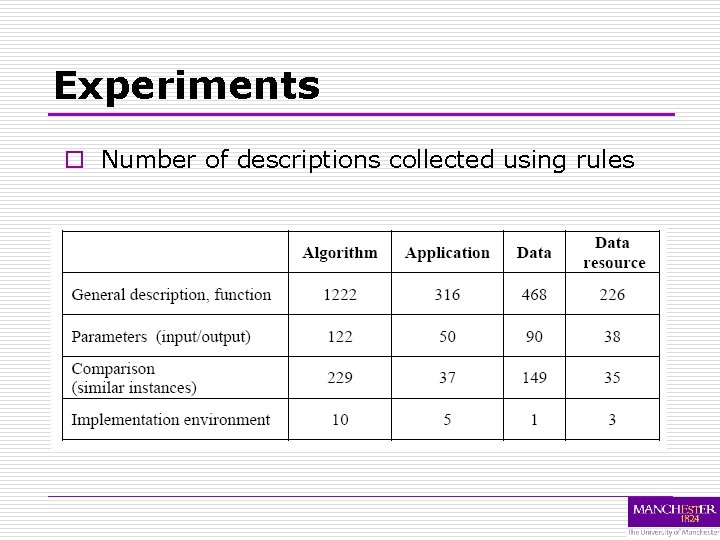
Experiments o Number of descriptions collected using rules
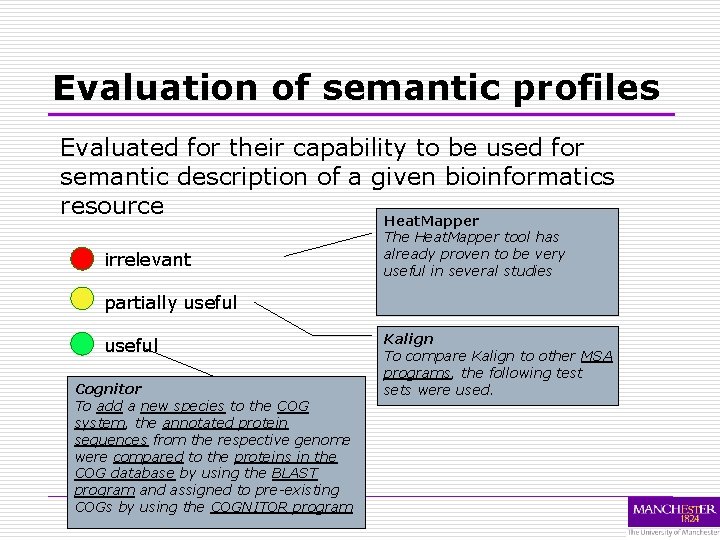
Evaluation of semantic profiles Evaluated for their capability to be used for semantic description of a given bioinformatics resource Heat. Mapper irrelevant The Heat. Mapper tool has already proven to be very useful in several studies partially useful Cognitor To add a new species to the COG system, the annotated protein sequences from the respective genome were compared to the proteins in the COG database by using the BLAST program and assigned to pre-existing COGs by using the COGNITOR program Kalign To compare Kalign to other MSA programs, the following test sets were used.

Evaluation of semantic profiles o Two experiments: o 5 well-known resources with descriptions already available o excellent rating for sentences o average rating for semantic descriptors o predicate functions o 5 new, unknown resources o excellent rating for sentences o average rating for semantic descriptors o predicate functions

What next? o Good recall, poor precision n context needs a better model o Mining parameter values n sub-language of parameters o Candidate service/resource mentions n an entity whose profile looks like a service n comparison of semantic profiles n network of services [ISMB 2009] o Do we have good service ontologies? http: //gnode 1. mib. man. ac. uk/bioinf

Conclusion o Literature mining approach to service description and annotation o Aims n reduce curation efforts n provide semantic synopses of services for the Semantic Web o Potential of text mining n integration with other annotation approaches n extracting the entire service context is still challenging

Acknowledgements o gn. TEAM (text extraction, analitics, mining) H. Yang, I. Spasic, H. Afzal, A. Gledson, J. Eales, M. Greenwood, F. Sarafraz o my. Grid team: Franck Tanoh o BBSRC n “Mining term associations from literature to support knowledge discovery in biology” (2005 -2008) n “pubmed 2 ensembl” (2009 -2010) n “Bio. Catalogue” (2008 -2011)

Announcement o Journal of Bio. Medical Semantics n published by Bio. Med Central n launched at ISMB 2009 o Topics include n Infrastructure for biomedical semantics o semantic resources and repositories o meta-data management and resource description o knowledge representation and semantic frameworks o Biomedical Semantic Web o life-long management of semantic resources n Semantic mining, annotation and analysis
- Slides: 37