Mining Frequent Subgraphs COMP 790 90 Seminar Spring

Mining Frequent Subgraphs COMP 790 -90 Seminar Spring 2007
Overview Introduction Finding recurring subgraphs from graph databases. g. Span FFSM 1 L 06 2 6/18/2021
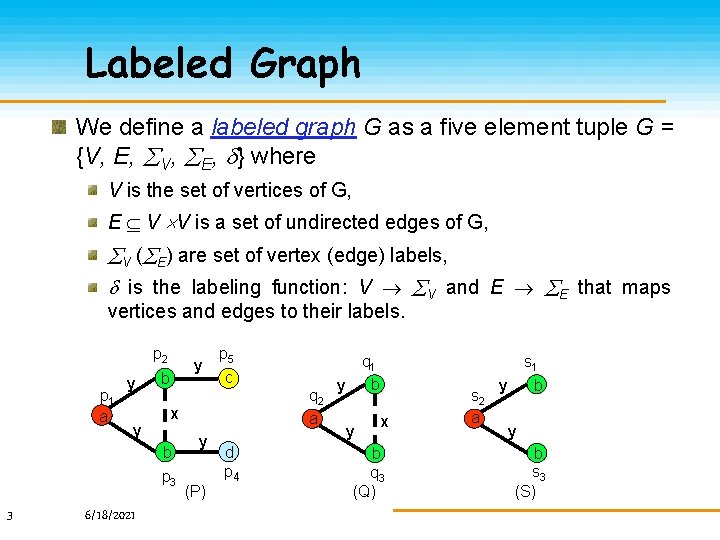
Labeled Graph We define a labeled graph G as a five element tuple G = {V, E, V, E, } where V is the set of vertices of G, E V V is a set of undirected edges of G, V ( E) are set of vertex (edge) labels, is the labeling function: V V and E E that maps vertices and edges to their labels. p 2 p 1 a y y y b 6/18/2021 q 2 a x b p 3 3 p 5 c y (P) d p 4 s 1 q 1 b y y x b q 3 (Q) s 2 a y b s 3 (S)
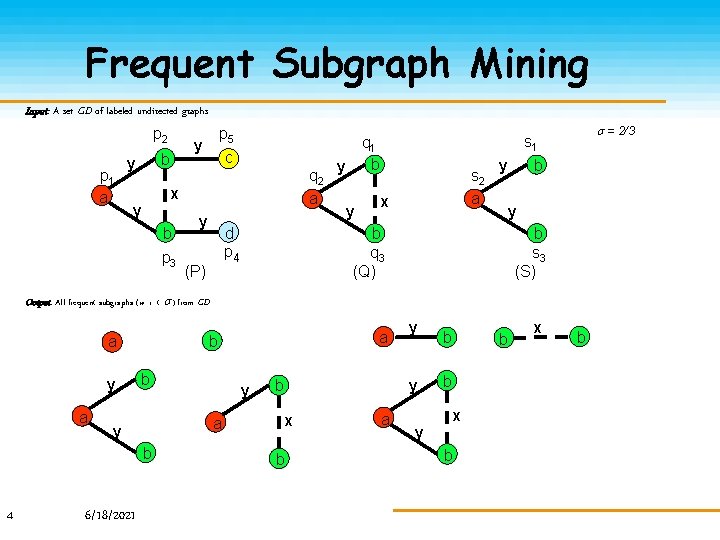
Frequent Subgraph Mining Input: A set GD of labeled undirected graphs p 2 p 1 a y b y p 5 c q 2 a x y b p 3 y d p 4 (P) y y = 2/3 s 1 q 1 b s 2 a x y b s 3 (S) b q 3 (Q) Output: All frequent subgraphs (w. r. t. ) from GD. a y a b y 6/18/2021 y a b 4 a b y y b x b a y b b b x b

Finding Frequent Subgraphs Given a graph database GD = {G 0, G 1, …, Gn}, find all subgraphs appearing in at least graphs. Isomorphic subgraphs are considered the same subgraph. Apriori approaches Generation of subgraph candidates is complicated and expensive. Subgraph isomorphism is an NP-complete problem, so pruning is expensive.

g. Span DFS without candidate generation Relabels graph representation to support DFS. Discovers all frequent subgraphs without candidate generation or pruning. DFS Representation Map each graph to a DFS code (sequence). Lexicographically order the codes. Construct a search tree based on the lexicographic order.
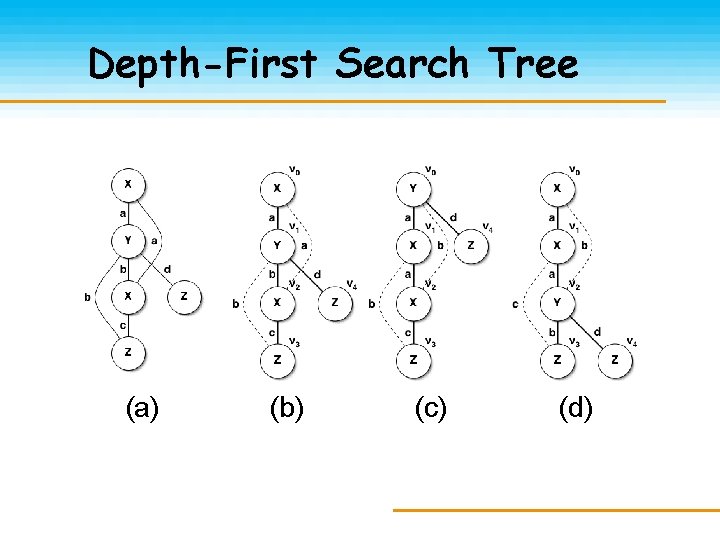
Depth-First Search Tree (a) (b) (c) (d)
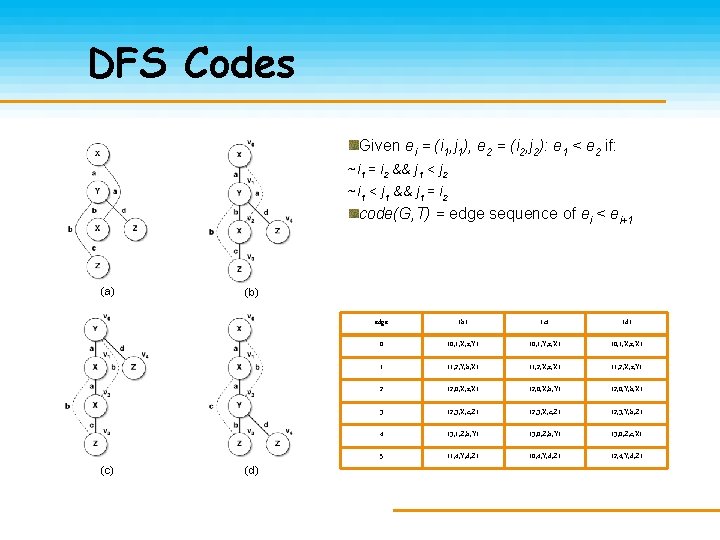
DFS Codes Given ei = (i 1, j 1), e 2 = (i 2, j 2): e 1 < e 2 if: ~i 1 = i 2 && j 1 < j 2 ~i 1 < j 1 && j 1 = i 2 code(G, T) = edge sequence of ei < ei+1 (a) (c) (b) (d) edge (b) (c) (d) 0 (0, 1, X, a, Y) (0, 1, Y, a, X) (0, 1, X, a, X) 1 (1, 2, Y, b, X) (1, 2, X, a, Y) 2 (2, 0, X, a, X) (2, 0, X, b, Y) (2, 0, Y, b, X) 3 (2, 3, X, c, Z) (2, 3, Y, b, Z) 4 (3, 1, Z, b, Y) (3, 0, Z, c, X) 5 (1, 4, Y, d, Z) (0, 4, Y, d, Z) (2, 4, Y, d, Z)

DFS Lexicographic Order ∂ = code(G∂, T∂) = (a 0, a 1, …, am) ß = code(Gß, Tß) = (b 0, b 1, …, bn) ∂ ≤ ß iff (1) or (2): (1) (2) Minimum DFS code The minimum DFS code min(G), in DFS lexicographic order, is the canonical label of graph G. Graphs A and B are isomorphic if min(A) = min(B).

DFS Codes: Parents and Children If ∂ = (a 0, a 1, …, am) and ß = (a 0, a 1, …, am, b): ß is the child of ∂. ∂ is the parent of ß. A valid DFS code requires that b grows from a vertex on the rightmost path.
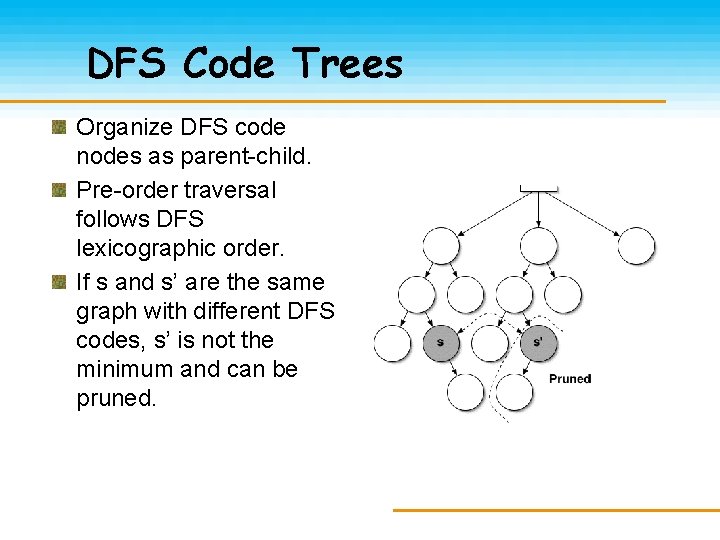
DFS Code Trees Organize DFS code nodes as parent-child. Pre-order traversal follows DFS lexicographic order. If s and s’ are the same graph with different DFS codes, s’ is not the minimum and can be pruned.

g. Span D is the set of all graphs. S is the result set. Algorithm 1: Graph. Set_Projection(D, S) 1: sort labels in D by frequency 2: remove infrequent vertices and edges 3: relabel remaining vertices and edges 4: S’ = all frequent 1 -edge graphs in D 5: sort S’ in DFS lexicographic order 6: S = S’ 7: foreach edge e in S’ do 8: s = graph defined by e 9: s. D = subgraphs in D containing e 10: Subgraph_Mining(D, S, s) 11: D=D-e 12: if |D| < min. Sup 13: break Subprocedure 1: Subgraph_Mining(D, S, s) 1: if s != min(s) 2: return 3: S = S U {s} 4: s’ = +1 -edge children of s in s. D 5: foreach child c of s’ do 6: if support(c) ≥ min. Sup 7: Subgraph_Mining(Ds, S, c)
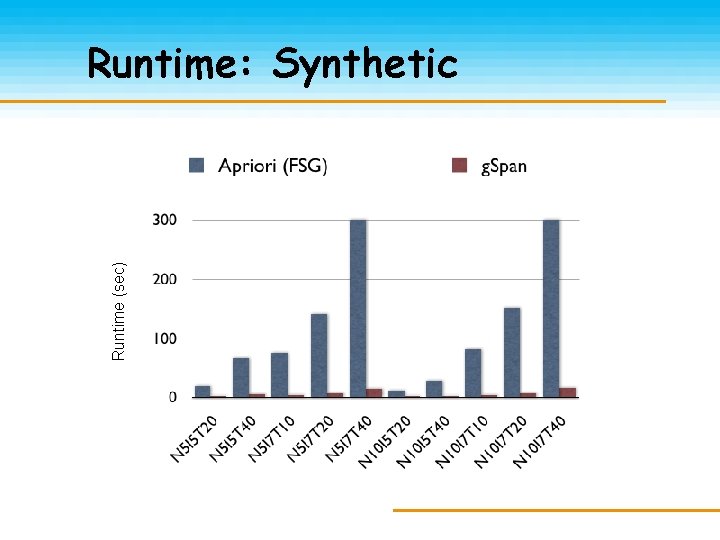
Runtime (sec) Runtime: Synthetic
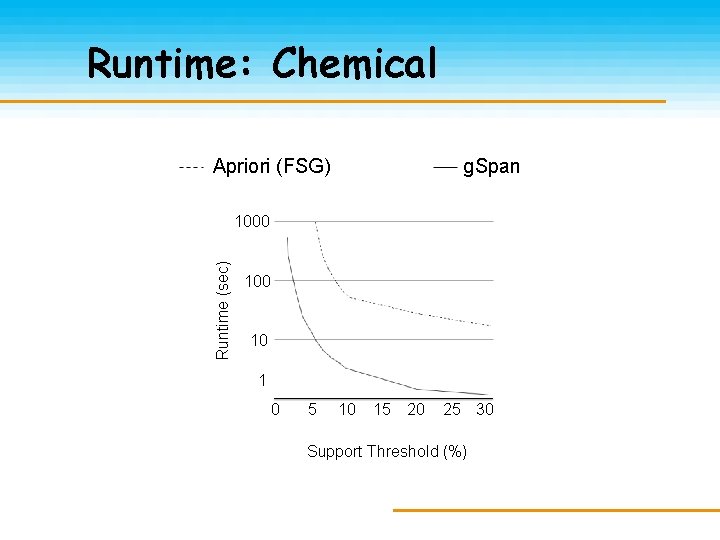
Runtime: Chemical Apriori (FSG) g. Span Runtime (sec) 1000 10 1 0 5 10 15 20 25 30 Support Threshold (%)

g. Span Advantages Lower memory requirements. Faster than naïve FSG by an order of magnitude. No candidate generation. Lexicographic ordering minimizes search tree. False positives pruning. Any disadvantage?

FFSM: Fast Frequent Subgraph Mining -- An Overview: How to solve graph isomorphism problem? A Novel Graph Canonical Form: CAM How to tackle subgraph isomorphism problem (NP-complete)? Incrementally maintained embeddings How to enumerate subgraphs: An Efficient Data Structure: CAM Tree Two Operations: CAM-join, CAM-extension. 16 6/18/2021
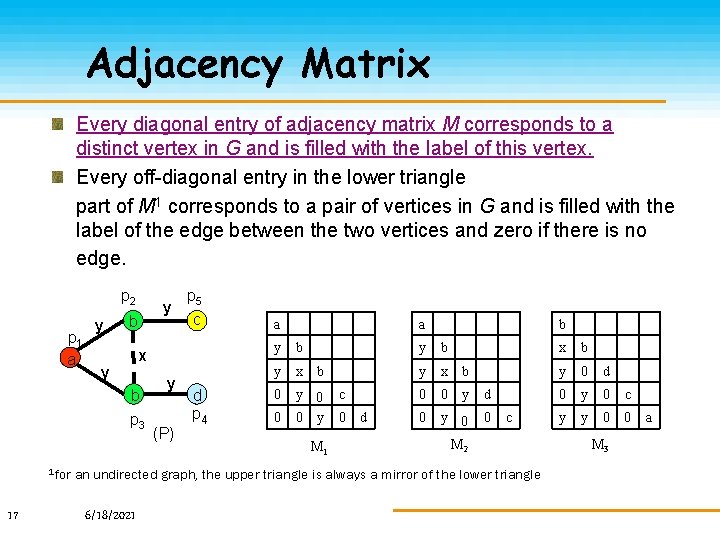
Adjacency Matrix Every diagonal entry of adjacency matrix M corresponds to a distinct vertex in G and is filled with the label of this vertex. Every off-diagonal entry in the lower triangle part of M 1 corresponds to a pair of vertices in G and is filled with the label of the edge between the two vertices and zero if there is no edge. p 2 p 1 a y y b x y b p 3 1 for 17 p 5 c y (P) d p 4 a y b y x 0 y 0 b a b M 1 c 0 d y b y x b 0 0 y d 0 y 0 0 c M 2 an undirected graph, the upper triangle is always a mirror of the lower triangle 6/18/2021 x b y 0 d 0 y 0 c y y 0 0 M 3 a

Code A Code of n n adjacency matrix M is defined as sequence of lower triangular entries (including the diagonal entries) in the order: M 1, 1 M 2, 2 … Mn, 1 Mn, 2 …Mn, n-1 Mn, n a y b y x 0 y 0 b M 1 c 0 d Code(M 1): aybyxb 0 y 0 c 00 y 0 d > Code(M 2): aybyxb 00 yd 0 y 00 c > Code(M 3): bxby 0 d 0 y 0 cyy 00 a b a y b y x b 0 0 y 0 0 M 2 d c x b y 0 d 0 y 0 c y y 0 0 a M 3 The Canonical Adjacency Matrix is the one produces the maximal code, using lexicographic order. 18 6/18/2021

MP Submatrix For an m m matrix A, an n n matrix B is A’s maximal proper submatrix (MP Submatrix), iff N is obtained by removing the last none-zero entry from M. a a a M 1 y b M 2 a y b y 0 M 3 b y x M 4 b y x 0 y 0 b M 5 We define a CAM is connected iff the corresponding graph is connected. Theorem I: A CAM’s MP submatrix is CAM Theorem II: A connected CAM’s MP submatrix is connected 19 6/18/2021 c y b y x 0 y 0 b M 6 c 0 d
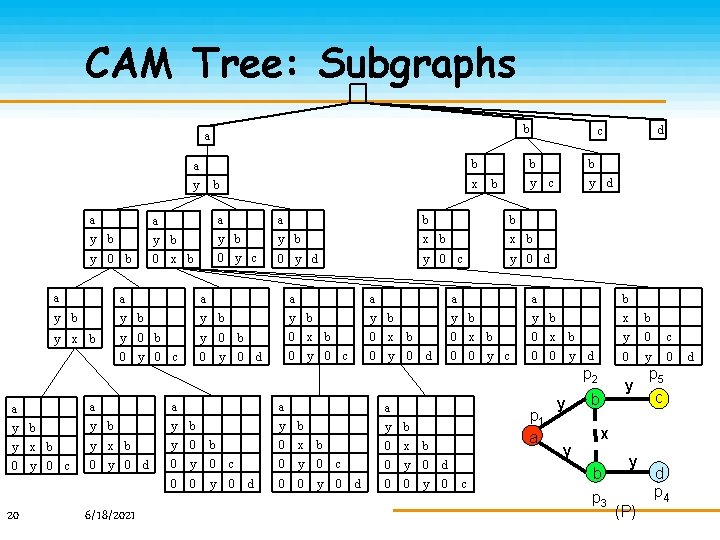
CAM Tree: Subgraphs b a y x b y b b c a a b b y b y b x b 0 y c 0 y d y 0 c y 0 d y 0 b a a a y b y b y 0 b 0 y x b b a b 0 y 0 c a a y b 0 x b 0 y 0 d c d y a a y b 0 x b 0 0 y c d b d p 2 a a a y b y b y x b y 0 b 0 y 0 d 0 y 0 c 0 0 y 0 20 c 6/18/2021 a d a y b 0 x b 0 y 0 c 0 0 y 0 d p 1 a y b 0 x b 0 y 0 d 0 0 y 0 c y y x b y 0 c 0 y b p 5 c x b p 3 y (P) d p 4 d
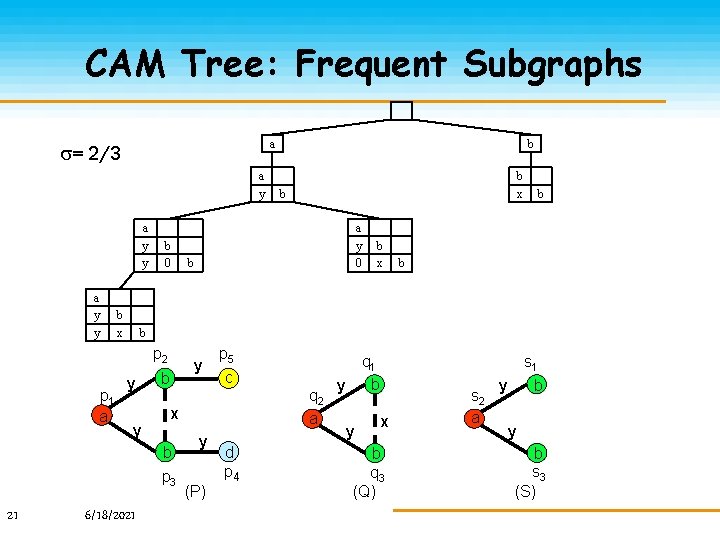
CAM Tree: Frequent Subgraphs a = 2/3 a y y b x b 0 b a y 0 b x b b b y y y b 6/18/2021 p 5 c q 2 a x b p 3 21 b x b p 2 p 1 a b y (P) d p 4 s 1 q 1 b y y x b q 3 (Q) s 2 a y b s 3 (S)

How to Enumerate Nodes in a CAM Tree? Two operations to explore CAM tree: CAM-Join CAM-Extension Augmenting CAM tree with Suboptimal CAMs Objectives: none false dismissal no redundancy Plus: We want to this efficiently! 22 6/18/2021
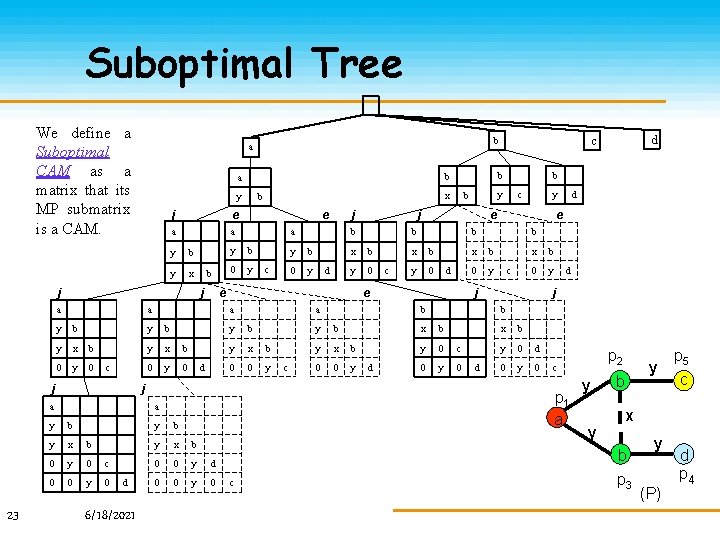
Suboptimal Tree We define a Suboptimal CAM as a matrix that its MP submatrix is a CAM. y j e a a y b y x j j a y b y x b 0 y 0 c 0 0 y 0 d 6/18/2021 c b y x b 0 y 0 d j b y b 0 y d b y 0 c a y b y x b 0 0 y c b y x b 0 0 y d y x b y 0 d d e b b x b 0 y c x b 0 y j b y c e b x b y b e a y j a b d j b x b y 0 c 0 y 0 d x b y 0 d 0 y b y x b 0 0 y d 0 0 y 0 c p 2 c p 1 a a y e y j a x b e a j 23 b b b a d c b a y y y b p 5 c x b p 3 y (P) d p 4

Summary Theorem: For a graph G, let CK-1 (Ck) be set of the suboptimal CAMs of all the size (K-1) (K) subgraphs of G (K ≥ 2). Every member of set CK can be enumerated unambiguously either by joining two members of set CK-1 or by extending a member in CK-1. 24 6/18/2021

Experimental Study Predictive Toxicology Evaluation Competition (PTE) Contains: 337 compounds Each graph contains 27 nodes and 27 edges on average NIH DTP Anti-Viral Screen Test (DTP CA/CM) Chemicals are classified to be Confirmed Active (CA), Confirmed Moderate Active (CM) and Confirmed Inactive (CI). We formed a dataset contains CA (423) and CM (1083). Each graph contains 25 nodes and 27 edges on average 25 6/18/2021
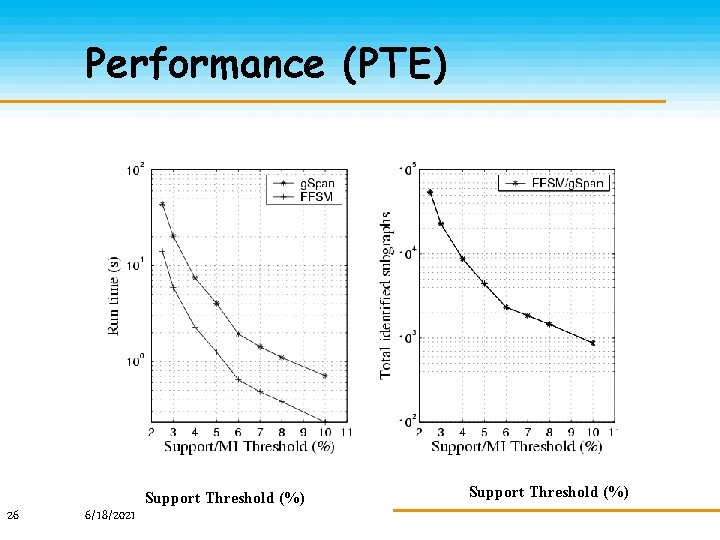
Performance (PTE) Support Threshold (%) 26 6/18/2021 Support Threshold (%)
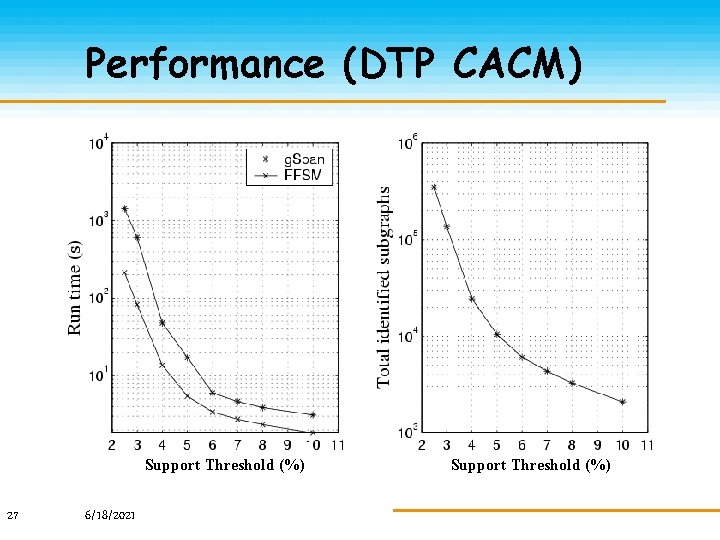
Performance (DTP CACM) Support Threshold (%) 27 6/18/2021 Support Threshold (%)
- Slides: 27