Minimum PCR Primer Set Selection with Amplification Length



![Motivation for Primer Set Selection (1) • Spotted microarray synthesis [Fernandes and Skiena’ 02] Motivation for Primer Set Selection (1) • Spotted microarray synthesis [Fernandes and Skiena’ 02]](https://slidetodoc.com/presentation_image_h2/9e8b866dc769fb3a538c77a4e3badd92/image-6.jpg)


![Previous Work (1) • [Pearson et al. 96][Linhart&Shamir’ 02][Souvenir et al. ’ 03] - Previous Work (1) • [Pearson et al. 96][Linhart&Shamir’ 02][Souvenir et al. ’ 03] -](https://slidetodoc.com/presentation_image_h2/9e8b866dc769fb3a538c77a4e3badd92/image-9.jpg)
![Previous Work (2) • [Fernandes&Skiena’ 02] model primer selection as a minimum multicolored subgraph Previous Work (2) • [Fernandes&Skiena’ 02] model primer selection as a minimum multicolored subgraph](https://slidetodoc.com/presentation_image_h2/9e8b866dc769fb3a538c77a4e3badd92/image-10.jpg)









- Slides: 24

Minimum PCR Primer Set Selection with Amplification Length and Uniqueness Constraints Ion Mandoiu University of Connecticut CS&E Department May 25, 2004 GSU Biotech Symposium 1

Combinatorial Optimization Applications in Bioinformatics • Fast growing number of applications – – Dynamic Programming & Integer Programming in sequence alignment TSP and Euler paths in DNA sequencing Integer Programming in Haplotype inference Integer Programming & approximation algorithms for efficient pathogen identification (string barcoding) – … May 25, 2004 GSU Biotech Symposium 2

High-Thrughput Assay Design • New source of combinatorial problems – – – – Microarray probe selection Mask design for Affy arrays Universal tag arrays Self-assembling microarrays Quality control … This talk: Multiplex PCR primer set selection • Optimization goals – Improved speed – High reliability – Reduced COST May 25, 2004 GSU Biotech Symposium 3

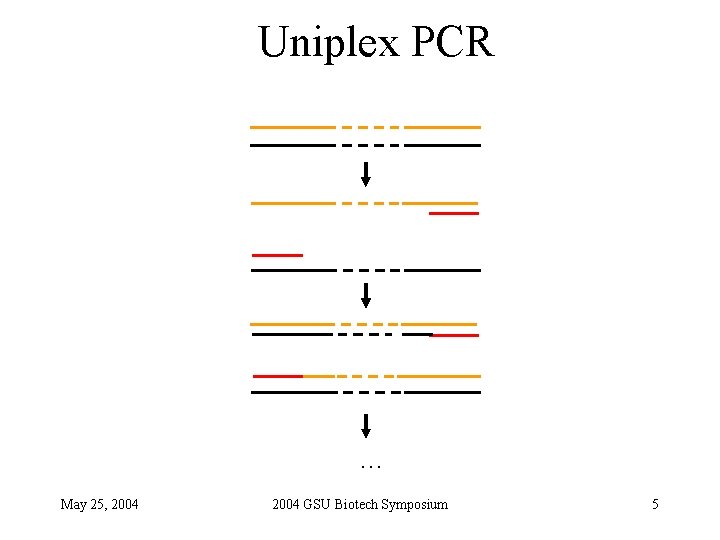
Uniplex PCR … May 25, 2004 GSU Biotech Symposium 5

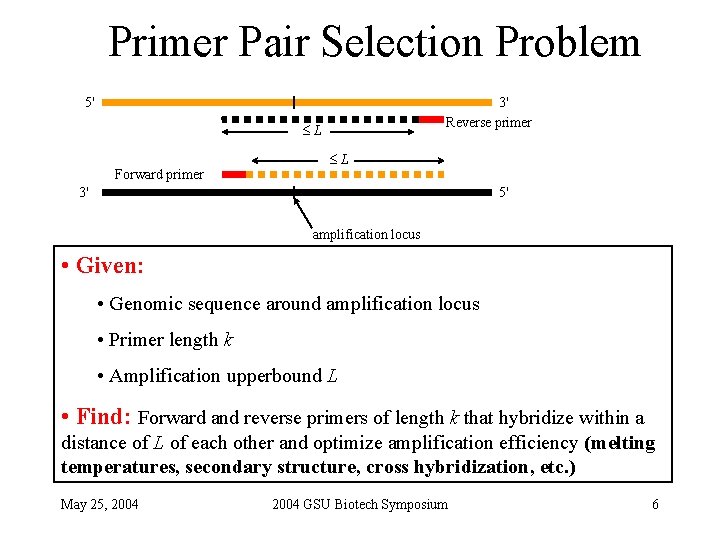
Primer Pair Selection Problem 5' 3' Reverse primer L Forward primer L 3' 5' amplification locus • Given: • Genomic sequence around amplification locus • Primer length k • Amplification upperbound L • Find: Forward and reverse primers of length k that hybridize within a distance of L of each other and optimize amplification efficiency (melting temperatures, secondary structure, cross hybridization, etc. ) May 25, 2004 GSU Biotech Symposium 6
![Motivation for Primer Set Selection 1 Spotted microarray synthesis Fernandes and Skiena 02 Motivation for Primer Set Selection (1) • Spotted microarray synthesis [Fernandes and Skiena’ 02]](https://slidetodoc.com/presentation_image_h2/9e8b866dc769fb3a538c77a4e3badd92/image-6.jpg)
Motivation for Primer Set Selection (1) • Spotted microarray synthesis [Fernandes and Skiena’ 02] – Need unique pair for each amplification product, but primers can be reused to minimize cost – Potential to reduce #primers from O(n) to O(n 1/2) for n products May 25, 2004 GSU Biotech Symposium 7

Motivation for Primer Set Selection (2) • SNP Genotyping – Thousands of SNPs that must genotyped using hybridization based methods (e. g. , SBE) – Selective PCR amplification needed to improve accuracy of detection steps (whole-genome amplification not appropriate) – No need for unique amplification! – Primer minimization is critical • Fewer primers to buy • Fewer multiplex PCR reactions May 25, 2004 GSU Biotech Symposium 8

Primer Set Selection Problem • Given: • Genomic sequences around each amplification locus • Primer length k • Amplification upperbound L • Find: • Minimum size set of primers S of length k such that, for each amplification locus, there are two primers in S hybridizing to the forward and reverse sequences within a distance of L of each other • For some applications: S should contain a unique pair of primers amplifying each locus May 25, 2004 GSU Biotech Symposium 9
![Previous Work 1 Pearson et al 96LinhartShamir 02Souvenir et al 03 Previous Work (1) • [Pearson et al. 96][Linhart&Shamir’ 02][Souvenir et al. ’ 03] -](https://slidetodoc.com/presentation_image_h2/9e8b866dc769fb3a538c77a4e3badd92/image-9.jpg)
Previous Work (1) • [Pearson et al. 96][Linhart&Shamir’ 02][Souvenir et al. ’ 03] - Separately select forward and reverse primers - To enforce bound of L on amplification length, select only primers that are within a distance of L/2 of the target SNP • Ignores half of the feasible primer pairs • Solution can increase by a factor of O(n) by ignoring them! • Greedy set cover algorithm gives O(ln n) approximation factor for this formulation • Cannot approximate better unless P=NP May 25, 2004 GSU Biotech Symposium 10
![Previous Work 2 FernandesSkiena 02 model primer selection as a minimum multicolored subgraph Previous Work (2) • [Fernandes&Skiena’ 02] model primer selection as a minimum multicolored subgraph](https://slidetodoc.com/presentation_image_h2/9e8b866dc769fb3a538c77a4e3badd92/image-10.jpg)
Previous Work (2) • [Fernandes&Skiena’ 02] model primer selection as a minimum multicolored subgraph problem: • Vertices of the graph correspond to candidate primers • There is an edge colored by color i between primers u and v if they hybridize to i-th forward and reverse sequences within a distance of L • Goal is to find minimum size set of vertices inducing edges of all colors • No non-trivial approximation factor known previously May 25, 2004 GSU Biotech Symposium 11

Selection w/o Uniqueness Constraints • Can be seen as a “simultaneous set covering” problem: - The ground set is partitioned into n disjoint sets, each with 2 L elements - The goal is to select a minimum number of sets (== primers) that cover at least half of the elements in each partition • Naïve modifications of the greedy set cover algorithm do not work • Key idea: use potential function for a partial solution P = minium number of elements that are not yet covered as measure of infeasibility • Initially, = n. L • For feasible solutions, = 0 May 25, 2004 GSU Biotech Symposium 12

Potential-Function Driven Greedy 1. Select a primer that decreases the potential function by the largest amount (breaking ties arbitrarily) 2. Repeat until feasibility is achived • Lemma: Each greedy selection reduces by a factor of at least (1 -1/OPT) • Theorem: The number of primers selected by the greedy algorithm is at most ln(n. L) larger than the optimum May 25, 2004 GSU Biotech Symposium 13

Selection w/ Uniqueness Constraints • Can be modeled as minimum multicolored subgraph problem: add edge colored by color i between two primers if they amplify ith SNP and do not amplify any other SNP • Trivial approximation algorithm: select 2 primers for each SNP • O(n 1/2) approximation since at least n 1/2 primers required by every solution • Non-trivial approximation? May 25, 2004 GSU Biotech Symposium 14

Integer Program Formulation • Variable xu for every vertex (candidate primer) u - xu set to 1 if u is selected, and to 0 otherwise • Variable ye for every edge e - ye set to 1 if corresponding primer pair selected to amplify one of the SNPs • Objective: minimize sum of xu’s • Constraints: - for each i, sum of {ye : e amplifying SNP i} 1 - ye xu for every e incident to u May 25, 2004 GSU Biotech Symposium 15

LP-Rounding Algorithm 1. Solve linear programming relaxation 2. Select node u with probability xu • Theorem: With probability of at least 1/3, the number of selected nodes is within a factor of O(m 1/2 lnn) of the optimum, where m is the maximum number of edges sharing the same color. • For primer selection, m L 2 approximation factor is O(Lln n) May 25, 2004 GSU Biotech Symposium 16

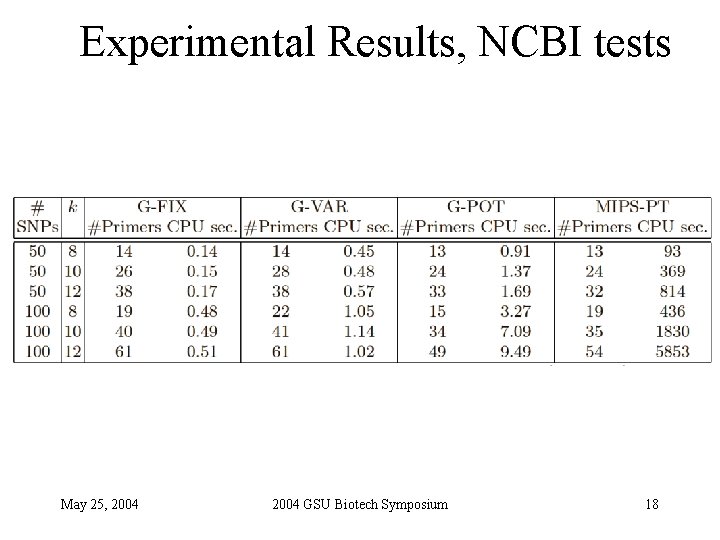
Experimental Setting • SNP sets extracted from NCBI databases + randomly generated • C/C++ code run on a 2. 8 GHz Dell Power. Edge running Linux • Compared algorithms • G-FIX: greedy primer cover algorithm of Pearson et al. - Primers restricted to be within L/2 of amplified SNPs • G-VAR: naïve modification of G-FIX - For each SNP, first selected primer can be L bases away from SNP - If first selected primer is L 1 bases away from the SNP, opposite sequence is truncated to a length of L- L 1 • G-POT: potential function driven greedy algorithm • MIPS-PT: iterative beam-search heuristic of Souvenir et al (WABI’ 03) May 25, 2004 GSU Biotech Symposium 17

Experimental Results, NCBI tests May 25, 2004 GSU Biotech Symposium 18

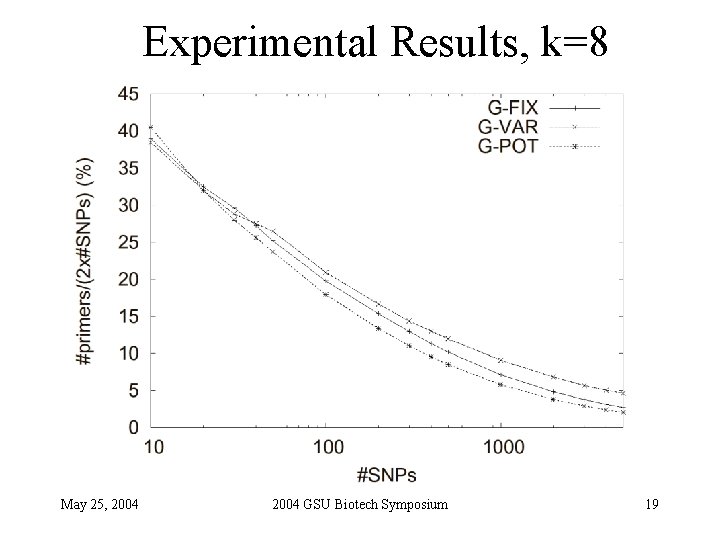
Experimental Results, k=8 May 25, 2004 GSU Biotech Symposium 19

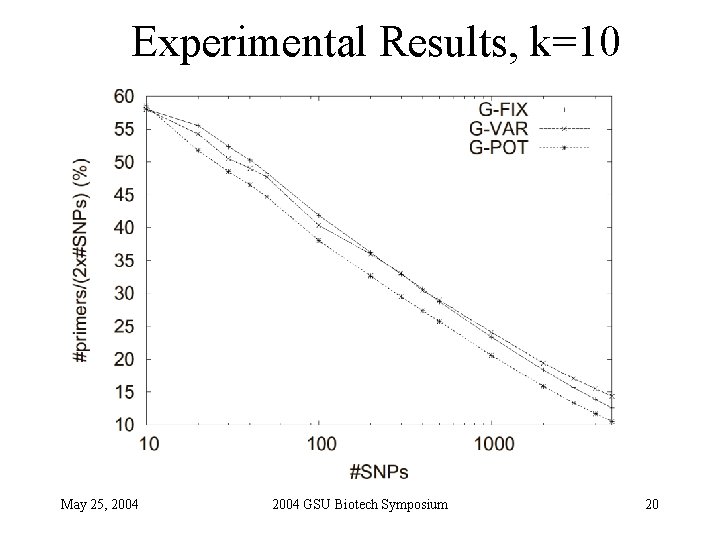
Experimental Results, k=10 May 25, 2004 GSU Biotech Symposium 20

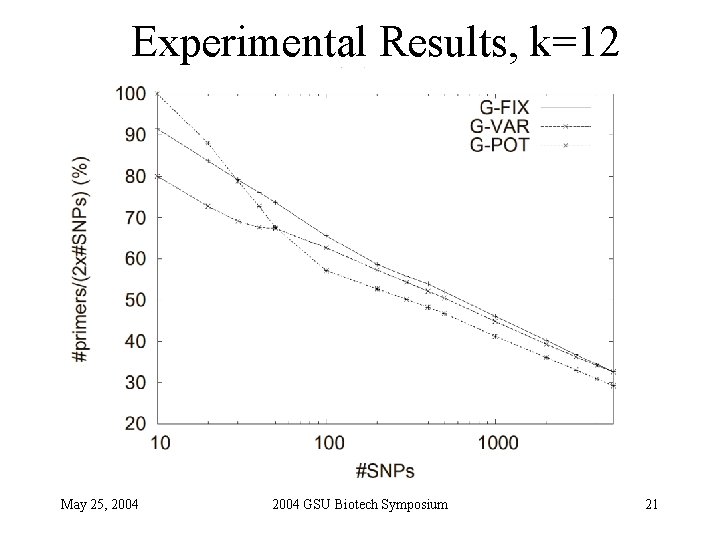
Experimental Results, k=12 May 25, 2004 GSU Biotech Symposium 21

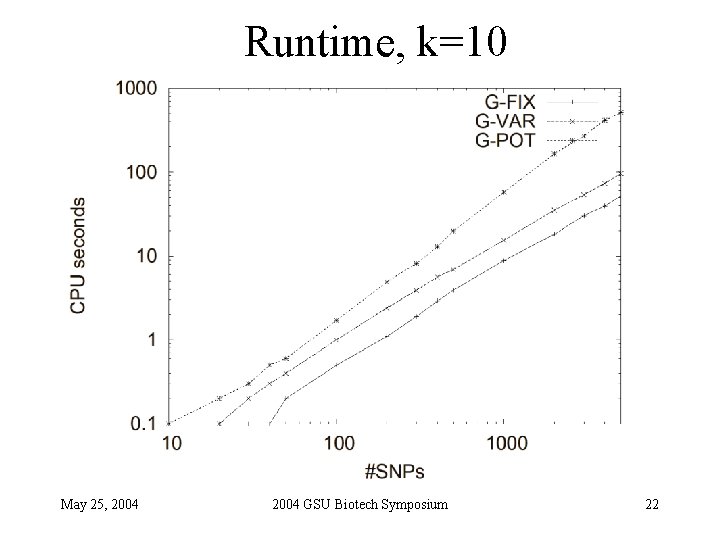
Runtime, k=10 May 25, 2004 GSU Biotech Symposium 22

Conclusions • New combinatorial optimization problems arising in the area of high-throughput assay design • Theoretical insights (such as approximation results) give algorithms with significant practical improvements • Choosing the proper problem model is critical to solution efficiency May 25, 2004 GSU Biotech Symposium 23

Ongoing Work & Open Problems • Allow degenerate primers • Incorporate more biochemical constraints into the model (melting temperature, secondary structure, cross hybridization, etc. ) • Close gap between O(lnn) inapproximability bound and O(L lnn) approximation factor for minimum multi-colored subgraph problem • Approximation algorithms for partition into multiplexed PCR reactions (Aumann et al. WABI’ 03) May 25, 2004 GSU Biotech Symposium 24

Acknowledgments • • Kishori Konwar Alex Russell Alex Shvartsman Financial support from UCONN Research Foundation May 25, 2004 GSU Biotech Symposium 25