Minimal Models for Quantum Decoherence in Coupled Biomolecules

Minimal Models for Quantum Decoherence in Coupled Biomolecules Joel Gilmore Ross H. Mc. Kenzie University of Queensland, Brisbane, Australia Gilmore and Mc. Kenzie, J. Phys. : Cond. Matt. 17, 1735 (2005) and quant-ph/0412170, to appear in Chem. Phys. Lett

Why should physicists biologists be be interestedininquantum? biology? Quantum They’re allmechanics highly efficient, plays ahighly critical refined, role selfinassembling much of biology! quantum nanoscale devices. • Retinal, responsible for vision – Ultrafast vision receptor • Light harvesting complexes in photosynthesis – Ultraefficient collection & conversion of light • Green Fluorescent Protein – Highly efficient marker
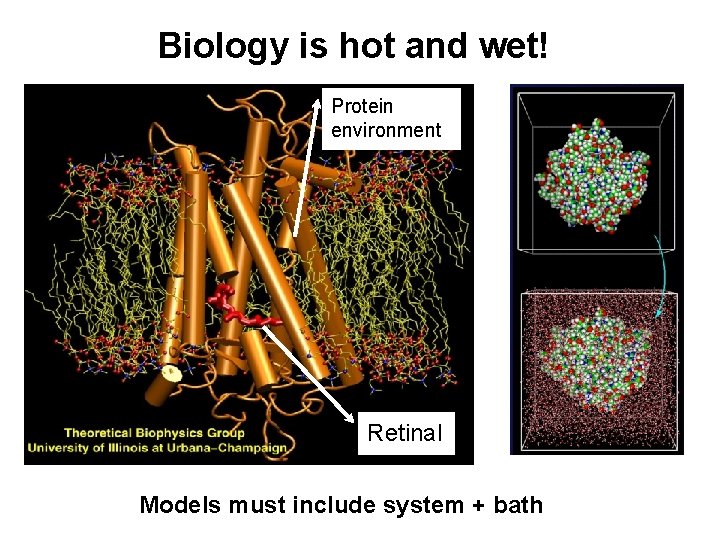
Biology is hot and wet! Protein environment Retinal Models must include system + bath

The spin-boson model • Popular model for describing decoherence – Extensively studies by Leggett, Weiss, Saleur, Costi, et al. – Applications to SQUIDS, decoherence of qubits • Describes the coupling of a two level system to a bath of harmonic oscillators – Works for many, very different, environments • All coupling to enviornment is in the spectral density: We can apply this to systems of coupled biomolecules!

Experimental realisation of spin-boson model What is the two level system? • Two molecules • Each with two energy levels If only one excitation is available, effectively a two level system
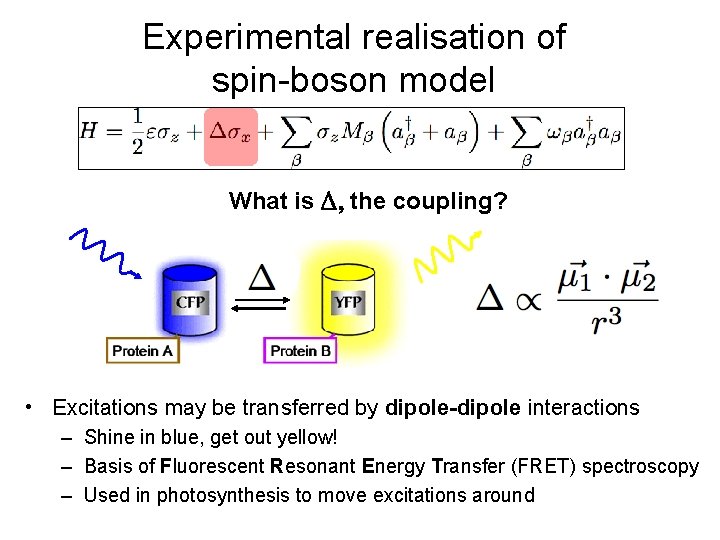
Experimental realisation of spin-boson model What is the coupling? • Excitations may be transferred by dipole-dipole interactions – Shine in blue, get out yellow! – Basis of Fluorescent Resonant Energy Transfer (FRET) spectroscopy – Used in photosynthesis to move excitations around
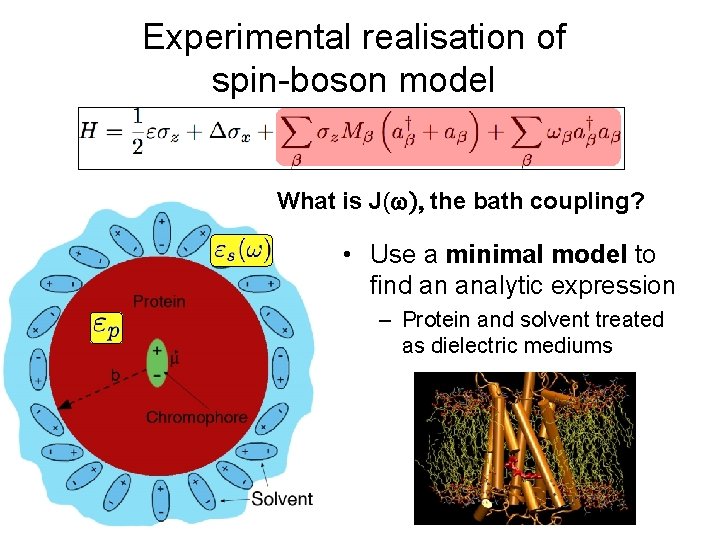
Experimental realisation of spin-boson model What is J( the bath coupling? • Use a minimal model to find an analytic expression – Protein and solvent treated as dielectric mediums
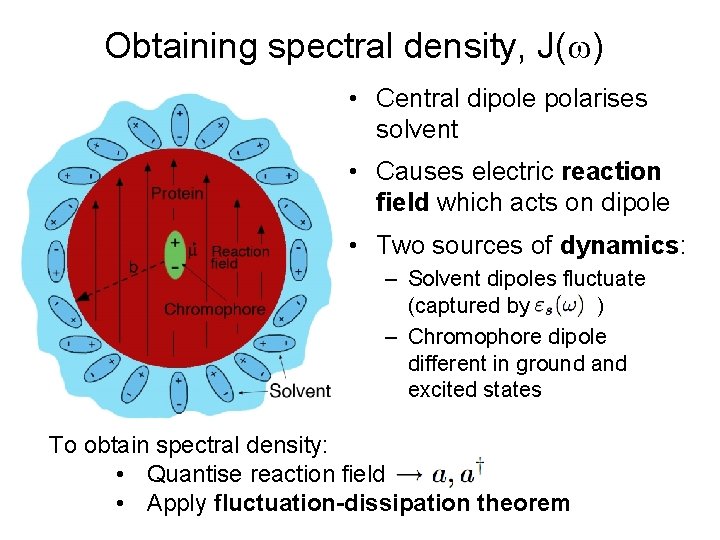
Obtaining spectral density, J( ) • Central dipole polarises solvent • Causes electric reaction field which acts on dipole • Two sources of dynamics: – Solvent dipoles fluctuate (captured by ) – Chromophore dipole different in ground and excited states To obtain spectral density: • Quantise reaction field • Apply fluctuation-dissipation theorem

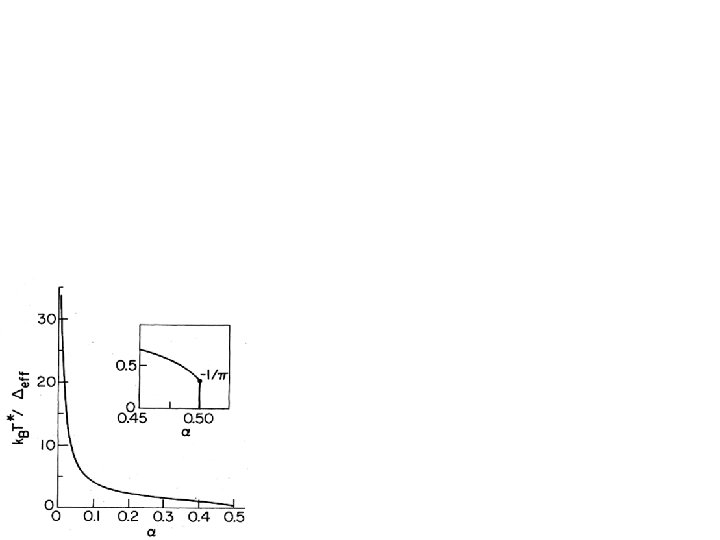
Spectral density for the minimal model = chromophore dipole diff. b = protein radius s( ) = solvent dielectric constant p = protein dielectric constant • Ohmic spectral density - Cut-off determined by solvent dielectric relaxation time, 8 ps • Microscopic derivation of spin-boson model and spectral density • Slope is critical parameter - For chromophore in water, - Protein can shield chromophore, so - c. f. , for Joesephson Junction qubits - Strong decoherence - quantum consciousness unlikely!
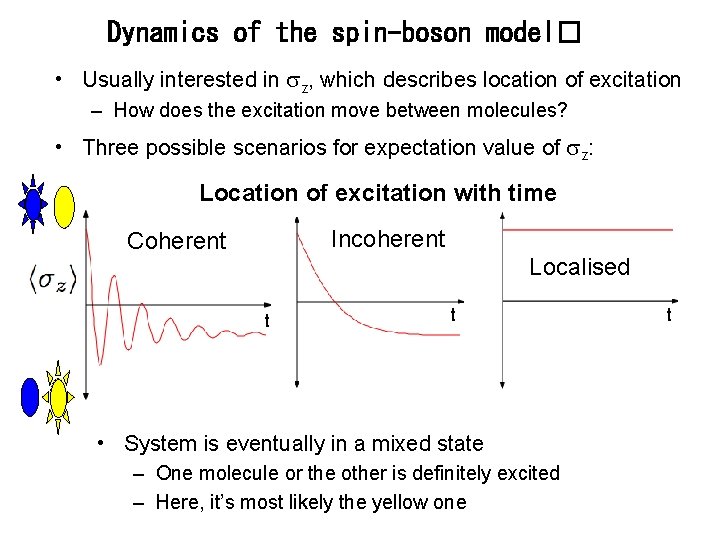
Dynamics of the spin-boson model� • Usually interested in z, which describes location of excitation – How does the excitation move between molecules? • Three possible scenarios for expectation value of z: Location of excitation with time Incoherent Coherent Localised t t • System is eventually in a mixed state – One molecule or the other is definitely excited – Here, it’s most likely the yellow one t

Dynamics of the spin-boson model • Behaviour depends on and relative size of parameters: - c - - k. BT c All known in terms of experimental parameters • Rich, non-trivial dynamics • Cross-over from coherent-incoherent in many ways For identical ( ) molecules and c For c, coherent oscillations remain even for high T, Bias can help or hinder coherent oscillations
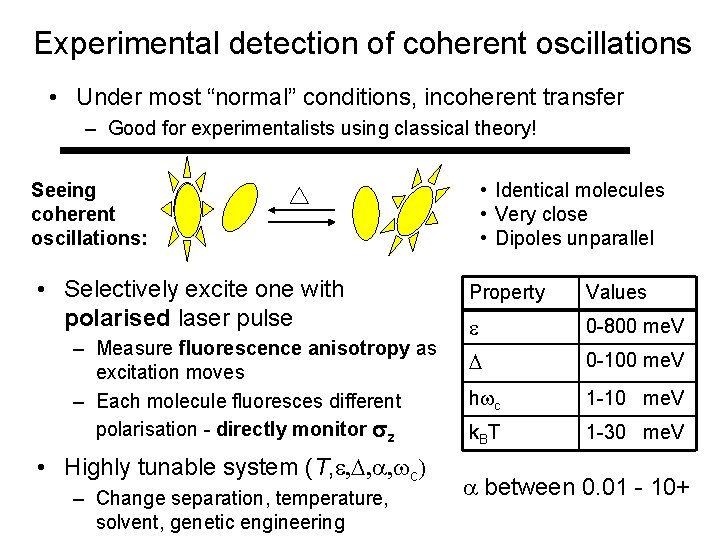
Experimental detection of coherent oscillations • Under most “normal” conditions, incoherent transfer – Good for experimentalists using classical theory! • Identical molecules • Very close • Dipoles unparallel Seeing coherent oscillations: • Selectively excite one with polarised laser pulse – Measure fluorescence anisotropy as excitation moves – Each molecule fluoresces different polarisation - directly monitor z • Highly tunable system (T, c – Change separation, temperature, solvent, genetic engineering Property Values 0 -800 me. V 0 -100 me. V h c 1 -10 me. V k. BT 1 -30 me. V between 0. 01 - 10+

Key Results & Conclusions • Demonstrated an experimental realisation of the spin-boson model in terms of coupled biomolecules • Microscopic derivation of the spectral density through minimal models of the surrounding protein and solvent • Dynamics can be observed directly through experiment • Model applicable to other scenarios – Retinal in vision – Photosynthesis – More complex protein models • Molecular biophysics may be a useful testing ground for models of quantum decoherence – Complex but tuneable systems - self assembling too! – It doesn’t always have to be physics helping advance biology! Sometimes, biology can help physics too!

Acknowledgements • Ross Mc. Kenzie (UQ) • Paul Meredith (UQ) • Ben Powell (UQ) • Andrew Briggs & all at QIPIRC (Oxford) Gilmore and Mc. Kenzie, J. Phys. : Cond. Matt. 17, 1735 (2005) and quant-ph/0412170, to appear in Chem. Phys. Lett

Quantum mechanics in biology Classical biology! Quantum biology! • Ball and stick models • Highly efficient photosynthesis • DNA • Ultrafast vision receptors • No quantum courses for biologists… • Tunneling in enzymes • Quantum consciousness? ! (Okay, probably not) Quantum or classical What decides?
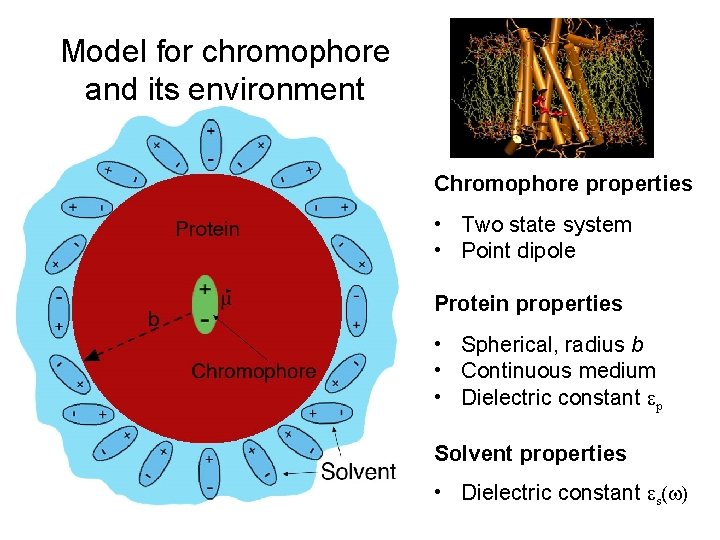
Model for chromophore and its environment Chromophore properties • Two state system • Point dipole Protein properties • Spherical, radius b • Continuous medium • Dielectric constant p Solvent properties • Dielectric constant s( )
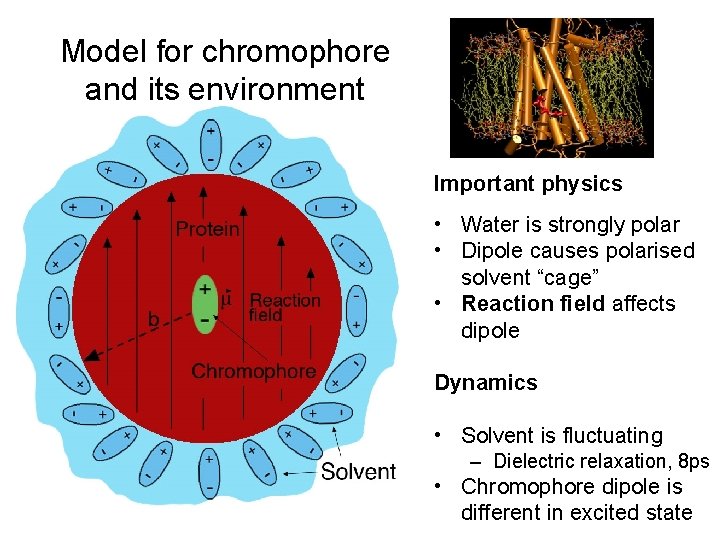
Model for chromophore and its environment Important physics • Water is strongly polar • Dipole causes polarised solvent “cage” • Reaction field affects dipole Dynamics • Solvent is fluctuating – Dielectric relaxation, 8 ps • Chromophore dipole is different in excited state
- Slides: 18