mini PCR Antibiotic Resistance Lab Monitoring resistant organisms

mini. PCR™ Antibiotic Resistance Lab Monitoring resistant organisms in the environment

Instructor note—This case study represents a fictional outbreak based on the synthesis of data representing different infections: 1) an outbreak in Denmark of Cipro-resistant Salmonella traced to consumption of pork that originated from one farm 2) the first patient isolate ever reported containing the NDM-1 gene 3) subsequent sampling of water samples from around New Delhi for presence of the NDM-1 gene 4) a multi-state outbreak of E. coli traced to lettuce from the Yuma, Arizona region and, ultimately, to water from an irrigation canal. 5) identification of carbapenem resistant bacteria in agricultural settings
The vast majority of our food is safe, but sometimes…

When this happens • Public health officials try to find and contain the source of the infection. • Doctors treat patients as quickly as possible using antibiotics. – But more and more often, the bacteria people are infected with are resistant to antibiotic treatments, meaning it is harder and harder ensure that sick people will get well again.

Where does antibiotic resistance come from? • Antibiotics will kill bacteria • But if any bacteria has resistance to an antibiotic, it is more likely to survive and reproduce. Antibiotics

Where does antibiotic resistance come from? • Antibiotics will kill bacteria • But if any bacteria has resistance to an antibiotic, it is more likely to survive and reproduce. • Over time, natural selection leads to more and more antibiotic resistant bacteria.
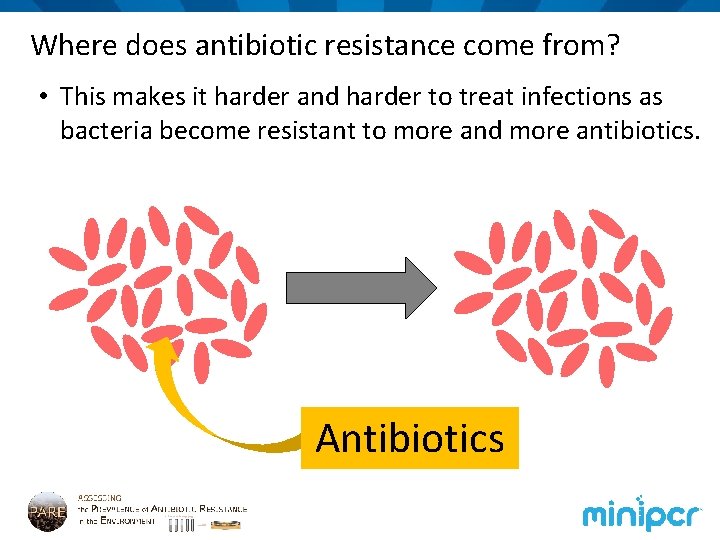
Where does antibiotic resistance come from? • This makes it harder and harder to treat infections as bacteria become resistant to more and more antibiotics. Antibiotics
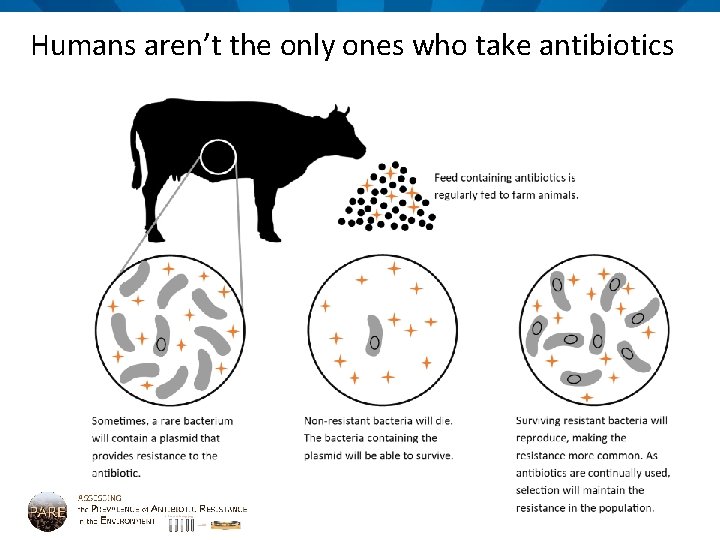
Humans aren’t the only ones who take antibiotics
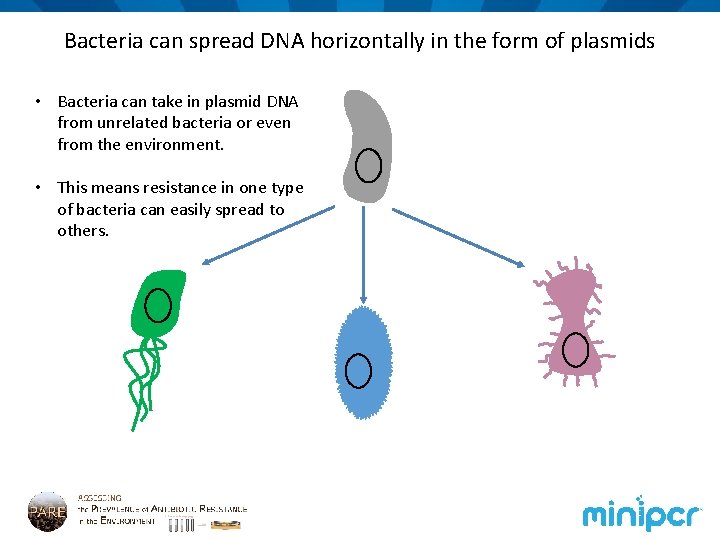
Bacteria can spread DNA horizontally in the form of plasmids • Bacteria can take in plasmid DNA from unrelated bacteria or even from the environment. • This means resistance in one type of bacteria can easily spread to others.

Today’s Lab: • There has been a recent multi-state outbreak of E. coli food poisoning. • Patients hospitalized, all treated with a carbapenem type antibiotic which usually works. • Treatment was ineffective—patients were not improving. • Ultimately a different antibiotic worked and most patients recovered… • Unfortunately, some died.

Today’s Lab: • Public health officials identified the source of the infection as tainted pork originating from a single farm. • Neighboring farms are worried. They want to know if the antibiotic resistant bacteria is a risk to them. • Your Job: Establish whether antibiotic resistant bacteria is spreading in the environment.
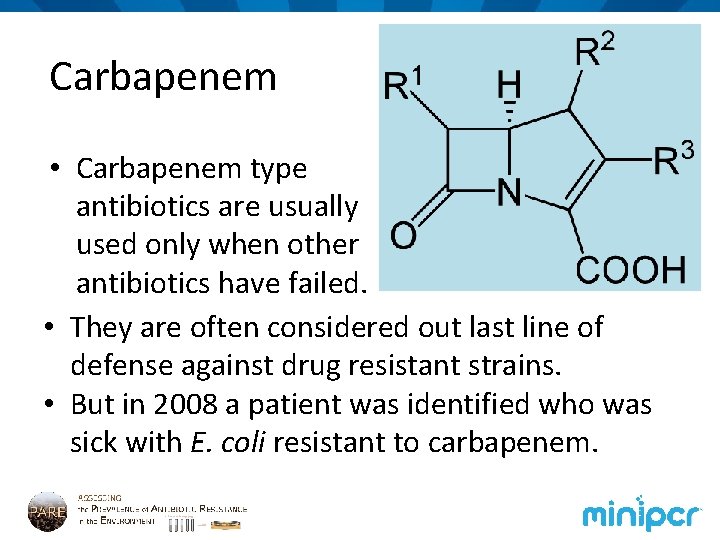
Carbapenem • Carbapenem type antibiotics are usually used only when other antibiotics have failed. • They are often considered out last line of defense against drug resistant strains. • But in 2008 a patient was identified who was sick with E. coli resistant to carbapenem.

Carbapenem resistance Bacteria resistant to carbapenem contain the enzyme NDM-1. • NDM-1 (New Delhi metallo-beta-lactamase 1) is an enzyme that hydrolyzes carbapenems. • First described in 2008 in a Swedish man after traveling to India. • It has since been detected in bacterial samples from many regions of the world. • Spread through horizontal gene transfer – bacteria picking up genes from the environment.

Where did the Carbapenem resistance come from? If carbapenem is only used in last resort medical type settings, how did the NDM-1 carbapenemase, end up in an agricultural setting? • Antibiotic resistant bacteria are becoming more common in agricultural environments because farmers often use antibiotics on their animals. • This leads to selection for resistant bacteria on some farms. • But carbapenem is not allowed to be used on farms, so where did the resistance come from? • It turns out that NDM-1 carbapenemase gives bacteria resistance to more antibiotics other than just carbapenem. • Use of other common antibiotics could select for the spread of NDM-1.

Can you help local farmers establish their risk? • The source of the infection came from a single farm. • Two neighboring farms, Barrow Creek Farm and Apple Point Farm, are now worried that the antibiotic resistance may be spreading in the environment.

Presence of the NDM-1 gene causes E. coli to be resistant to the antibiotic carbapenem Carbapenem-resistant bacteria present? Apple Point Farm Photo: Walter Baxter Barrow Creek Farm Photo: Mat Fascione
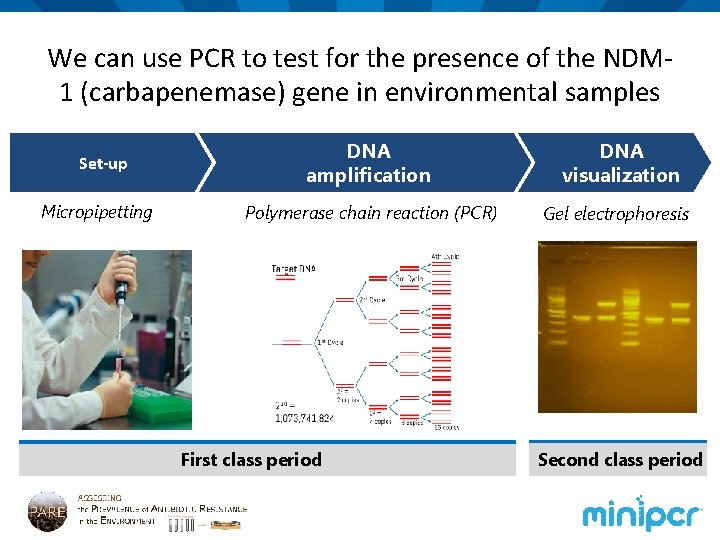
We can use PCR to test for the presence of the NDM 1 (carbapenemase) gene in environmental samples Set-up Micropipetting DNA amplification DNA visualization Polymerase chain reaction (PCR) Gel electrophoresis First class period Second class period
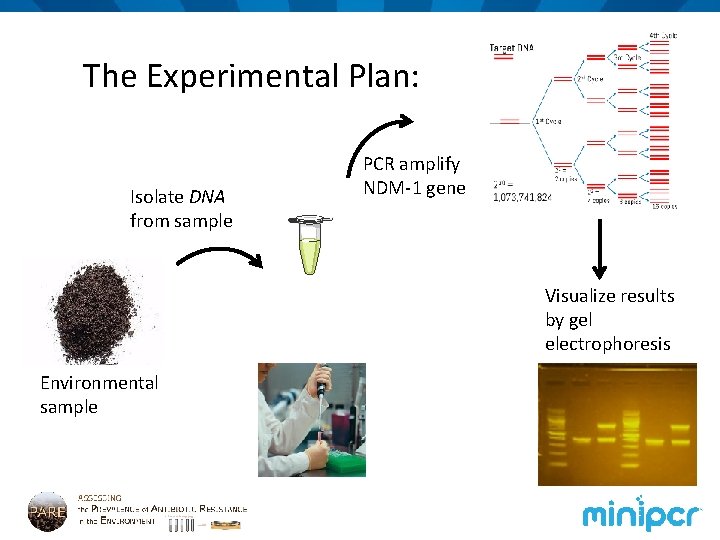
The Experimental Plan: Isolate DNA from sample PCR amplify NDM-1 gene Visualize results by gel electrophoresis Environmental sample
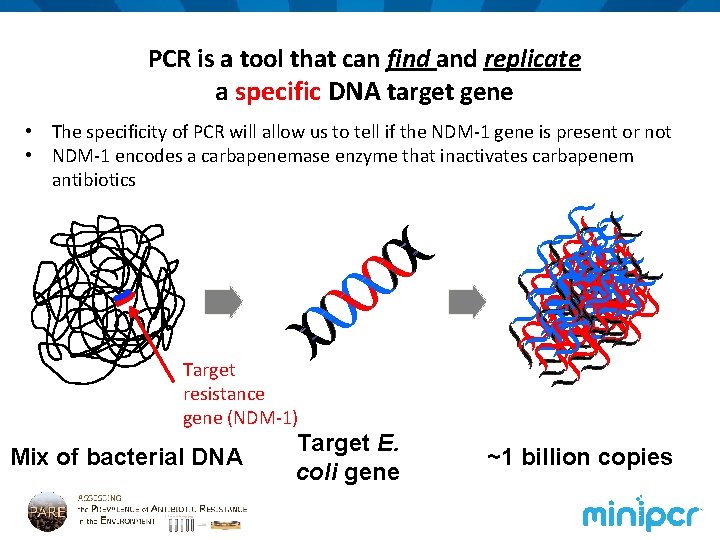
PCR is a tool that can find and replicate a specific DNA target gene • The specificity of PCR will allow us to tell if the NDM-1 gene is present or not • NDM-1 encodes a carbapenemase enzyme that inactivates carbapenem antibiotics Target resistance gene (NDM-1) Mix of bacterial DNA Target E. coli gene ~1 billion copies
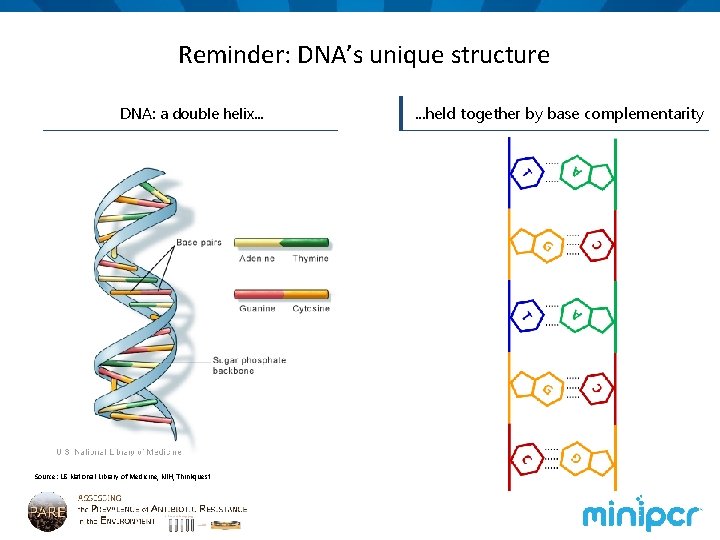
Reminder: DNA’s unique structure DNA: a double helix. . . Source: US National Library of Medicine, NIH, Thinkquest . . . held together by base complementarity
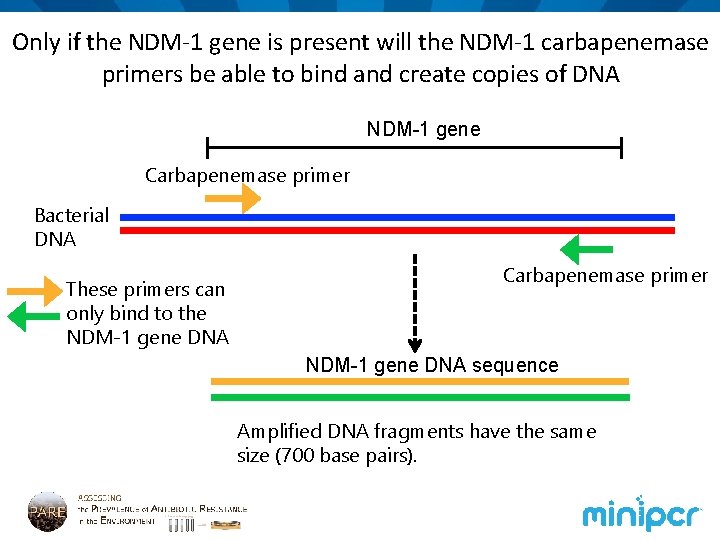
Only if the NDM-1 gene is present will the NDM-1 carbapenemase primers be able to bind and create copies of DNA NDM-1 gene Carbapenemase primer Bacterial DNA These primers can only bind to the NDM-1 gene DNA Carbapenemase primer NDM-1 gene DNA sequence Amplified DNA fragments have the same size (700 base pairs).
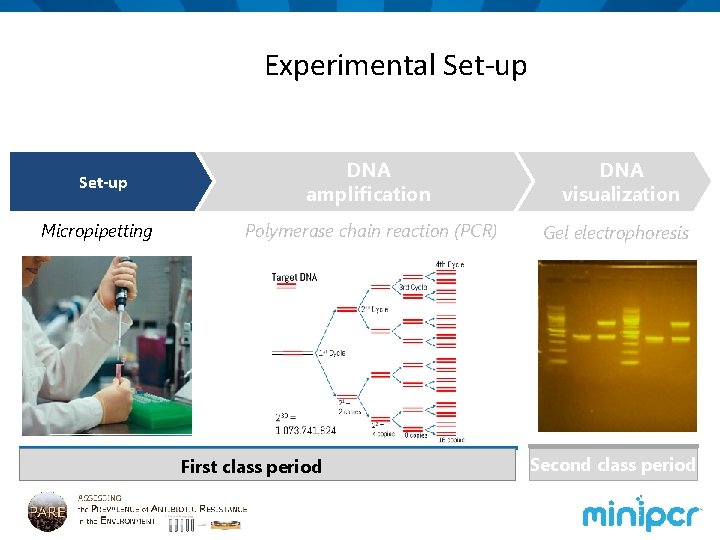
Experimental Set-up Micropipetting DNA amplification DNA visualization Polymerase chain reaction (PCR) Gel electrophoresis First class period Second class period
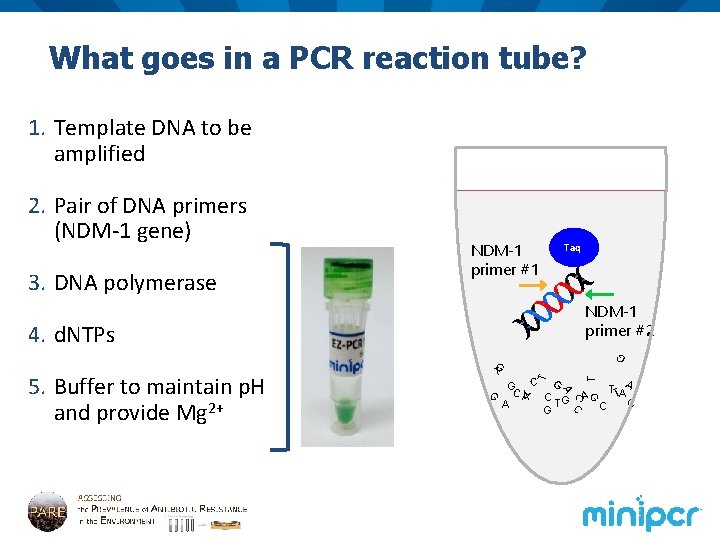
What goes in a PCR reaction tube? 1. Template DNA to be amplified 2. Pair of DNA primers (NDM-1 gene)? ? 3. DNA polymerase NDM-1 primer #1 Taq NDM-1 primer #2 T GC C G TTAA A C G AG T C A C G C C A A G 5. Buffer to maintain p. H and provide Mg 2+ G T T G 4. d. NTPs
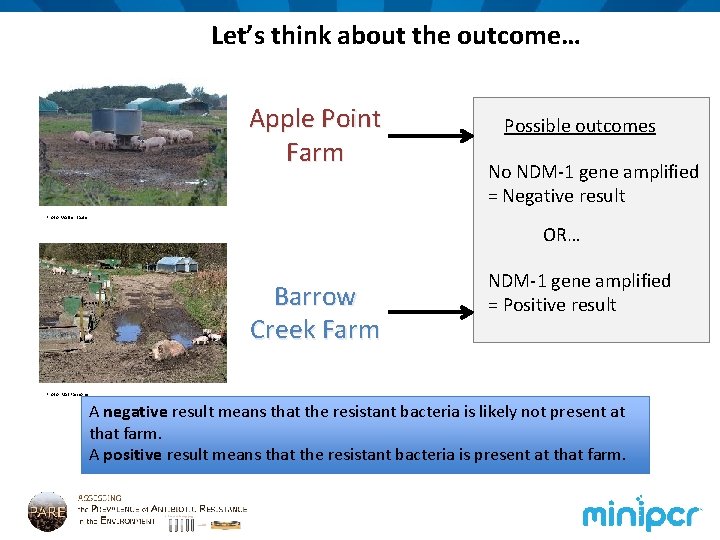
Let’s think about the outcome… Apple Point Farm Possible outcomes No NDM-1 gene amplified = Negative result Photo: Walter Baxter OR… Barrow Creek Farm NDM-1 gene amplified = Positive result Photo: Mat Fascione A negative result means that the resistant bacteria is likely not present at that farm. A positive result means that the resistant bacteria is present at that farm.

Let’s think about the outcome… How do you know that an NDM-1 negative result isn’t just because you forgot to add the Taq polymerase? We use a control to know if the reaction worked! Controls help to interpret the results

16 S ribosomal RNA gene as a control • 16 S ribosomal RNA gene is a sequence present in virtually all bacteria. • We can use second a set of primers that amplifies this sequence. • If the 16 S sequence amplifies, we know that we did our DNA extraction and PCR correctly.
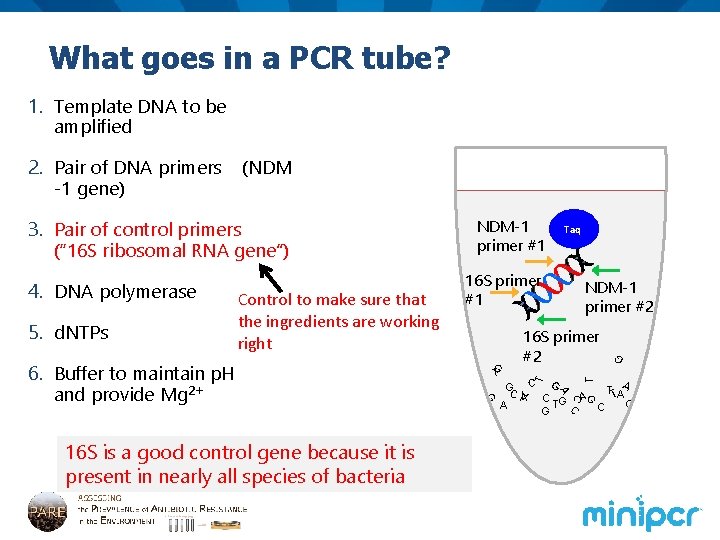
What goes in a PCR tube? 1. Template DNA to be amplified (NDM 16 S primer #2 GC C G TTAA A C G AG T C A C G C C A 16 S is a good control gene because it is present in nearly all species of bacteria G T G 6. Buffer to maintain p. H and provide Mg 2+ NDM-1 primer #2 G 5. d. NTPs Control to make sure that the ingredients are working right 16 S primer #1 Taq T 4. DNA polymerase NDM-1 primer #1 T 3. Pair of control primers (“ 16 S ribosomal RNA gene”) A 2. Pair of DNA primers -1 gene)

Experimental set up Environmental samples Apple Point Tube B Tube A Template DNA = DNA samples from Barrow Creek. A Template DNA = DNA samples from Apple Point. B Tube P (pos control) Tube N (neg control) Barrow Creek Reference lab controls P Control DNA from carbapenemresistant E. coli (NDM-1 gene present) N Control DNA from nonresistant E. coli (no NDM-1 gene)

Experimental set up More controls • We also use a positive an negative control reaction. • In the positive control we put DNA from E. coli that we know has the NDM-1 gene. • The negative control has DNA from E. coli that we know does not have the NDM-1 gene. Reference lab controls Tube P (pos control) P Control DNA from carbapenemresistant E. coli (NDM-1 gene present) Tube N (neg control) N Control DNA from nonresistant E. coli (no NDM-1 gene)

Why do we use so many controls? • Controls tell us that the experiment is working the way we expect it to, and give us something to compare out results to. This lab has two basic controls: • If the control gene doesn’t amplify, we know that there was a problem with the PCR. • The lab controls tell us that the carbapenemase primers work as we expect them to.

Warming up: Micropipetting practice You will be working with tiny volumes. For the PCR to work, it is very important that the precise amount be added to each tube.

Micropipetting practice 1. Identify volume range 2. Adjust volume 3. Press plunger to FIRST STOP 4. Press plunger to SECOND STOP

Now, transfer real liquids: 20, 10, and 5µl 1. Adjust volume 2. Get a tip 3. Press plunger to FIRST STOP 4. Collect liquid. Release plunger 5. Transfer liquid 6. Press plunger to SECOND STOP 7. Eject tip
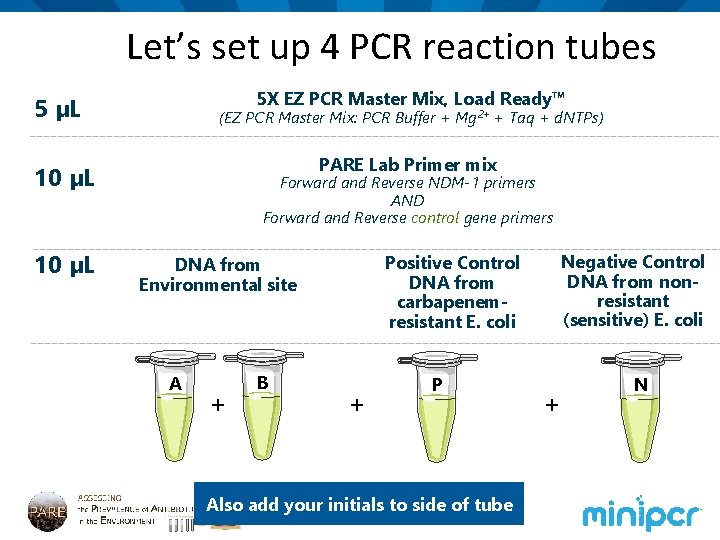
Let’s set up 4 PCR reaction tubes 5 X EZ PCR Master Mix, Load Ready™ 5 µL (EZ PCR Master Mix: PCR Buffer + Mg 2+ + Taq + d. NTPs) PARE Lab Primer mix 10 µL Forward and Reverse NDM-1 primers AND Forward and Reverse control gene primers A + B Negative Control DNA from nonresistant (sensitive) E. coli Positive Control DNA from carbapenemresistant E. coli DNA from Environmental site + P Also add your initials to side of tube + N
Experimental Set-up Micropipetting DNA amplification DNA visualization Polymerase chain reaction (PCR) Gel electrophoresis First class period Second class period
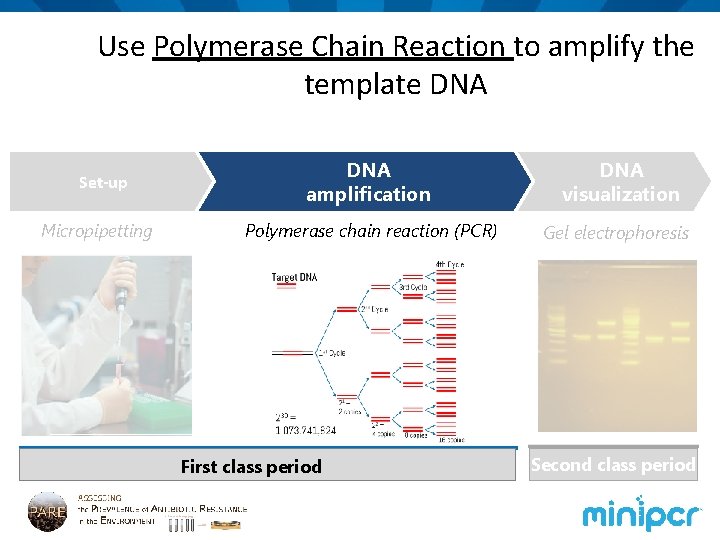
Use Polymerase Chain Reaction to amplify the template DNA Set-up Micropipetting DNA amplification DNA visualization Polymerase chain reaction (PCR) Gel electrophoresis First class period Second class period

PCR parameters Initial denaturation: 94°C 30 seconds Denaturation: Annealing Extension 94°C 55°C 72°C 10 seconds x 28 cycles Final extension 72°C 30 seconds
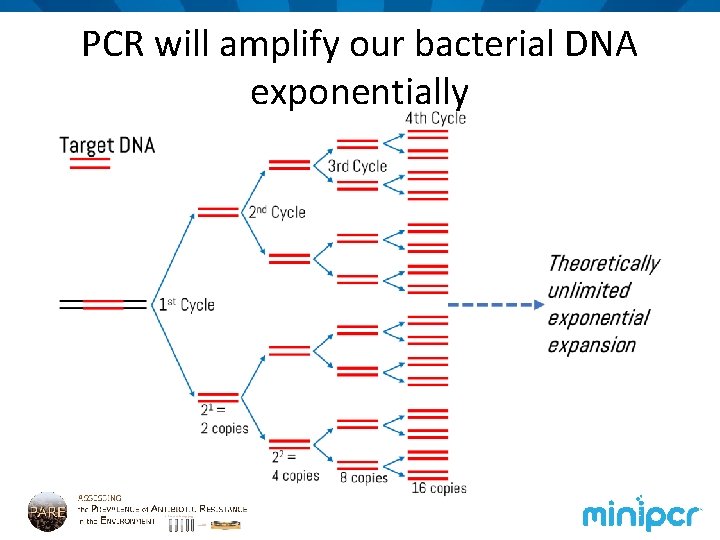
PCR will amplify our bacterial DNA exponentially
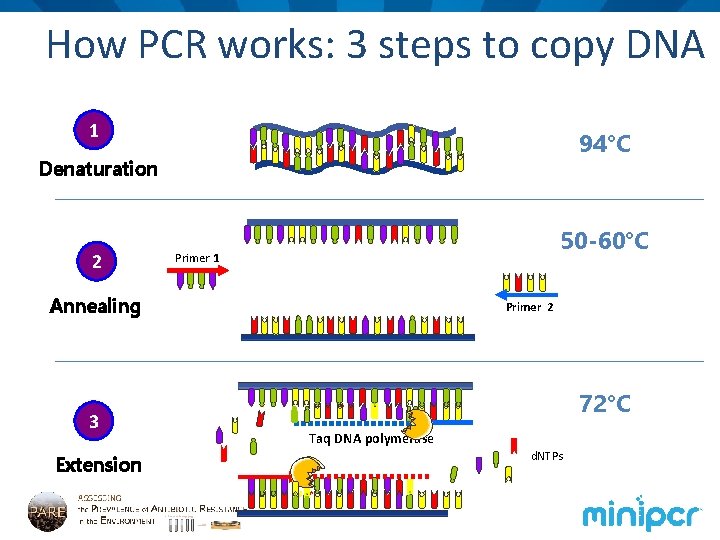
How PCR works: 3 steps to copy DNA 1 94°C Denaturation 2 50 -60°C Primer 1 Annealing 3 Extension Primer 2 72°C Taq DNA polymerase d. NTPs
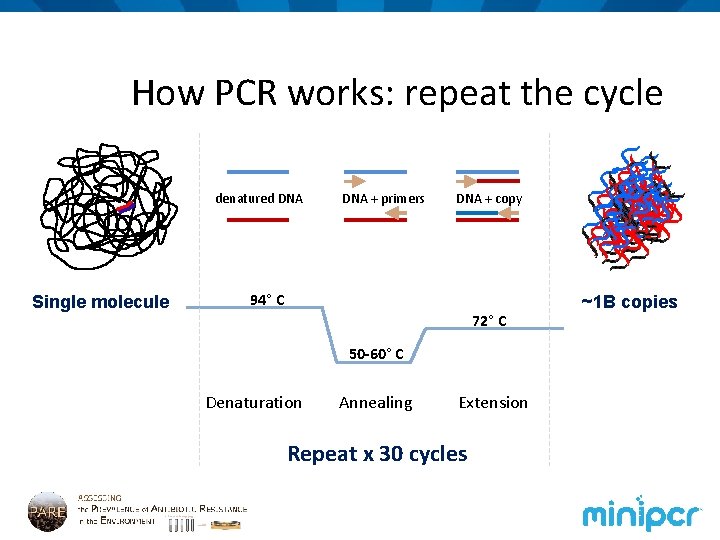
How PCR works: repeat the cycle denatured DNA Single molecule DNA + primers DNA + copy 94° C 72° C 50 -60° C Denaturation Annealing Extension Repeat x 30 cycles ~1 B copies
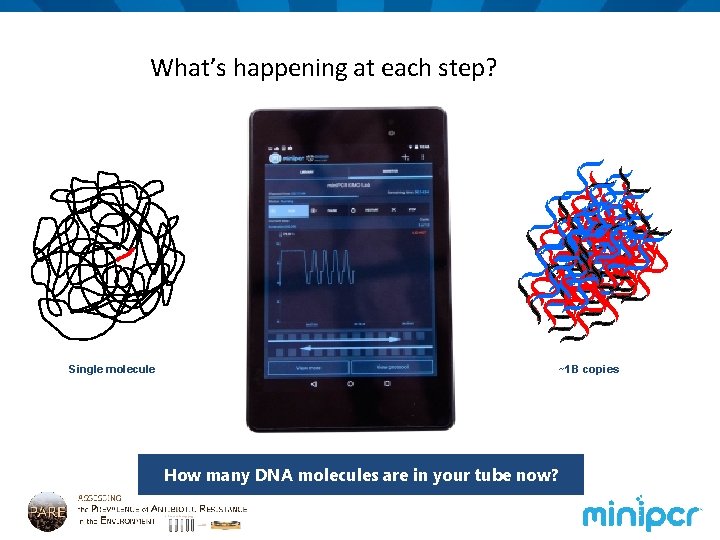
What’s happening at each step? ATP d GTP d C TP d ? primers TTP d Taq ~1 B copies Single molecule How many DNA molecules are in your tube now?

Discussion: observe mini. PCR cycling • What is happening to DNA molecules at each PCR step? • Denaturation • Annealing • Extension • What is unique about Taq DNA polymerase? • What temperature is optimal for most enzymes? • How many molecules of DNA will we have after each PCR cycle? • And at the end of the entire PCR reaction? • We call this exponential amplification
Use Polymerase Chain Reaction to amplify the template DNA Set-up Micropipetting DNA amplification DNA visualization Polymerase chain reaction (PCR) Gel electrophoresis First class period Second class period
Second class period: DNA visualization using electrophoresis Set-up Micropipetting DNA amplification DNA visualization Polymerase chain reaction (PCR) Gel electrophoresis First class period Second class period

How does electrophoresis work? DNA moves through gel polymer to positive electrode Dye added helps track movement through the polymer - Pole DNA “bands” can be visualized under blue light e + Pole DNA is negatively charged Longer molecules move more slowly Smaller molecules move faster
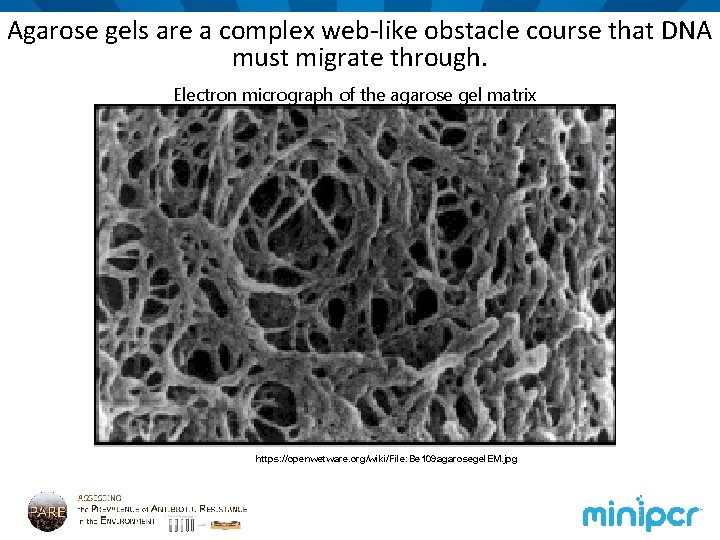
Agarose gels are a complex web-like obstacle course that DNA must migrate through. Electron micrograph of the agarose gel matrix https: //openwetware. org/wiki/File: Be 109 agarosegel. EM. jpg
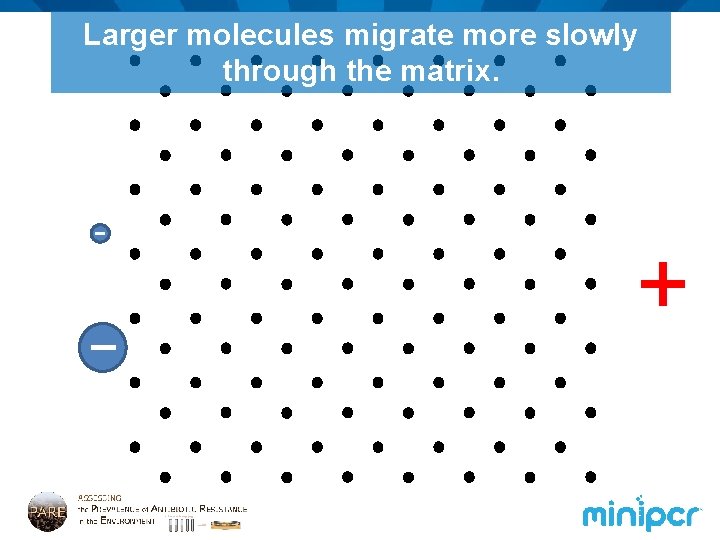
Larger molecules migrate more slowly through the matrix. +
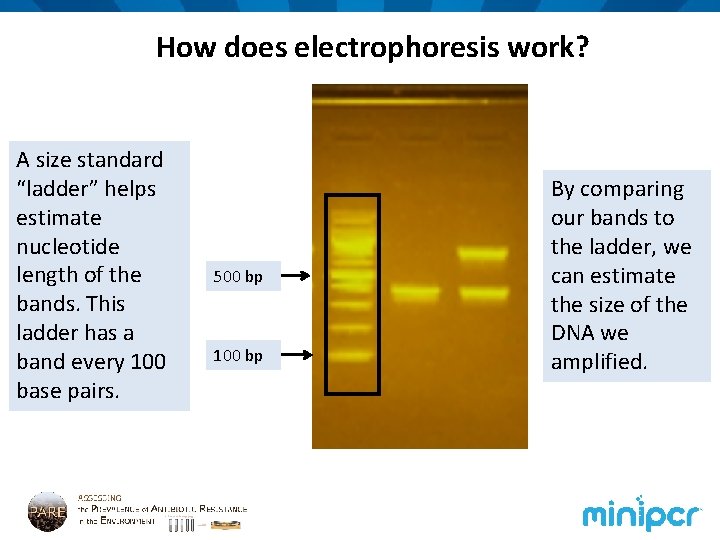
How does electrophoresis work? A size standard “ladder” helps estimate nucleotide length of the bands. This ladder has a band every 100 base pairs. 500 bp 100 bp By comparing our bands to the ladder, we can estimate the size of the DNA we amplified.
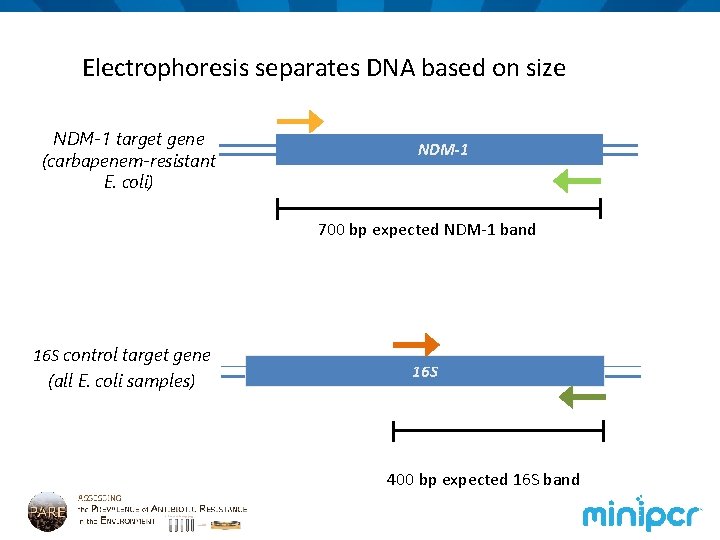
Electrophoresis separates DNA based on size NDM-1 target gene (carbapenem-resistant E. coli) NDM-1 700 bp expected NDM-1 band 16 S control target gene (all E. coli samples) 16 S 400 bp expected 16 S band

Loading the gel Load 15 µl of PCR product Negative control Environmental Sample A or B A B N Positive control P

Predict the expected results • On your worksheet: • Draw the expected band for the negative and positive controls

Expected results Apple Point: What size band(s) do you see? Barrow Creek: What size band(s) do you see? What interpretation can you draw from these results?
- Slides: 52