Microarrays Lawrence Berkeley National Lab Center for Environmental

Microarrays Lawrence Berkeley National Lab Center for Environmental Biotechnology Todd De. Santis, Sonya Murray, Jordan Moberg, Gary Andersen

The ponderings of a pre-schooler How will this lactose impact the diversity in my lower G. I. bacterial community? Will the swings be wet at the park? Will I inhale any archaeal microorganisms when I visit the hot springs? Will I be as bald as my Dad? Vincent De. Santis Why can’t I watch Shrek 3 times per day? Microarrays can help answer these types of questions.

What can they do? • Determine if a particular biological macromolecule is in a complex sample. • Examples of biological macromolecules: • Microarrays can also quantify the abundance.

One-of-a-kind foot • Molecule of interest is identified by some unique feature. • Cinderella is identified by her unique foot. • The foot is the “target” to be sought. • If her foot was common to all women, then would her foot be useful target? • Unique molecular features (targets). – – DNA or RNA sequence of ACGT(U) Proteins AA, Epitopes Lipids ? Carbohydrates branch structure

Need a perfect slipper • Molecule of interest (target) must capture, or be captured by, a second molecule (probe). • Many feet had to be evaluated by an easy test. • The specific, not common, glass slipper was used as the test (probe). • Unique molecular probes. – – DNA or RNA (Oligo nt). Proteins (Antibodies) Lipids ? Carbohydrates ? • My focus: DNA targets probed with complementary DNA
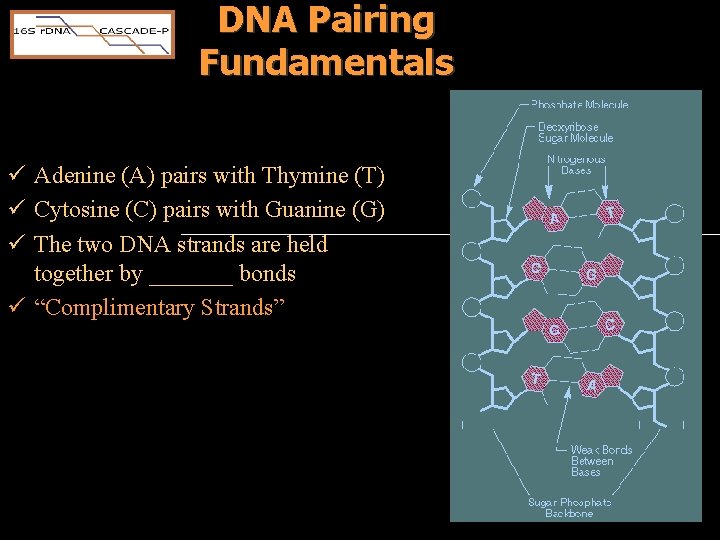
DNA Pairing Fundamentals ü Adenine (A) pairs with Thymine (T) ü Cytosine (C) pairs with Guanine (G) ü The two DNA strands are held together by _______ bonds ü “Complimentary Strands”
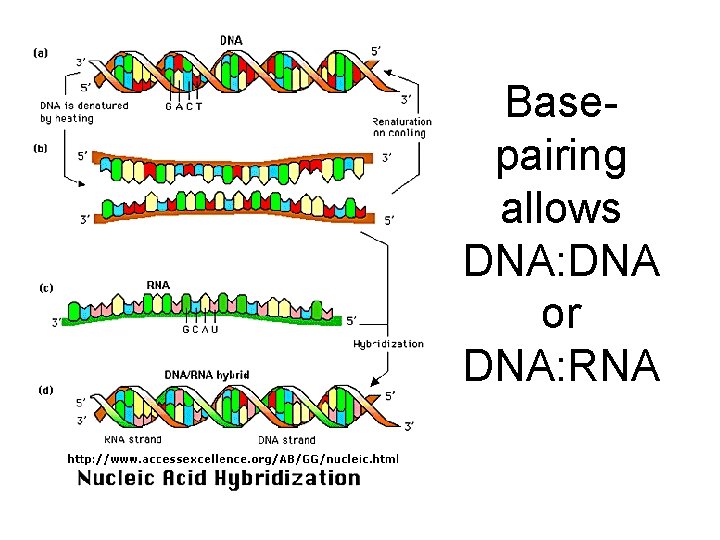
Basepairing allows DNA: DNA or DNA: RNA
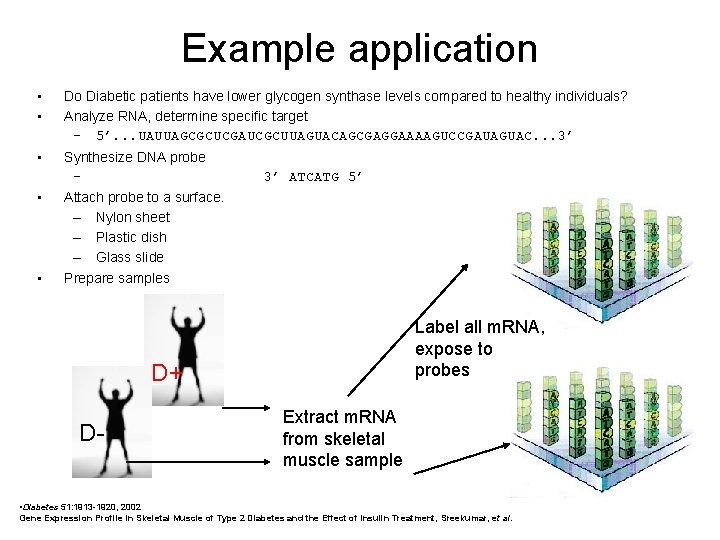
Example application • • Do Diabetic patients have lower glycogen synthase levels compared to healthy individuals? Analyze RNA, determine specific target – 5’. . . UAUUAGCGCUCGAUCGCUUAGUACAGCGAGGAAAAGUCCGAUAGUAC. . . 3’ • Synthesize DNA probe – • • 3’ ATCATG 5’ Attach probe to a surface. – Nylon sheet – Plastic dish – Glass slide Prepare samples Label all m. RNA, expose to probes D+ D- Extract m. RNA from skeletal muscle sample • Diabetes 51: 1913 -1920, 2002 Gene Expression Profile in Skeletal Muscle of Type 2 Diabetes and the Effect of Insulin Treatment, Sreekumar, et al.

Hybridization • Notice all m. RNA is labeled (florescence) • Non-binding m. RNA is washed away • If surface “glows”, then target was captured by probe. • What does it mean if no “glow” is detected? • http: //www. affymetrix. com
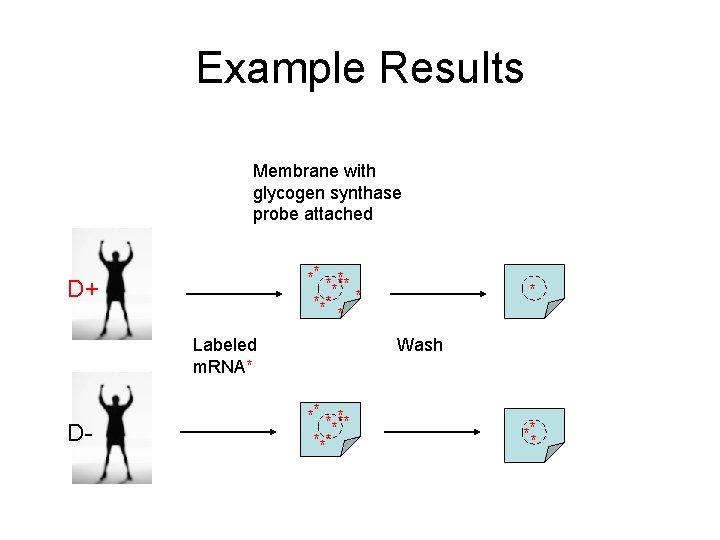
Example Results Membrane with glycogen synthase probe attached ** * *** * * D+ Labeled m. RNA* D- * Wash ** * *** ***
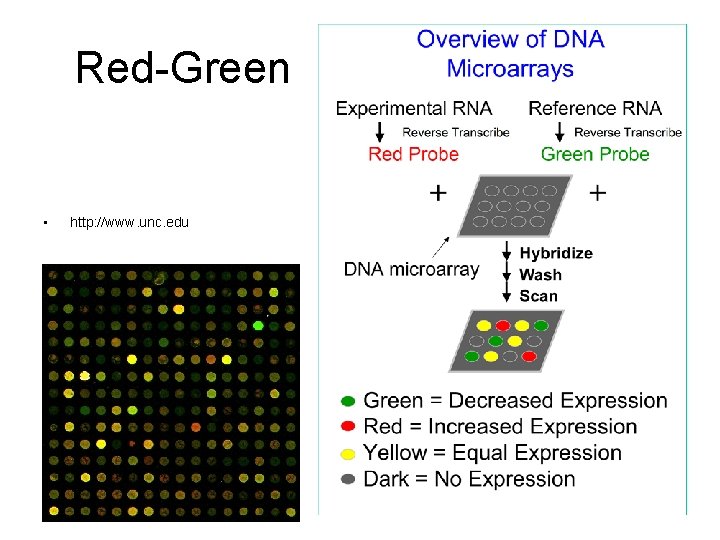
Red-Green • http: //www. unc. edu

Millions of copies per feature
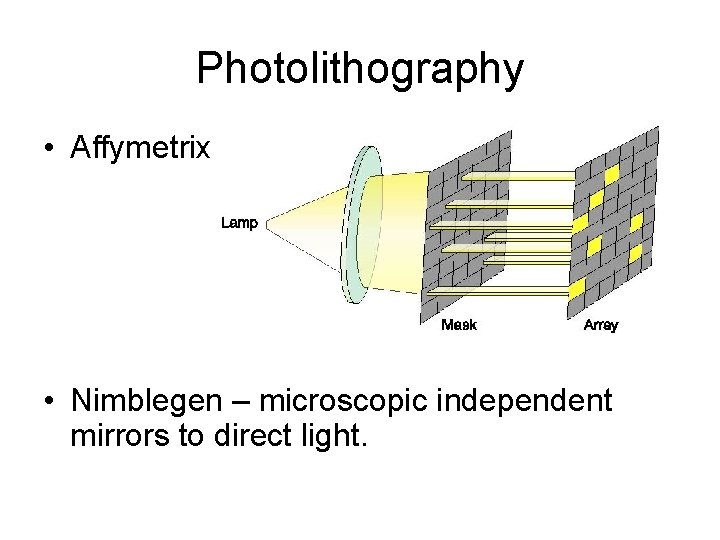
Photolithography • Affymetrix - Photolithography • Nimblegen – microscopic independent mirrors to direct light.
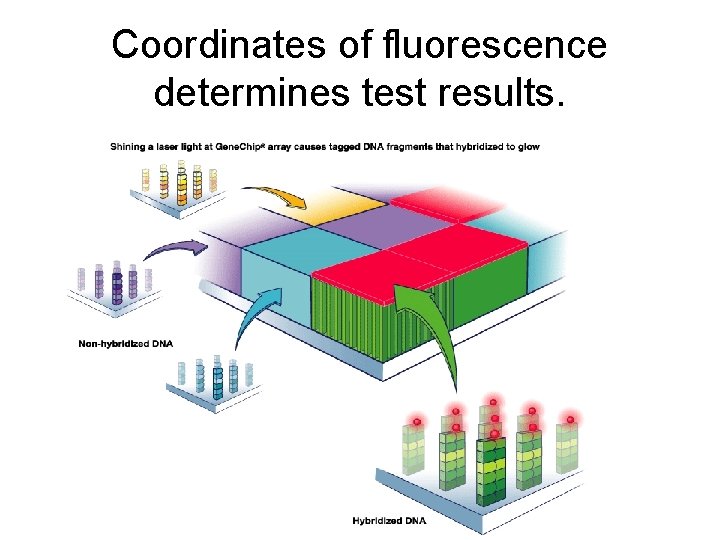
Coordinates of fluorescence determines test results.
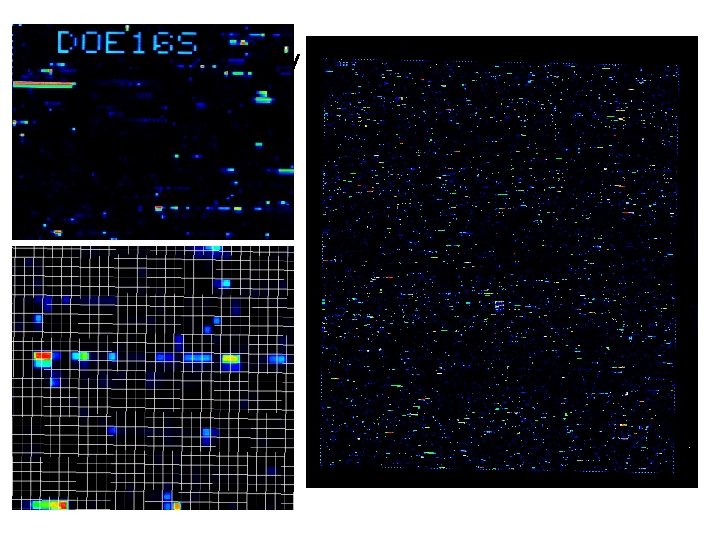
500, 000 Probe 16 S array (DOE 16 S Chip)

What “bugs” are in my sample? Rapid taxonomic classification of complex consortia of environmental r. DNA using a microarray. Environmental Surveys Counter Bio-terrorism Bioremediation Clinical Investigations
Example Microorganisms • C. immitis • B. anthracis • Plus thousands more. . Lung
Project Overview • Goal – Create a single microarray capable of detecting and categorizing the bacteria in a complex sample. • Approach – Gene. Chip targeted at 16 S r. DNA sequence variations to distinguish taxa.
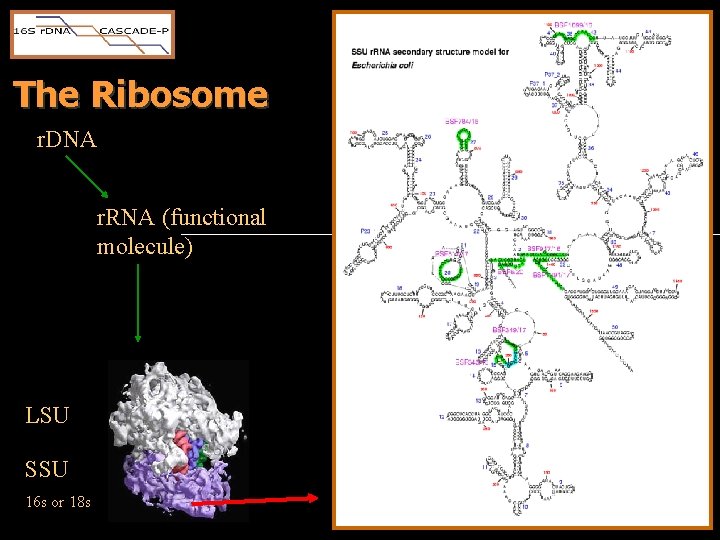
The Ribosome r. DNA r. RNA (functional molecule) LSU SSU 16 s or 18 s

• Foundations: – Maintain the largest 16 S gene library (~83, 000). – Cluster sequences into taxa (~10, 000). – Create algorithm for picking probes for each taxa (~500, 000).
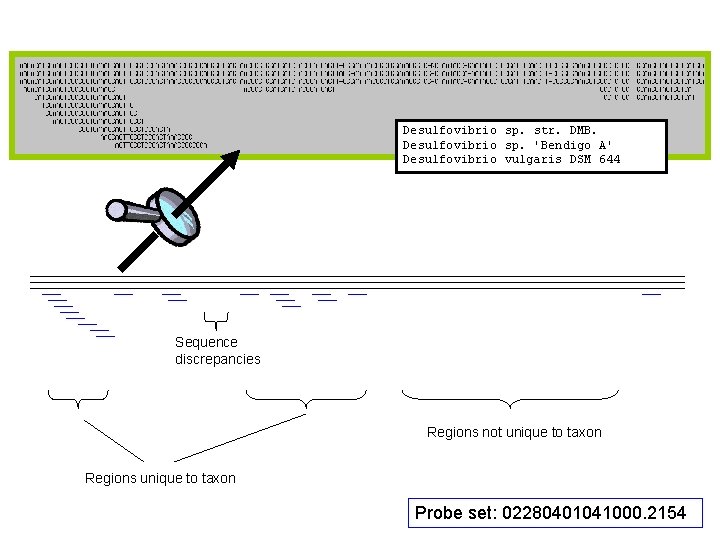
Desulfovibrio sp. str. DMB. Desulfovibrio sp. 'Bendigo A' Desulfovibrio vulgaris DSM 644 Sequence discrepancies Regions not unique to taxon Regions unique to taxon Probe set: 02280401041000. 2154
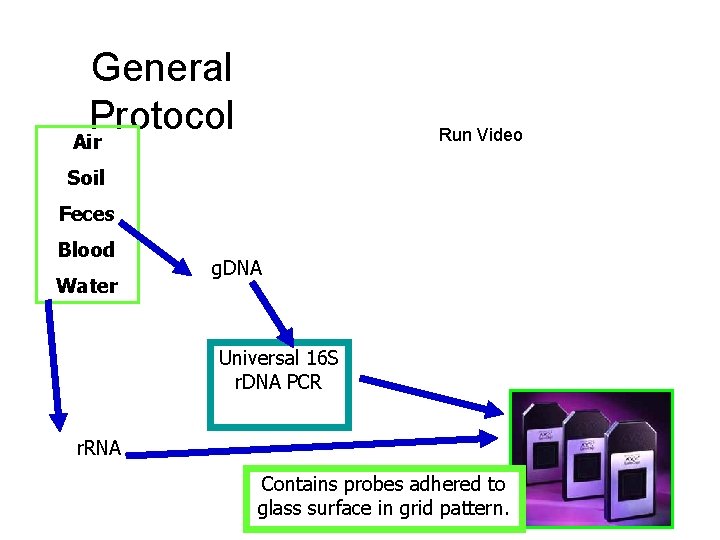
General Protocol Run Video Air Soil Feces Blood Water g. DNA Universal 16 S r. DNA PCR r. RNA Contains probes adhered to glass surface in grid pattern.
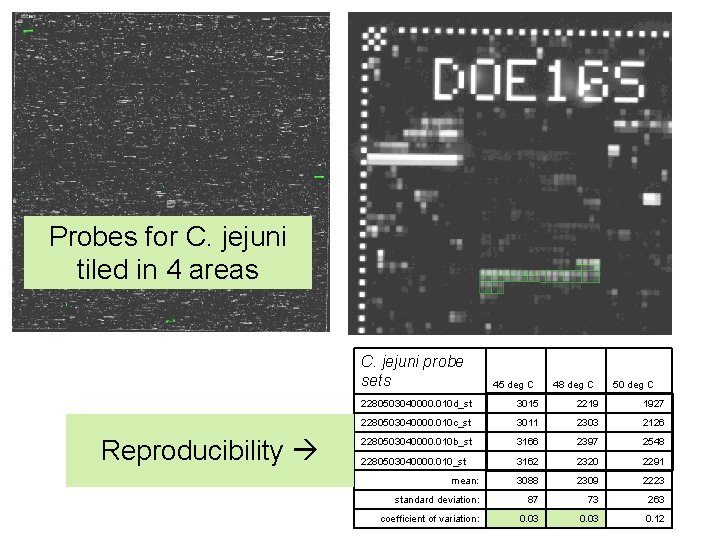
Probes for C. jejuni tiled in 4 areas C. jejuni probe sets Reproducibility 45 deg C 48 deg C 50 deg C 2280503040000. 010 d_st 3015 2219 1927 2280503040000. 010 c_st 3011 2303 2126 2280503040000. 010 b_st 3166 2397 2548 2280503040000. 010_st 3162 2320 2291 mean: 3088 2309 2223 standard deviation: 87 73 263 coefficient of variation: 0. 03 0. 12


Counter Bio-Terrorism -Sampling the Air -Extracting DNA -Hybridization -PCR -Single Organism Detection -Detection Arrays for Multiple Organisms

Traditional Detection • Collect air sample • Swab sample onto petri dish • Wait 2 days to 4 weeks for organisms to grow – Isolate – Biochemical tests – Visual Inspection

PCR • Polymerase Chain Reaction – Makes many DNA copies from few
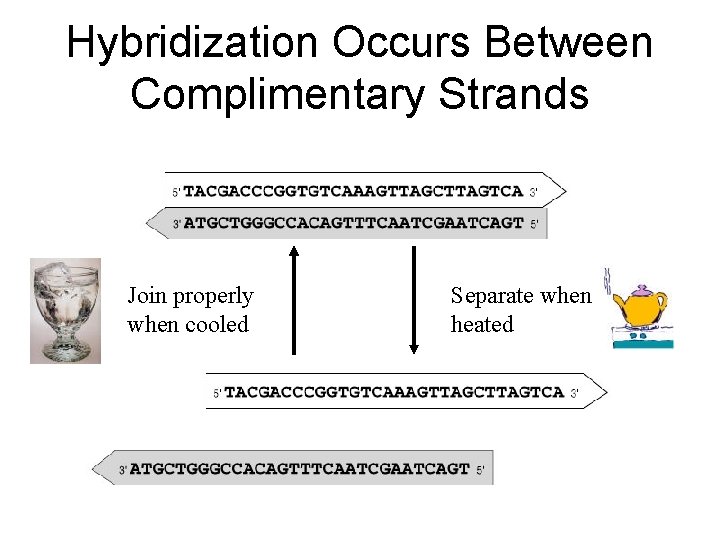
Hybridization Occurs Between Complimentary Strands Join properly when cooled Separate when heated
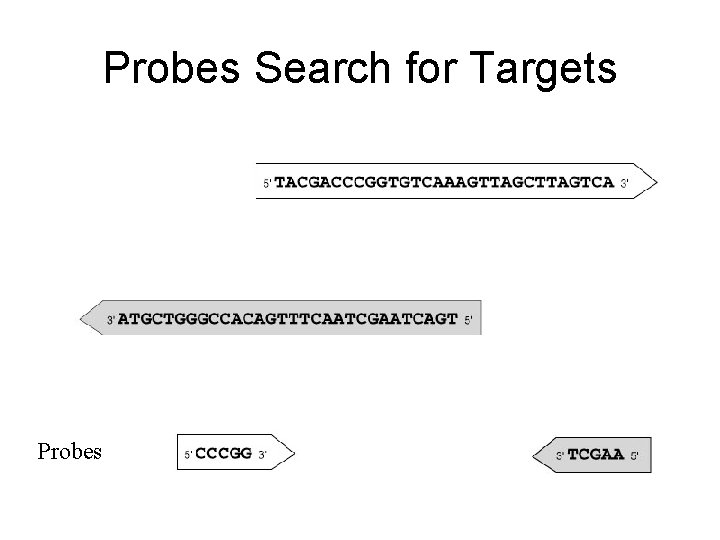
Probes Search for Targets Probes

Probes Search for Targets

DNA Replication Foundations • DNA Polymerase – enzyme which “xeroxes” a single strand in to a double strand – assembles d. NTPs monomers into a polymeric strand G G A A C TG – adds d. NTPs to 3’ end of polymer G C T G C A T Poly

Polymerase Chain Reaction (PCR) • An in vitro technique for creating many copies of a gene segment • Components – polymerase – template DNA – d. NTPs (individual A, C, G, or T) – Small probes called “primers” G Let’s do it. . . G A A C TG G T C A TG C Poly
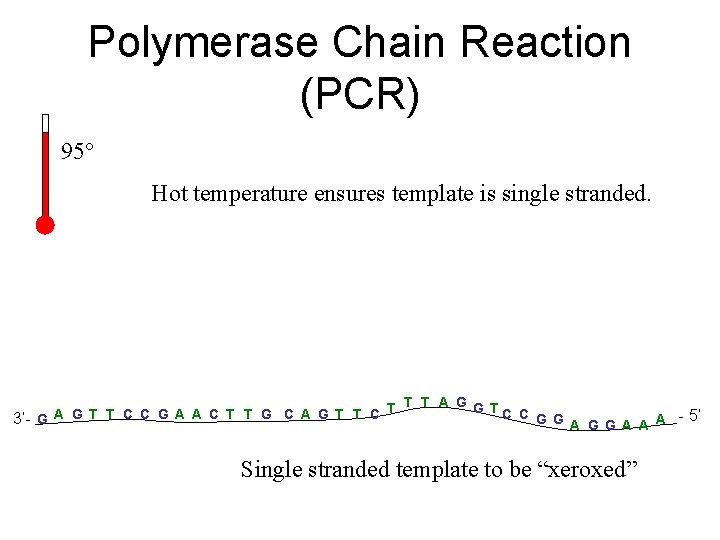
Polymerase Chain Reaction (PCR) 95° Hot temperature ensures template is single stranded. T T T A G G TC C A G T T C C G A A C T T G C A G T T C 3’- G G G A A A - 5’ Single stranded template to be “xeroxed”
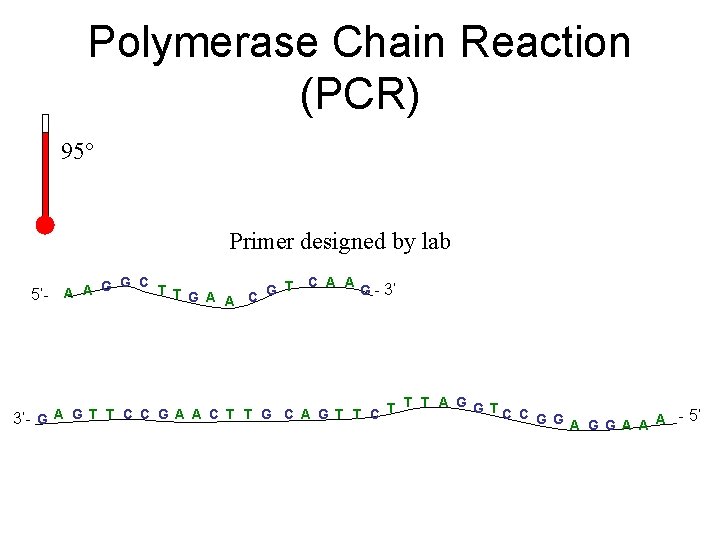
Polymerase Chain Reaction (PCR) 95° Primer designed by lab 5’- C A A G G C T G - 3’ A A T G A A C T T T A G G TC C A G T T C C G A A C T T G C A G T T C 3’- G G G A A A - 5’
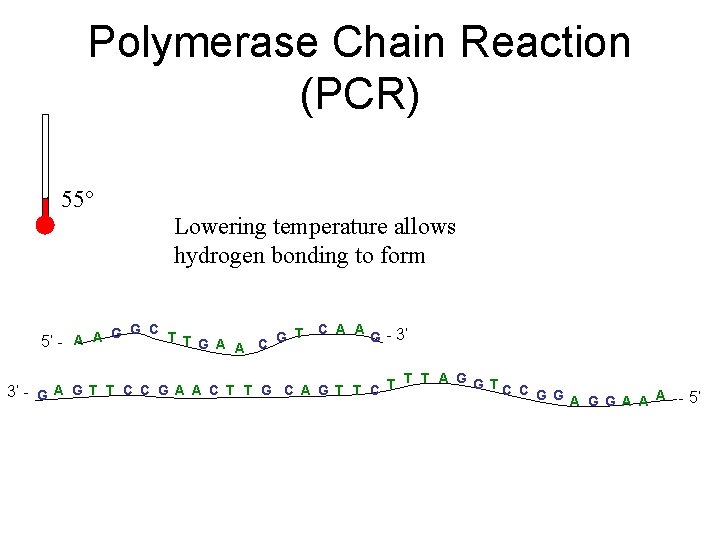
Polymerase Chain Reaction (PCR) 55° Lowering temperature allows hydrogen bonding to form C A A G G C T G - 3’ T G A 5’ - A A C A T T T A G G TC C A G T T C C G A A C T T G C A G T T C 3’ - G G G A A A - 5’
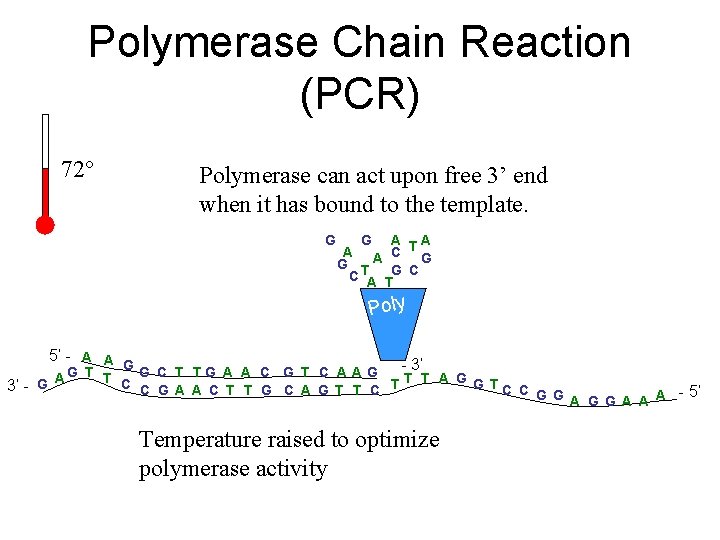
Polymerase Chain Reaction (PCR) 72° Polymerase can act upon free 3’ end when it has bound to the template. G G A A C TG G T C A TG C Poly 5’ - A A A G - 3’ G T T G C T TG A A C G T C A A G A C C G A A C T T G C A G T T C T T T A G G TC C 3’ - G G G A A A - 5’ Temperature raised to optimize polymerase activity
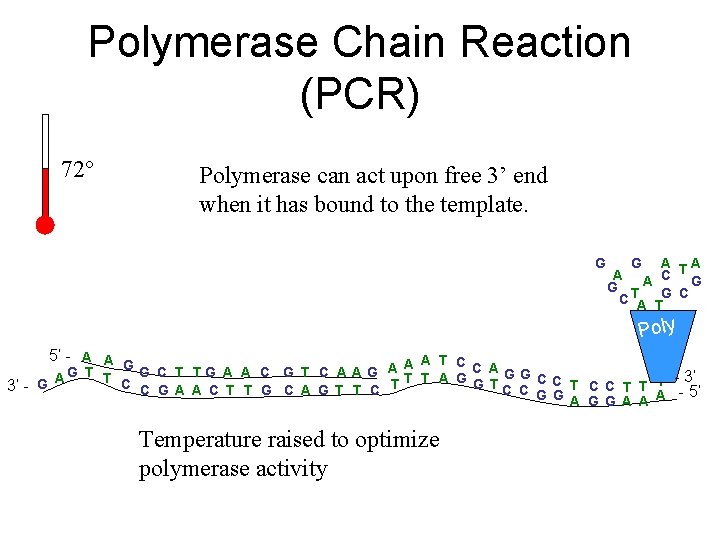
Polymerase Chain Reaction (PCR) 72° Polymerase can act upon free 3’ end when it has bound to the template. G G A A C TG G T C A TG C Poly 5’ - A A A T C G A A C AGG A G T G C T T G A A C G T C A G A T T T A G G T C C T T T - 3’ C C G A A C T T G C A G T T C T 3’ - G G G A A A - 5’ Temperature raised to optimize polymerase activity

career preparation • Do a Senior Project that involves both CS and BIO students • Find Mentor • Interview, Interview (on campus, off-campus, maintain contacts) • Learn HTML/Perl/Java/CGI

Attractive Perl Properties: · Forgiving syntax · Interpreted, not compiled · Platform independent · Text manipulation · Libraries, modules, etc. · Object oriented optional

Why Perl is the leading language of Bioinformatics? • • • Perl easy to learn Perl is powerful Perl is free Large support group: www. bio. Perl. org Interfaces easily to relational databases In fact, it has been claimed that Perl saved the Human Genome Project

Examples: • Find all the genomic 20 -mers in common between Vibrio cholera str. 14 and Vibrio mimicus V. choler 14 V. mimicus . . . TTGTACACACCGCCCGTCACACCATGGGAGTGGNCTGCAAAAGA-GCAGGTAGTTTAACC. . . TTGTACACACCGCCCGTCACACCATGGGAGTGGGCTGCAAAAGAAGCAGGTAGTTTAACC. . . • This could take a long time by hand.

Probe Finding Project • Given: – one microbial taxon • Purpose: – Describe its taxonomic placement. – Find two interesting things about the organisms in that taxa. – Find a probe that is specific for a group of organisms. • Method: – Obtain 16 S Sequences. – Align them to each other (MSA) – Determine best target from a short list (provided). • Verify that probe exists in all/most orgs of taxa • Check for X-hybe (non-specificity) – Check w/in sub-division, not all sequences
Start Here

Answers (Targets) • Oceanospirillum – TGCTACTTCGCCGGCGAGCGGCGGA • Streptococcus – CTTGACATCCTTCTGACCGGCCTAG • Rhodococcus – CGGGTCTCTGGGAAACAACTGACGC – TGGGAAACAACTGACGCTGAGGAAC • Methanocorpusculum – TGGAGAATACTCCCGGGAAACTGGG • Desulfovibrio – GCGTGAAAGGACTTCGGTCCGAGTA

Multi Microarray Analysis • Track a bio-marker over many experiments. • Download file – Fluorescence intensity for 118 taxa reported for 5 experimental conditions, and a negative control. • Find – – Taxa with highest intensity overall. Possible PCR contaminants Taxa with greatest intensity fluctuation overall Condition producing the brightest signal for Paracoccus yeeii?
- Slides: 45