MICROARRAY TECHNOLOGY By Sehreen Saddique Ibtesam Riaz Bakhtawer

MICROARRAY TECHNOLOGY By Sehreen Saddique Ibtesam Riaz Bakhtawer Nasrullah University of Gujrat

OUTLINE 1. Introduction of microarray 2. Types of microarray 3. Applications of microarray

Y A R R DN A O R C I M A
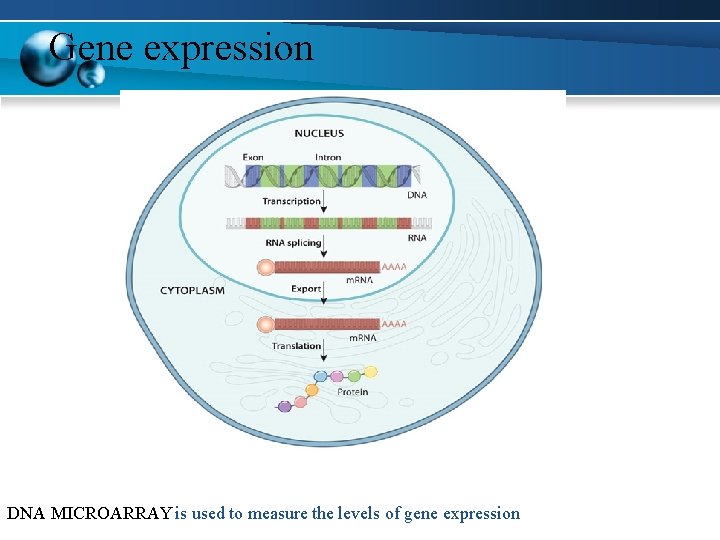
Gene expression DNA MICROARRAY is used to measure the levels of gene expression

If the hybridised DNA or RNA is labelled fluorescently it can be quantified by scanning of the chip.

DNA microarrays can be manufactured by: • Photolithography • Robot spotting
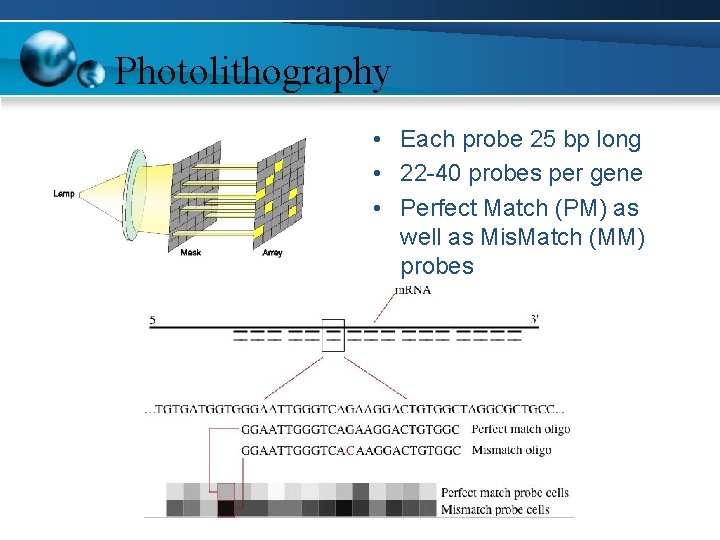
Photolithography • Each probe 25 bp long • 22 -40 probes per gene • Perfect Match (PM) as well as Mis. Match (MM) probes
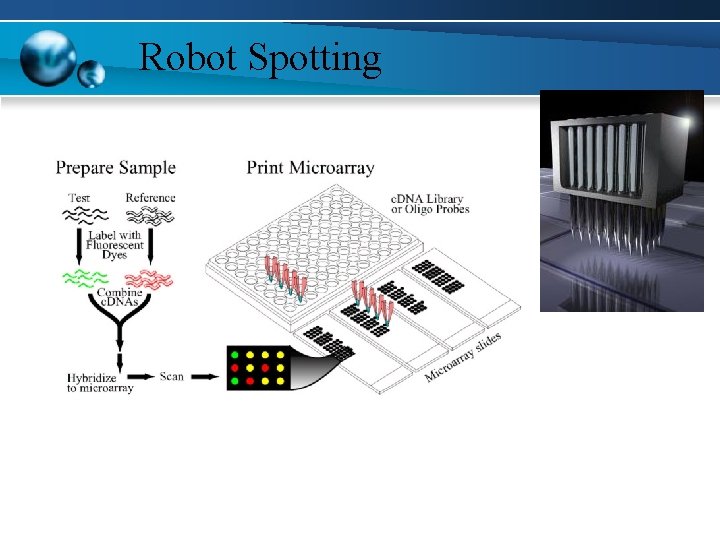
Robot Spotting

DNA MICROARRAY TECHNIQUE • Presynthesized DNA strands or synthesis of oligonuleotide sequence • Spotting of DNA strands to plastic plate or glass plate

DNA MICROARRAY TECHNIQUE • The two samples to be compared are grown/acquired. In this example treated sample (case) and untreated sample (control). • Extraction of m. RNA material from cells • Formation of complementary DNA through reverse transcription • Labeling with fluorescent dyes with Cy 3 or Cy 5 • Mixing of labeled samples & presenting to microarray spots again by spotters • Hybridization to a DNA Microarray • Scanning the Hybridized Array
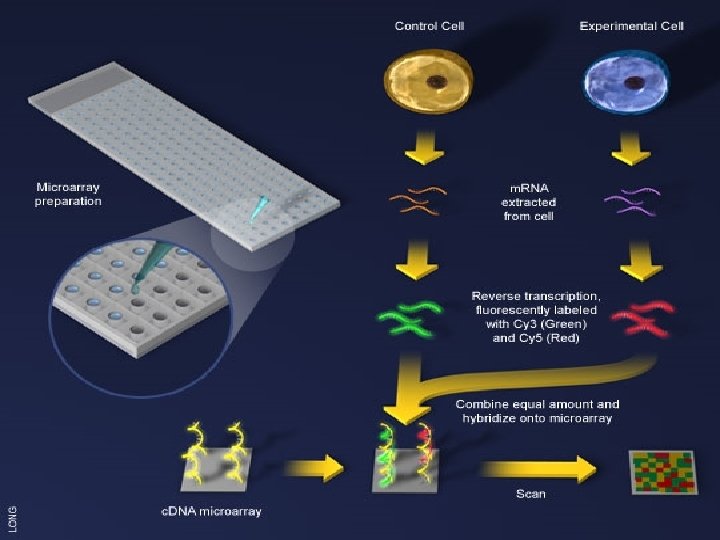


ANALYSIS OF GENE EXPRESSION DATA • Analyzing issues • Different environmental conditions

TYPES OF MICROARRAY SEHREEN SADDIQUE ROLL NO: 08030614 -045

Types of microarray • There are two types of DNA microarray • c DNA • Oligoneuclotide array
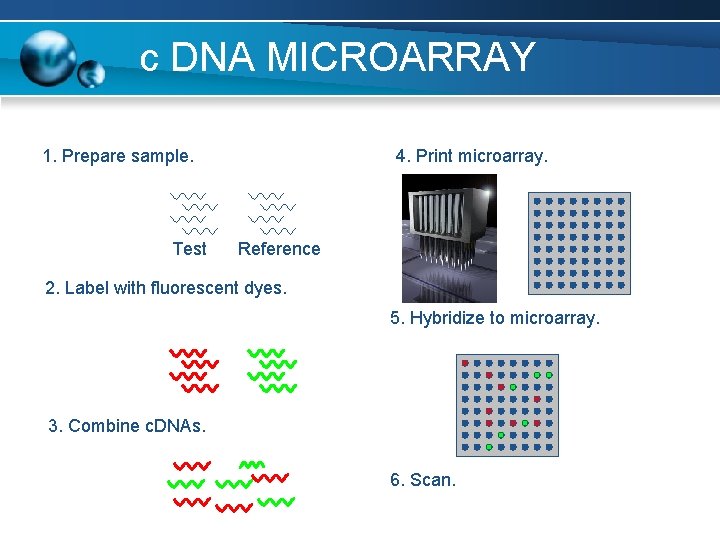
c DNA MICROARRAY 1. Prepare sample. Test 4. Print microarray. Reference 2. Label with fluorescent dyes. 5. Hybridize to microarray. 3. Combine c. DNAs. 6. Scan.
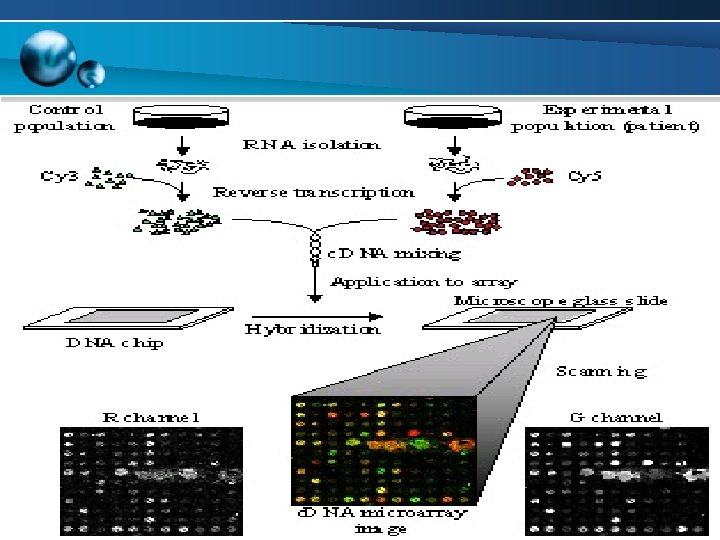
Nelzo Ereful Crop Bioinformatics
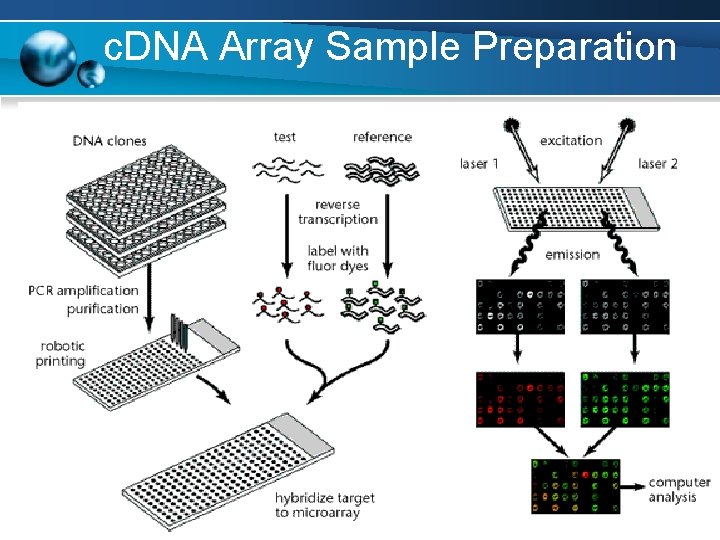
c. DNA Array Sample Preparation



APPLICATIONS OF MICROOARRAY BAKHTAWER ROLL: 08030614 -047
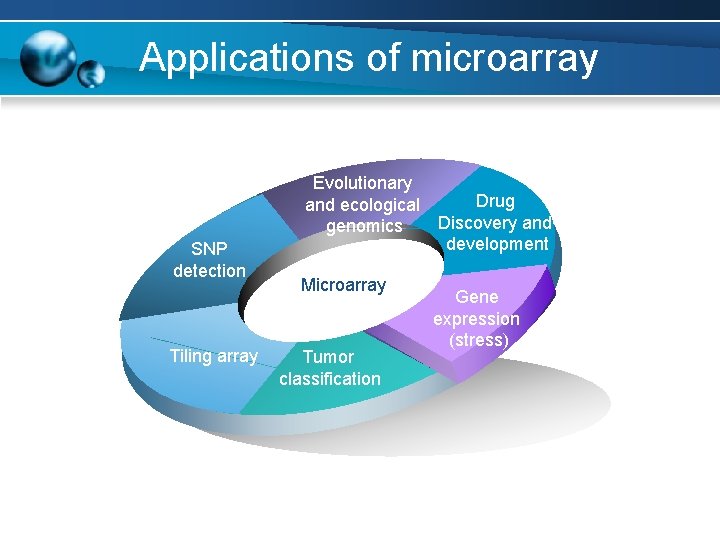
Applications of microarray Evolutionary and ecological genomics SNP detection Tiling array Microarray Tumor classification Drug Discovery and development Gene expression (stress)

Applications of microarray
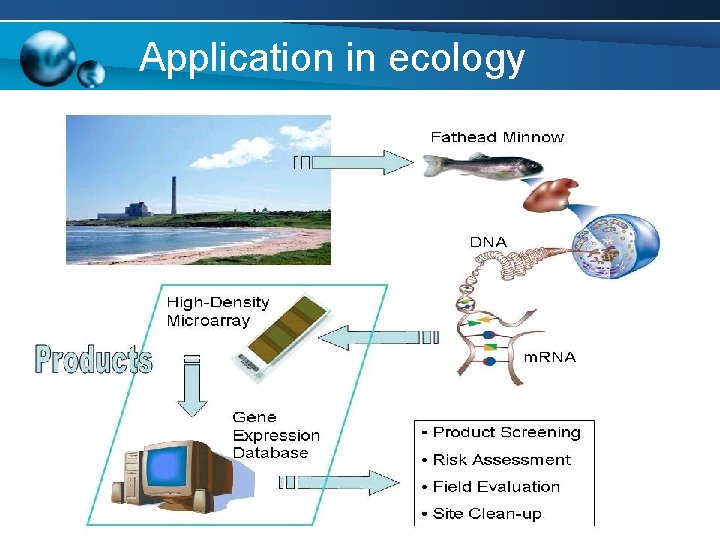
Application in ecology
Disease discovery Microarray and asthma
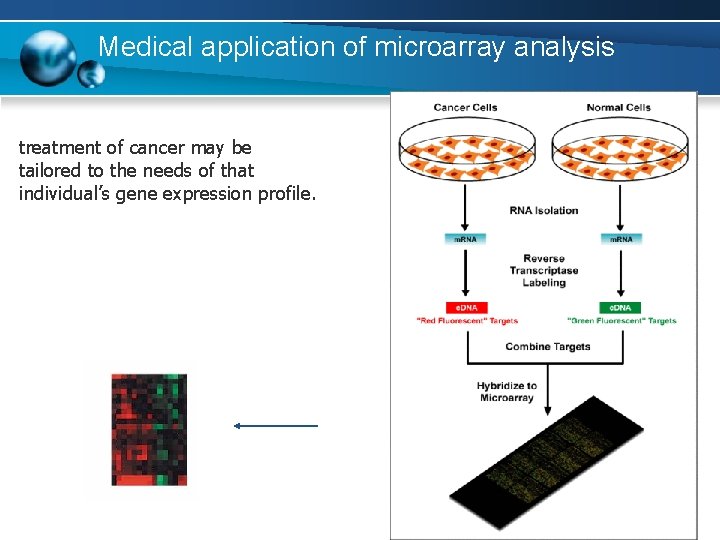
Medical application of microarray analysis treatment of cancer may be tailored to the needs of that individual’s gene expression profile.
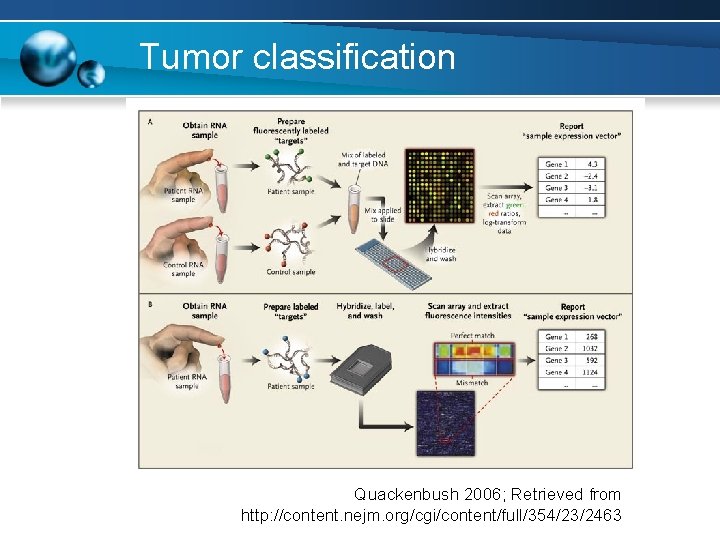
Tumor classification Quackenbush 2006; Retrieved from http: //content. nejm. org/cgi/content/full/354/23/2463
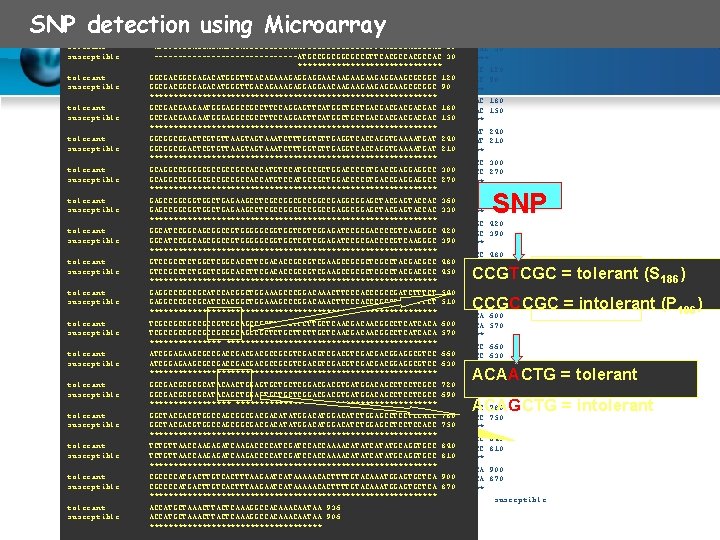
SNP detection using Microarray tolerant ATGTGTGGAGGAGAAGTGATCCCCGCCGACATGCCGGCGGCGCCGTTCACGCCAC 60 tolerant susceptible ATGTGTGGAGGAGAAGTGATCCCCGCCGACATGCCGGCGGCGCCGTTCACGCCAC 60 ---------------ATGCCGGCGGCGCCGTTCACGCCAC 30 susceptible ---------------ATGCCGGCGGCGCCGTTCACGCCAC 30 ****************************** tolerant GGCGACGGCGAGACATGGGTTGACAGAAAGAGGAGGAACAAGAAGAAGAGGAAGCGCGGC 120 tolerant susceptible GGCGACGGCGAGACATGGGTTGACAGAAAGAGGAGGAACAAGAAGAAGAGGAAGCGCGGC 120 GGCGACGGCGAGACATGGGTTGACAGAAAGAGGAGGAACAAGAAGAAGAGGAAGCGCGGC 90 susceptible GGCGACGGCGAGACATGGGTTGACAGAAAGAGGAGGAACAAGAAGAAGAGGAAGCGCGGC 90 ************************************************************ tolerant GCCGACGAAGAATGGGAGGCCGCCTTCCAGGAGTTCATGGCTGCTGACGACGAC 180 tolerant susceptible GCCGACGAAGAATGGGAGGCCGCCTTCCAGGAGTTCATGGCTGCTGACGACGAC 180 GCCGACGAAGAATGGGAGGCCGCCTTCCAGGAGTTCATGGCTGCTGACGACGAC 150 susceptible GCCGACGAAGAATGGGAGGCCGCCTTCCAGGAGTTCATGGCTGCTGACGACGAC 150 ************************************************************ tolerant GGCGGCGGACTCGTGTTAAGTAGTAAATCTTTGGTGTTGAGGTCACCAGGTGAAAATGAT 240 tolerant susceptible GGCGGCGGACTCGTGTTAAGTAGTAAATCTTTGGTGTTGAGGTCACCAGGTGAAAATGAT 240 GGCGGCGGACTCGTGTTAAGTAGTAAATCTTTGGTGTTGAGGTCACCAGGTGAAAATGAT 210 susceptible GGCGGCGGACTCGTGTTAAGTAGTAAATCTTTGGTGTTGAGGTCACCAGGTGAAAATGAT 210 ************************************************************ tolerant GCAGGCCGGGGCGCCGCCGCCACCATGTCCATGCCGCTGGACCCCGTGACCGAGGAGGCC 300 tolerant susceptible GCAGGCCGGGGCGCCGCCGCCACCATGTCCATGCCGCTGGACCCCGTGACCGAGGAGGCC 300 GCAGGCCGGGGCGCCGCCGCCACCATGTCCATGCCGCTGGACCCCGTGACCGAGGAGGCC 270 susceptible GCAGGCCGGGGCGCCGCCGCCACCATGTCCATGCCGCTGGACCCCGTGACCGAGGAGGCC 270 ************************************************************ tolerant GAGCCGGCGGTGGCTGAGAAGCCTCGCCGGCCGAGGCGGAGCTACGAGTACCAC 360 tolerant susceptible GAGCCGGCGGTGGCTGAGAAGCCTCGCCGGCCGAGGCGGAGCTACGAGTACCAC 360 GAGCCGGCGGTGGCTGAGAAGCCTCGCCGGCCGAGGCGGAGCTACGAGTACCAC 330 susceptible GAGCCGGCGGTGGCTGAGAAGCCTCGCCGGCCGAGGCGGAGCTACGAGTACCAC 330 ************************************************************ tolerant GGCATCCGGCAGCGGCCGTGGGGGCGGTGGTCGTCGGAGATCCGCGACCCCGTCAAGGGC 420 tolerant susceptible GGCATCCGGCAGCGGCCGTGGGGGCGGTGGTCGTCGGAGATCCGCGACCCCGTCAAGGGC 420 GGCATCCGGCAGCGGCCGTGGGGGCGGTGGTCGTCGGAGATCCGCGACCCCGTCAAGGGC 390 susceptible GGCATCCGGCAGCGGCCGTGGGGGCGGTGGTCGTCGGAGATCCGCGACCCCGTCAAGGGC 390 ************************************************************ tolerant GTCCGCCTCTGGCTCGGCACCTTCGACACCGCCGTCGAAGCCGCGCTCGCCTACGACGCC 480 tolerant susceptible GTCCGCCTCTGGCTCGGCACCTTCGACACCGCCGTCGAAGCCGCGCTCGCCTACGACGCC 480 GTCCGCCTCTGGCTCGGCACCTTCGACACCGCCGTCGAAGCCGCGCTCGCCTACGACGCC 450 susceptible GTCCGCCTCTGGCTCGGCACCTTCGACACCGCCGTCGAAGCCGCGCTCGCCTACGACGCC 450 ************************************************************ tolerant GAGGCCCGCCGCATCCACGGCTGGAAAGCCCGGACAAACTTCCCACCCGCCGATCTTTCT 540 tolerant susceptible GAGGCCCGCCGCATCCACGGCTGGAAAGCCCGGACAAACTTCCCACCCGCCGATCTTTCT 540 GAGGCCCGCCGCATCCACGGCTGGAAAGCCCGGACAAACTTCCCACCCGCCGATCTTTCT 510 susceptible GAGGCCCGCCGCATCCACGGCTGGAAAGCCCGGACAAACTTCCCACCCGCCGATCTTTCT 510 ************************************************************ tolerant TCGCCGCCGTCGCAGCCGCTCTGCTTCTTGCTCAACGACAACGGCCTCATCACA 600 tolerant susceptible TCGCCGCCGTCGCAGCCGCTCTGCTTCTTGCTCAACGACAACGGCCTCATCACA 600 TCGCCGCCGCAGCCGCTCTGCTTCTTGCTCAACGACAACGGCCTCATCACA 570 susceptible TCGCCGCCGCAGCCGCTCTGCTTCTTGCTCAACGACAACGGCCTCATCACA 570 ******************************************** tolerant ATCGGAGAAGCGCCGACGACGCCGCGTCGACGTCGACGACGGAGGCGTCC 660 tolerant susceptible ATCGGAGAAGCGCCGACGACGCCGCGTCGACGTCGACGACGGAGGCGTCC 660 ATCGGAGAAGCGCCGACGACGCCGCGTCGACGTCGACGACGGAGGCGTCC 630 susceptible ATCGGAGAAGCGCCGACGACGCCGCGTCGACGTCGACGACGGAGGCGTCC 630 ************************************************************ tolerant GGCGACGCGCGCATACAACTGGAGTGCTGCTCGGACGACGTGATGGACAGCCTCCTCGCC 720 tolerant susceptible GGCGACGCGCGCATACAACTGGAGTGCTGCTCGGACGACGTGATGGACAGCCTCCTCGCC 720 GGCGACGCGCGCATACAGCTGGAGTGCTGCTCGGACGACGTGATGGACAGCCTCCTCGCC 690 susceptible GGCGACGCGCGCATACAGCTGGAGTGCTGCTCGGACGACGTGATGGACAGCCTCCTCGCC 690 ****************************************** tolerant GGCTACGACGTGGCCAGCGGCGACGACATATGGACATCTGGAGCCTCCTCCACC 780 tolerant susceptible GGCTACGACGTGGCCAGCGGCGACGACATATGGACATCTGGAGCCTCCTCCACC 780 GGCTACGACGTGGCCAGCGGCGACGACATATGGACATCTGGAGCCTCCTCCACC 750 susceptible GGCTACGACGTGGCCAGCGGCGACGACATATGGACATCTGGAGCCTCCTCCACC 750 ************************************************************ tolerant TCTGTTAACCAAGAGATCAAGACCCCATCGATCCACCAAAACATATGCAGGTGCC 840 tolerant susceptible TCTGTTAACCAAGAGATCAAGACCCCATCGATCCACCAAAACATATGCAGGTGCC 840 TCTGTTAACCAAGAGATCAAGACCCCATCGATCCACCAAAACATATGCAGGTGCC 810 susceptible TCTGTTAACCAAGAGATCAAGACCCCATCGATCCACCAAAACATATGCAGGTGCC 810 ************************************************************ tolerant CGCCCCATGACTTGTCACTTTAAGAATCATAAAAACACTTTTGTACAAATGGAGTGCTCA 900 tolerant susceptible CGCCCCATGACTTGTCACTTTAAGAATCATAAAAACACTTTTGTACAAATGGAGTGCTCA 900 CGCCCCATGACTTGTCACTTTAAGAATCATAAAAACACTTTTGTACAAATGGAGTGCTCA 870 susceptible CGCCCCATGACTTGTCACTTTAAGAATCATAAAAACACTTTTGTACAAATGGAGTGCTCA 870 ************************************************************ tolerant ACCATGCTAAACTTACTCAAAGGCCACAATAA 936 susceptible tolerant ACCATGCTAAACTTACTCAAAGGCCACAAACAATAA 936 906 susceptible ACCATGCTAAACTTACTCAAAGGCCACAATAA 906 ************************************ SNP CCGTCGC = tolerant (S 186) CCGCCGC = intolerant (P 186) ACAACTG = tolerant ACAGCTG = intolerant
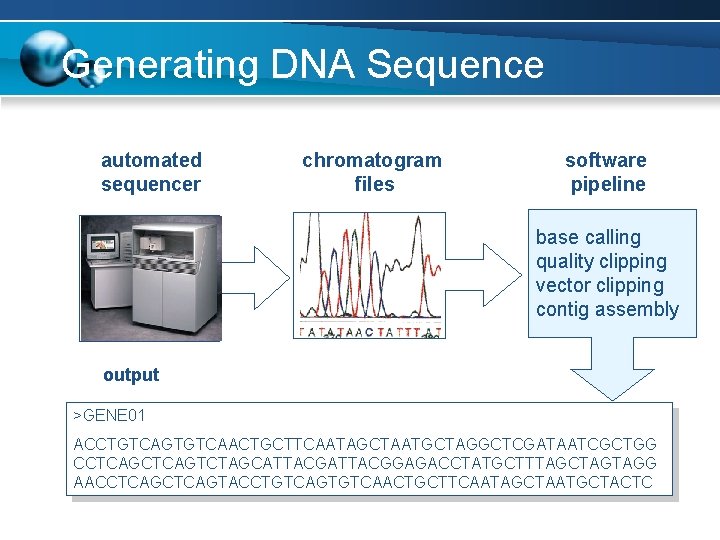
Generating DNA Sequence automated sequencer chromatogram files software pipeline base calling quality clipping vector clipping contig assembly output >GENE 01 ACCTGTCAGTGTCAACTGCTTCAATAGCTAATGCTAGGCTCGATAATCGCTGG CCTCAGTCTAGCATTACGGAGACCTATGCTTTAGCTAGTAGG AACCTCAGTACCTGTCAGTGTCAACTGCTTCAATAGCTAATGCTACTC

- Slides: 30