Microarray Introduction Content Biology background of microarray Design

Microarray Introduction

Content • • • Biology background of microarray Design of microarray The workflow of microarray Image analysis of microarray Data analysis of microarray Discussion

The Biology Background of Microarray • • The central dogma of life forms DNA RNA Monitoring the expression of genes
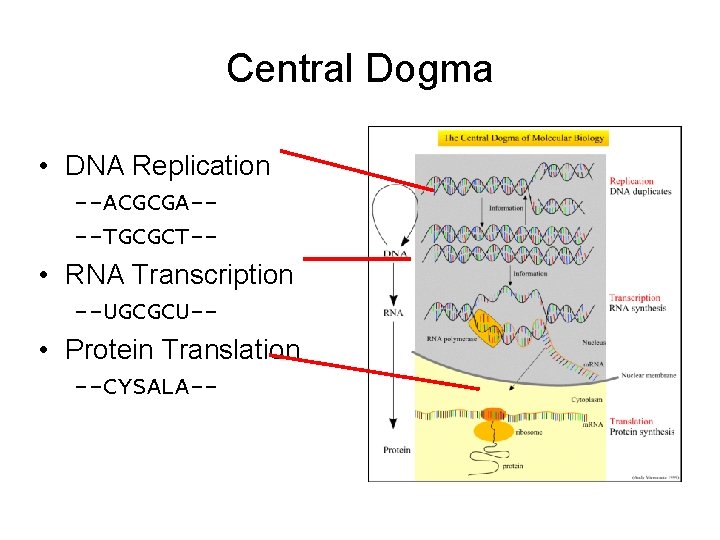
Central Dogma • DNA Replication --ACGCGA---TGCGCT-- • RNA Transcription --UGCGCU-- • Protein Translation --CYSALA--
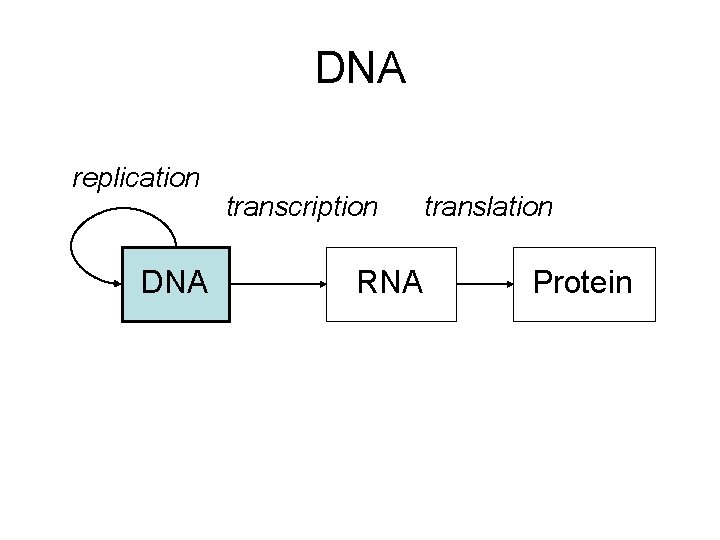
DNA replication DNA transcription RNA translation Protein
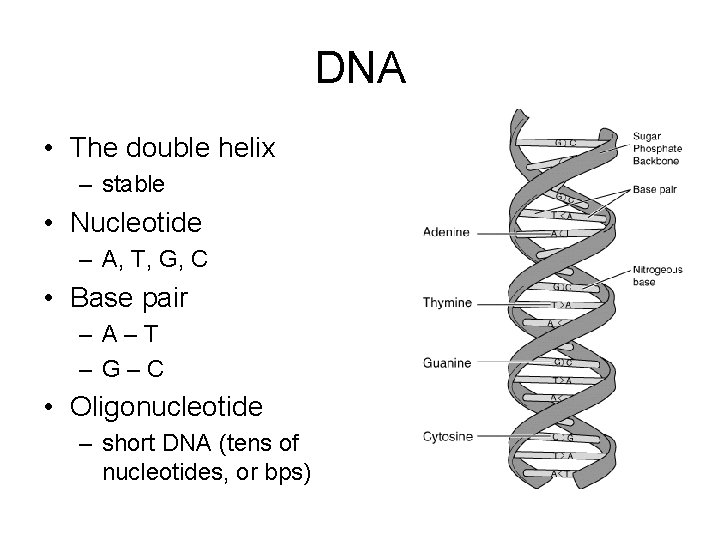
DNA • The double helix – stable • Nucleotide – A, T, G, C • Base pair – A–T – G–C • Oligonucleotide – short DNA (tens of nucleotides, or bps)

DNA Strand • DNA has canonical orientation – read from 5’ to 3’ – antiparallel: one strand has direction opposite to its complement’s 5’ … 3’ … TACTGAA … 3’ ATGACTT … 5’
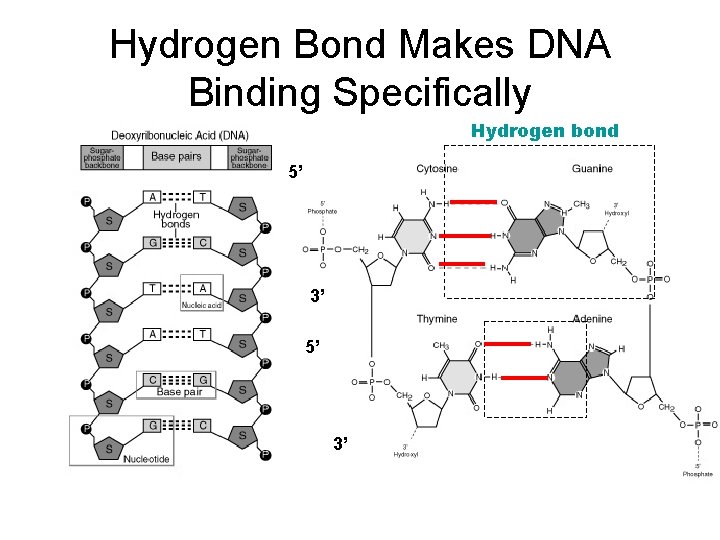
Hydrogen Bond Makes DNA Binding Specifically Hydrogen bond 5’ 3’

Hydrogen Bond Makes DNA Binding Specifically • The force between base pair is hydrogen bond, This force let A-T(U), C-G can specifically match together.
RNA replication DNA transcription RNA translation Protein

RNA • Types – messenger RNA – ribosomal RNA (r. RNA) – transfer RNA (t. RNA) Gene is expressed by transcribing DNA into single-stranded m. RNA
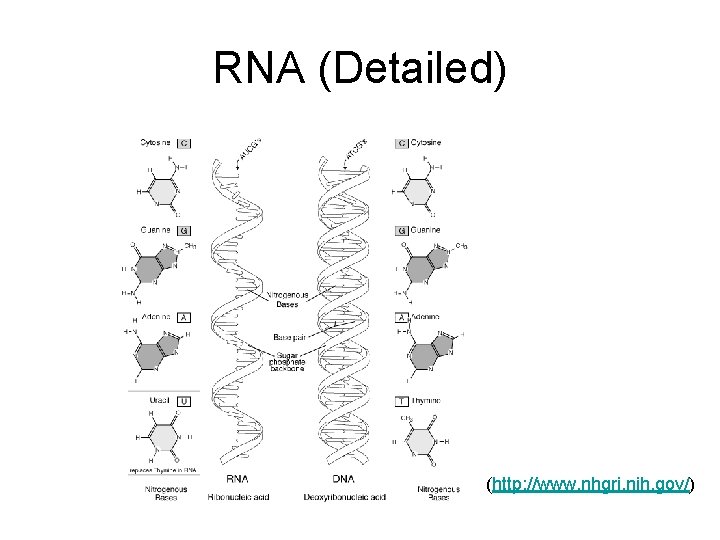
RNA (Detailed) (http: //www. nhgri. nih. gov/)
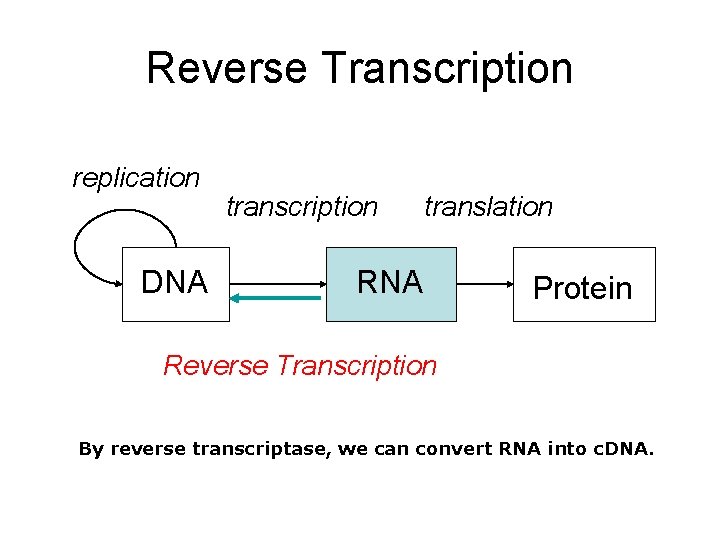
Reverse Transcription replication DNA transcription translation RNA Protein Reverse Transcription By reverse transcriptase, we can convert RNA into c. DNA.

The Southern Blot • Basic DNA detection technique that has been used for over 30 years, known as Southern blots: – – A “known” strand of DNA is deposited on a solid support (i. e. nitocellulose paper) An “unknown” mixed bag of DNA is labelled (radioactive or flourescent) “Unknown” DNA solution allowed to mix with known DNA (attached to nitro paper), then excess solution washed off If a copy of “known” DNA occurs in “unknown” sample, it will stick (hybridize), and labeled DNA will be detected on photographic film

m. RNA Represent Gene Function • When measure the level of a m. RNA, we are monitoring the activity of a gene. • Thus, if we can understand all the level of m. RNAs, we can study the expression of whole genome. • Microarray takes the advantage of getting over 10000 of blotting data in a single experiment, which makes monitoring the genome activity possible.

Content • • • Biology background of microarray Design of microarray The workflow of microarray Image analysis of microarray Data analysis of microarray Discussion

Design of Microarray • Microarray in different context • The idea of microarray • Main type of array chips

m. RNA Levels Compared in Many Different Contexts l Different tissues, same organism (brain v. liver) l Same tissue, same organism (tumor v. nontumor) l Same tissue, different organisms (wt v. mutant) l Time course experiments (development) l Other special designs (e. g. to detect spatial patterns).
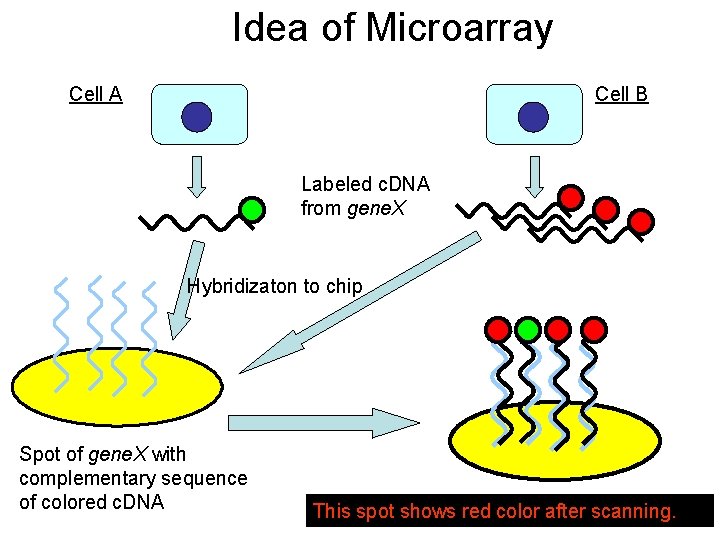
Idea of Microarray Cell A Cell B Labeled c. DNA from gene. X Hybridizaton to chip Spot of gene. X with complementary sequence of colored c. DNA This spot shows red color after scanning.
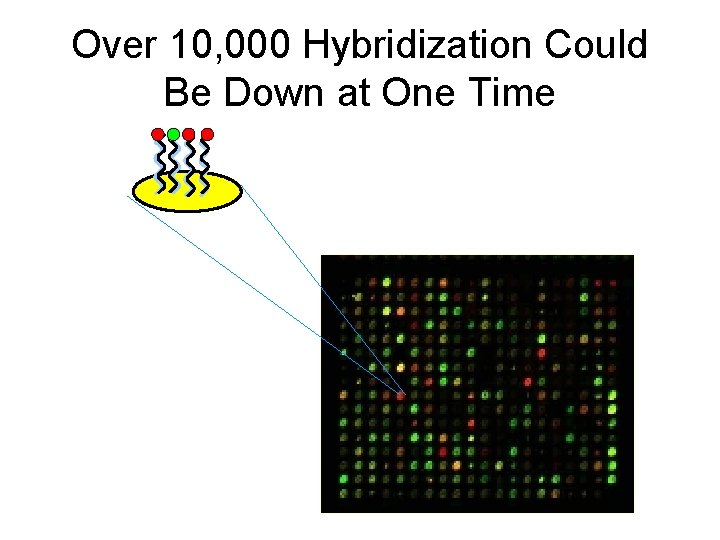
Over 10, 000 Hybridization Could Be Down at One Time

Several Types of Arrays • Spotted DNA arrays – Developed by Pat Brown’s lab at Stanford – PCR products of full-length genes (>100 nt) • Affymetrix gene chips – Photolithography technology from computer industry allows building many 25 -mers • Ink-jet microarrays from Agilent – 25 -60 -mers “printed directly on glass slides – Flexible, rapid, but expensive

Array Fabrication Spotting • Use PCR to amplify DNA • Robotic "pen" deposits DNA at defined coordinates • approximately 1 -10 ng per spot • Experimentation with oligos (40, 70 bp)

This machine can make 48 microarrays simultaneously.

Array Fabrication Photo-lithography • Light activated synthesis • synthesize oligonucleotides on glass slides • 107 copies per oligo in 24 x 24 um square • Use 20 pairs of different 25 -mers per gene • Perfect match and mismatch
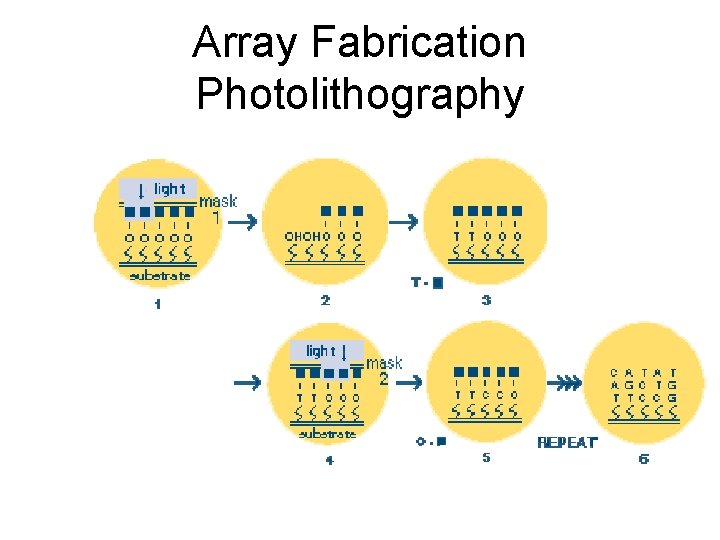
Array Fabrication Photolithography
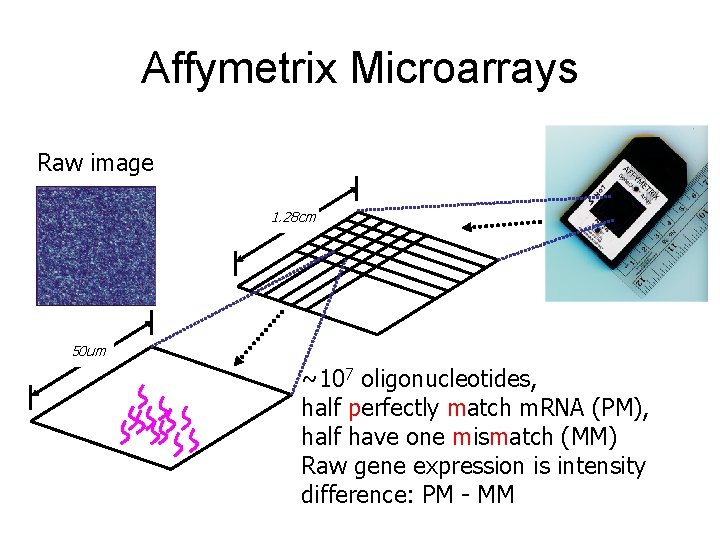
Affymetrix Microarrays Raw image 1. 28 cm 50 um ~107 oligonucleotides, half perfectly match m. RNA (PM), half have one mismatch (MM) Raw gene expression is intensity difference: PM - MM

Agilent c. DNA microarray and oligonucelotides microarray • Agilent delivering printed 60 -mer microarrays in addition to 25 -mer formats. • The inkjet process uses standard phosphoramidite chemistry to deliver extremely small volumes (picoliters) of the chemicals to be spotted.

Content • • • Biology background of microarray Design of microarray The workflow of microarray Image analysis of microarray Data analysis of microarray
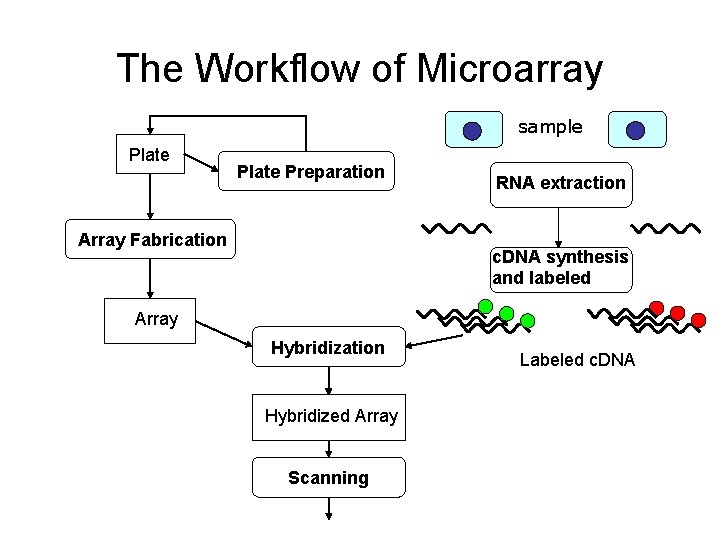
The Workflow of Microarray sample Plate Preparation Array Fabrication RNA extraction c. DNA synthesis and labeled Array Hybridization Hybridized Array Scanning Labeled c. DNA
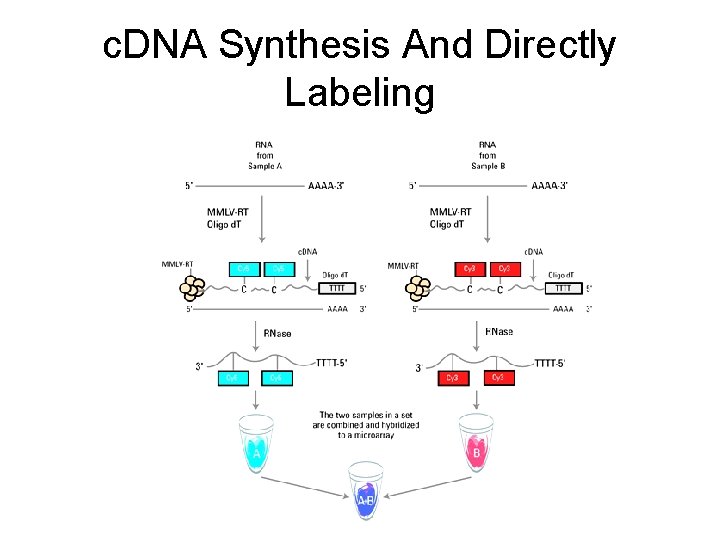
c. DNA Synthesis And Directly Labeling
![Cyanine [Cy 3 and Cy 5] c. DNA Hybridization On To The Chip e. Cyanine [Cy 3 and Cy 5] c. DNA Hybridization On To The Chip e.](http://slidetodoc.com/presentation_image_h2/2a49e23c32b1bbd6239b4d0208e277a7/image-31.jpg)
Cyanine [Cy 3 and Cy 5] c. DNA Hybridization On To The Chip e. g. treatment / control normal / tumor tissue Sample loading 1. Loading from the corner of the cover slip It is time consuming and easily producing bubbles. 1 2 Sample loading 3 Sample loading 2. Loading sample at the center of array then put the slip smoothly Faster, and have lower chance of bubble producing then the last one. 3. Loading sample at the side of the array then put the slip on. Solution would attach to the slip right after the slip contact with it, and would diffuse with the movement of slip when we slowly move down.

Scan Green: down regulate Red: up regulate Yellow: equal level

Content • • • Biology background of microarray Design of microarray The workflow of microarray Image analysis of microarray Data analysis of microarray Discussion

Image analysis • To find a spot • Convert feature into numeric data • Image normalization

The Algorithms 1. Find spots: Finds the location of each spot on the microarray. 2. Cookie cutter algorithm: (1). Suppose the distribution of pixels vs intensity is Gaussian curve (2). Using SD or IQR to identify the feature and background of each spot (3). Calculates statistics for the pixel population
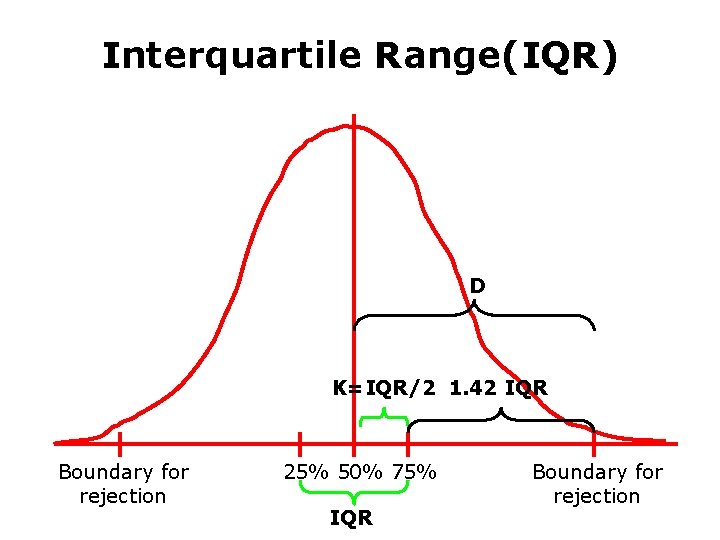
Interquartile Range(IQR) D K=IQR/2 1. 42 IQR Boundary for rejection 25% 50% 75% IQR Boundary for rejection
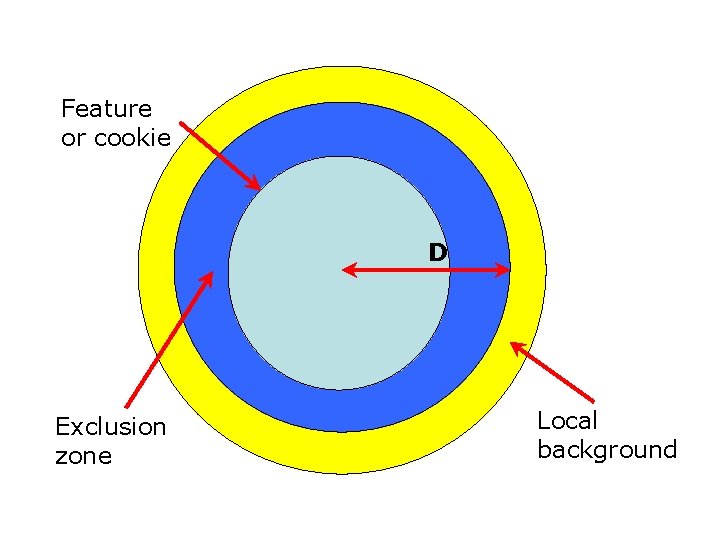
Feature or cookie D Exclusion zone Local background
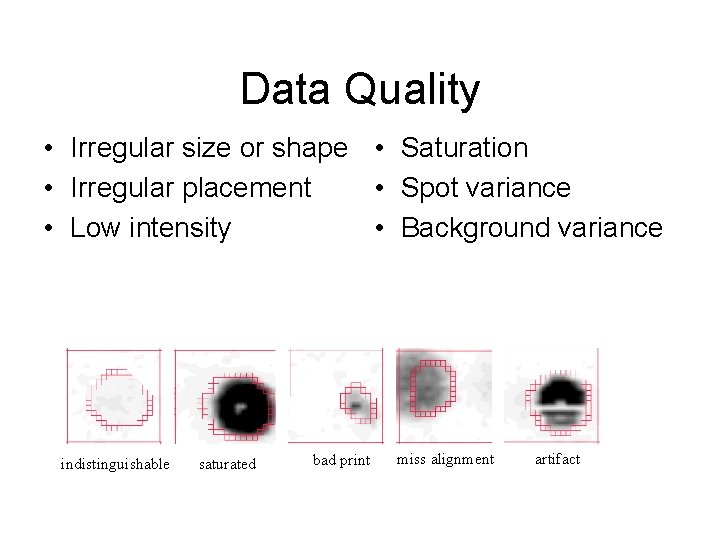
Data Quality • Irregular size or shape • Saturation • Irregular placement • Spot variance • Low intensity • Background variance indistinguishable saturated bad print miss alignment artifact

Convert Feature Into Numeric Value Green b. g. -corrected Red b. g. -corrected background (R. b. g. -c)/(G. b. g. Red intensity Green c) Systematic name intensity Red b. g. Gene function

Data Normalization • Normalize data to correct for variances – Dye bias – Location bias – Intensity bias – Pin bias – Slide bias • Control vs. non-control spots
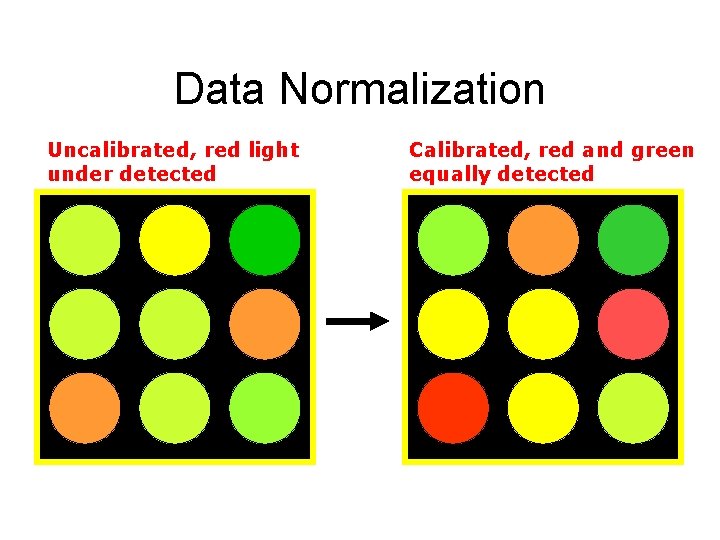
Data Normalization Uncalibrated, red light under detected Calibrated, red and green equally detected

Data Normalization • Assumptions – Overall mean average ratio should be 1 • Most genes are not differentially expressed – Total intensity of dyes are equivalent
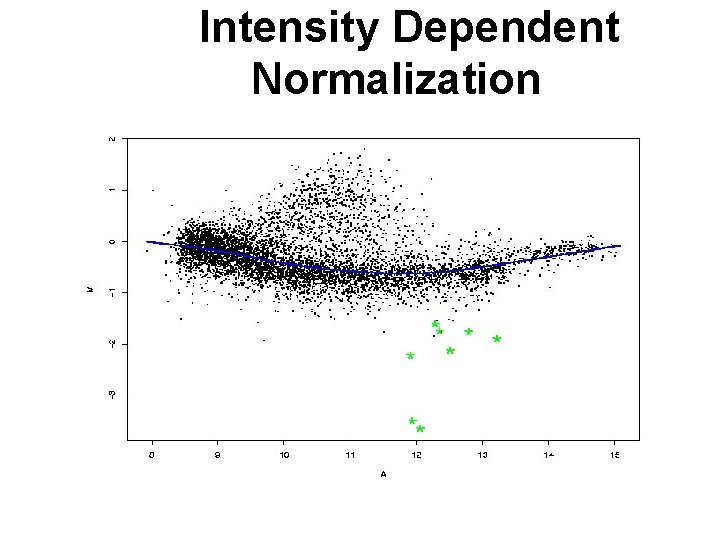
Intensity Dependent Normalization
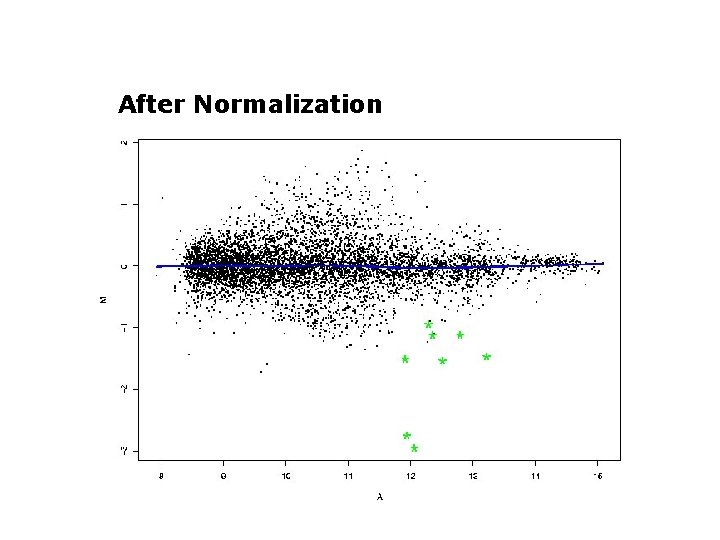
After Normalization

Additional Normalization • Pin dependent – Similar to intensity dependent fit. – Compute individual lowess fits for each pin group • Within slide normalization – After pin dependent normalization, log ratios for each pin are centered around 0 – Scale variance for each pin • Uses MAD (median absolute deviation)

Additional Normalization • Dye swap – Combine relative expression levels without explicit normalization – Compute lowess fit for log 2(RR’/GG’)/2 vs. log 2(A + A’)/2 – Normalized ratio is log 2(R/G) - c(A) where c(A) is the lowess prediction

Content • • • Biology background of microarray Design of microarray The workflow of microarray Image analysis of microarray Data analysis of microarray Discussion

Data analysis • • • Data filtering Fold change analysis Classification Clustering Future direction

Microarray Data Classification Microarray chips Images scanned by laser Value 193 -70 144 33 318 1764 1537 1204 707 Datasets New sample Prediction: Gene D 26528_at D 26561_cds 1_at D 26561_cds 2_at D 26561_cds 3_at D 26579_at D 26598_at D 26599_at D 26600_at D 28114_at Data Mining and analysis

The Threshold of Spots: selection/removing of genes • Filtering - remove genes with insufficient variation – Remove insufficient spot: saturated, None uniform, too high background… – Remove extreme signal: e. g. Max. Val - Min. Val < 500 and Max. Val/Min. Val < 5 – Statistical filtering (e. g. p-value<0. 01) – biological reasons – feature reduction for algorithmic

Microarray Data Analysis Types • Different gene expression – Fold change analysis • Classification (Supervised) – identify disease – predict outcome / select best treatment • Clustering (Unsupervised) – find new biological classes / refine existing ones – exploration • …

Differential Gene Expression • n-fold change – n typically >= 2 – May hold no biological relevance – Often too restrictive • 2 expression – Calculate standard deviation – Genes with expression more than 2 away are differentially expressed
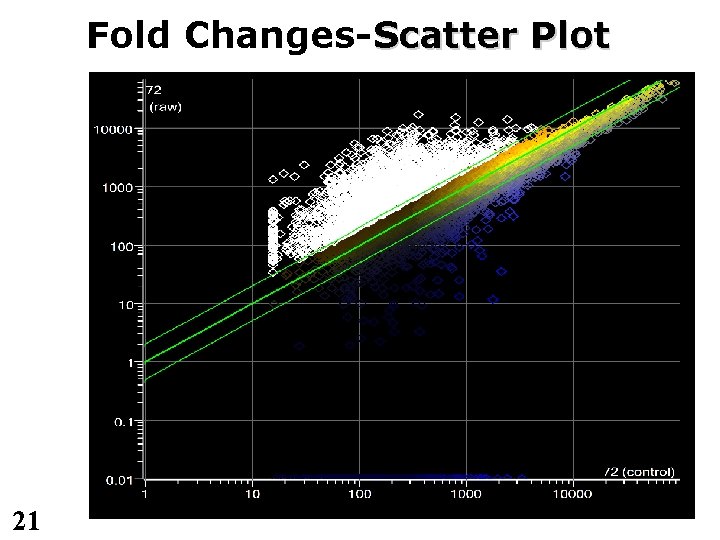
Fold Changes-Scatter Plot 21
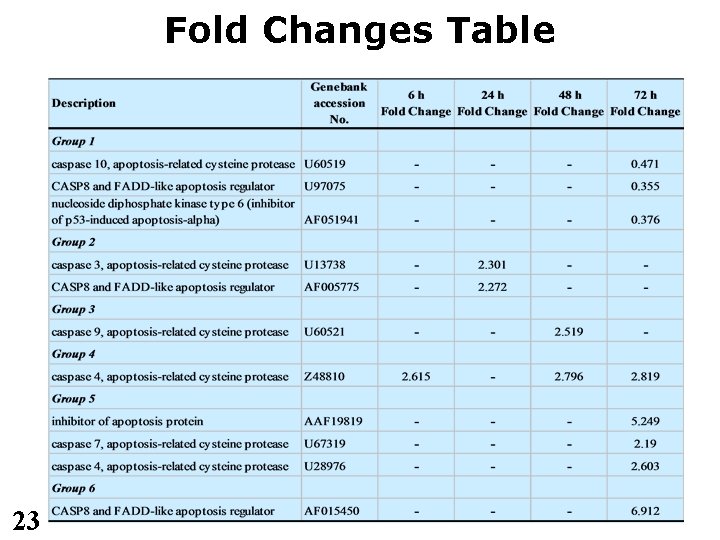
Fold Changes Table 23

Classification / categorization Similar Approach: • select top genes most correlated to each class • select best subset using cross-validation • build a single model separating all classes • Advanced: – build separate model for each class vs. rest – choose model making the strongest prediction

Clustering Goals • Find natural classes in the data • Identify new classes / gene correlations • Refine existing taxonomies • Support biological analysis / discovery • Different Methods – Hierarchical clustering, SOM's, etc

SOM clustering • SOM - self organizing maps • Preprocessing – filter away genes with insufficient biological variation – normalize gene expression (across samples) to mean 0, st. dev 1, for each gene separately. • Run SOM for many iterations • Plot the results
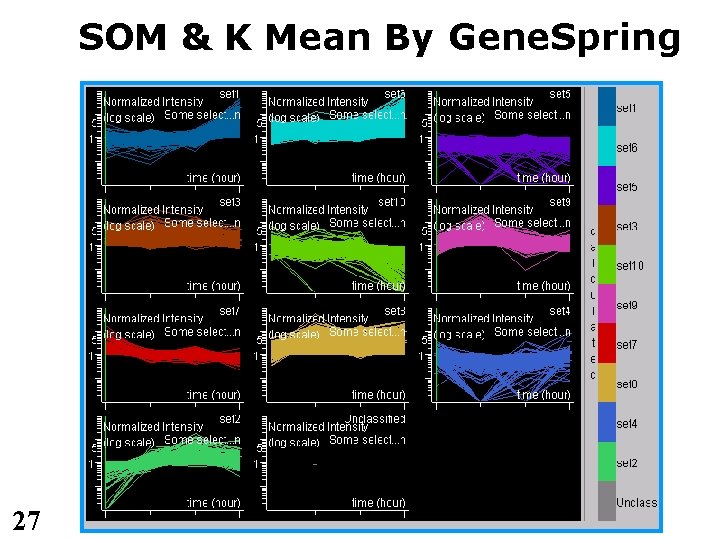
SOM & K Mean By Gene. Spring 27

Hierarchical Clustering • The most popular hierarchical clustering method used in microarray data analysis is the so called agglomerative method – works with the data in a bottom-up manner. • Initially, each data point forms a cluster and the algorithm works through the cluster sets by repeatedly merging the two which are the most similar or have the shortest distance. – algorithm involves the computation of the distance or similarity matrix • O(N^2) complexity and thus is not very efficient.

Hierarchical clustering
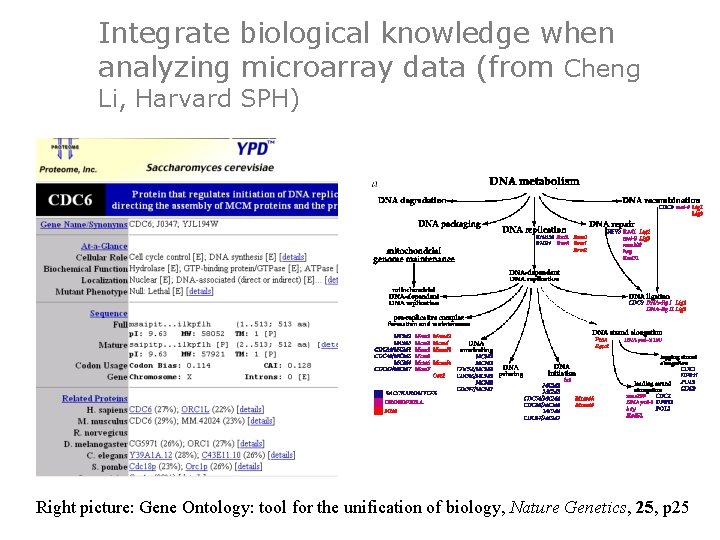
Integrate biological knowledge when analyzing microarray data (from Cheng Li, Harvard SPH) Right picture: Gene Ontology: tool for the unification of biology, Nature Genetics, 25, p 25

Content • • • Biology background of microarray Design of microarray The workflow of microarray Image analysis of microarray Data analysis of microarray Discussion

Microarray Potential Applications • Biological discovery – – new and better molecular diagnostics new molecular targets for therapy finding and refining biological pathways Mutation and polymorphism detection • Recent examples – molecular diagnosis of leukemia, breast cancer, . . . – appropriate treatment for genetic signature – potential new drug targets

Microarray Limitations n n Cross-hybridization of sequences with high identity Chip to chip variation True measure of abundance? Does m. RNA levels reflect protein levels? n Generally, do not “prove” new biology - simply suggest genes involved in a process, a hypothesis that will require traditional experimental verification. n What fold change has biological relevance? n Need cloned EST or some sequence knowledge -- rare messages may be undetected n Expensive!! Not every lab can afford experiment repeat. n The real limitation is Bioinformatics
- Slides: 64