Microarray An Introduction Gene Background Gene a region
Microarray: An Introduction

Gene Background • Gene: a region of the human genome coding for a protein • Biologists interested in gene function and disease • Studying multiple genes is time consuming and expensive

§ Genes are differentially expressed in. . . • Different cell types (e. g. muscle cells, fibroblasts) • Environmental conditions (e. g. heat shock, nutrient deprivation) • Developmental phases • Cell-cycle stages (e. g. G 1 phase) • Disease states (e. g. tumor cells, virus-infected cells)

• Gene expression is primarily regulated at the level of transcription § Hence, the number of m. RNA copies in a cell for a particular gene is a good indicator of that gene’s expression (number of proteins) § Dynamic range of m. RNA levels: • Highly expressed genes can have up to 9400 m. RNA copies per cell • Poorly expressed genes can have < 0. 3 m. RNA copies per cell
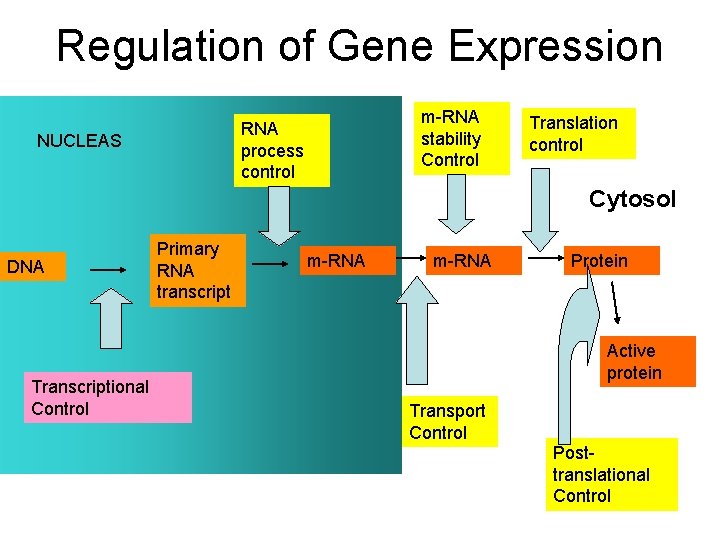
Regulation of Gene Expression m-RNA stability Control RNA process control NUCLEAS Translation control Cytosol DNA Transcriptional Control Primary RNA transcript m-RNA Protein Active protein Transport Control Posttranslational Control

Traditional Methods • Northern Blotting – Single RNA isolated – Probed with labeled c. DNA • Reverse Transcription – Polymerase Chain Reaction (RT-PCR) • Real time or quantitative RT-PCR – Each technique measures the m-RNA levels of one or a few genes at a time

Microarray Technology • Microarray: – New Technology (first paper: 1995) • Allows study of thousands of genes at same time • Tools used to measure simultaneously the transcription level of every gene in a cell

Advantages of DNA microarray experiments • Lack information a priori of which genes may have altered m. RNA levels in a particular experimental condition • Knowledge of patterns of genes with altered m. RNA levels is usually more informative than knowledge of single genes with altered m. RNA levels • Use statistical analysis of microarray data to test hypotheses or generate new hypotheses
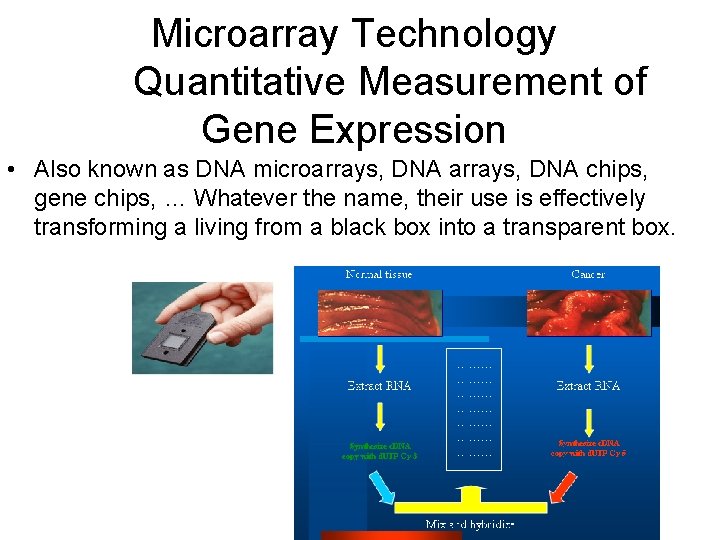
Microarray Technology Quantitative Measurement of Gene Expression • Also known as DNA microarrays, DNA chips, gene chips, … Whatever the name, their use is effectively transforming a living from a black box into a transparent box.
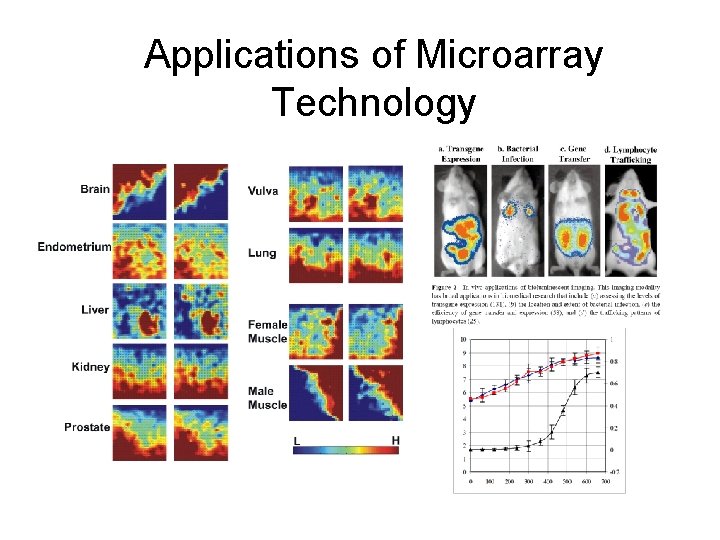
Applications of Microarray Technology

Applications of Microarray Technology • What type of data analysis is required to: – Identify Genes expressed in different cell types (e. g. Liver vs finger) – Learn how expression levels change in different developmental stages (embryo vs. adult) – Learn how expression levels change in different developmental stages (cancerous vs non-cancerous) – Learn how groups of genes inter-relate (gene-gene interactions) – Identify cellular processes that genes participate in (structure, repair, metabolism, replication, … etc) • Applications covered only as example contexts, emphasis is on analysis methods
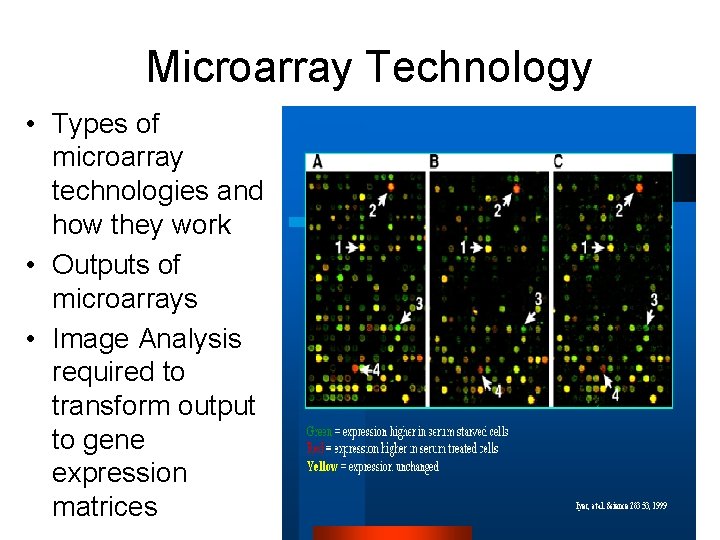
Microarray Technology • Types of microarray technologies and how they work • Outputs of microarrays • Image Analysis required to transform output to gene expression matrices

Fabrications of Microarrays • Size of a microscope slide Images: http: //www. affymetrix. com/
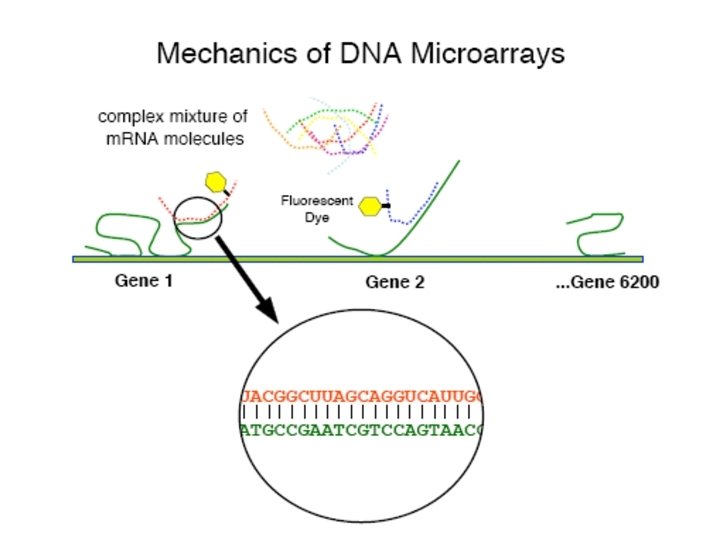

The Mechanics of a DNA Microarray Experiment • Isolate m. RNA from cell cultures • Reverse Transcribe m. RNA into c. DNA • Label c. DNA or c. RNA by incorporating fluorescently-labeled nucleotides • Hybridize labeled c. DNA (or c. RNA) to DNA microarray • Wash and scan microarray in confocal laser scanner • Analyze data

Differing Conditions • Ultimate Goal: – Understand expression level of genes under different conditions • Helps to: – Determine genes involved in a disease – Pathways to a disease – Used as a screening tool

Gene Conditions • • • Cell types (brain vs. liver) Developmental (fetal vs. adult) Response to stimulus Gene activity (wild vs. mutant) Disease states (healthy vs. diseased)

Expressed Genes • Genes under a given condition – m. RNA extracted from cells – m. RNA labeled – Labeled m. RNA is m. RNA present in a given condition – Labeled m. RNA will hybridize (base pair) with corresponding sequence on slide

Constructing DNA Microarrays • A DNA microarray is a collection of DNA probes separated in a regular array atop a solid support (glass slide, silicon chip, etc) • Affymetrix oligonucleotide microarrays Short (25 mers) oligonucleotide probes synthesized on the array • c. DNA microarrays c. DNA PCR products printed on coated glass slides • Oligonucleotide microarrays Long (70 - 100 mers) oligonucleotide probes are printed on coated glass slides

Custom Arrays • Mostly c. DNA arrays • 2 -dye (2 -channel) – RNA from two sources (c. DNA created) • Source 1: labeled with red dye (Cy 5) • Source 2: labeled with green dye (Cy 3)

Two Channel Microarrays • Microarrays measure gene expression • Two different samples: – Control (green label) – Sample (red label) • Both are washed over the microarray – Hybridization occurs – Each spot is one of 4 colors

Building the chip Ngai Lab arrayer , UC Berkeley Print-tip head
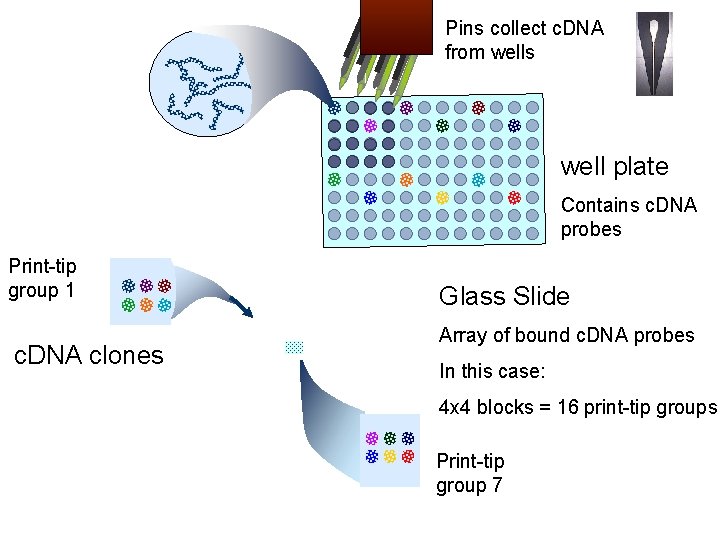
Pins collect c. DNA from wells well plate Contains c. DNA probes Print-tip group 1 c. DNA clones Glass Slide Array of bound c. DNA probes In this case: 4 x 4 blocks = 16 print-tip groups Print-tip group 7
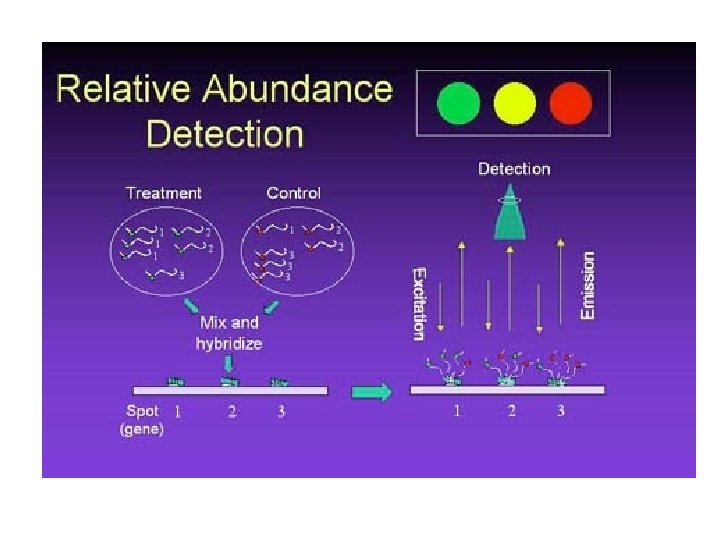

Microarray Image Analysis • Microarrays detect gene interactions: 4 colors: – – Green: high control Red: High sample Yellow: Equal Black: None • Problem is to quantify image signals

A two-channel microarray experiment


Microarray Animations • Davidson University: • http: //www. bio. davidson. edu/courses/genomics/chip. html • Imagecyte: • http: //www. imagecyte. com/array 2. html
- Slides: 28