mi RNAs and biomarkers Gabriella Sozzi q diagnostic

mi. RNAs and biomarkers Gabriella Sozzi

q diagnostic micro. RNAs in lung tumors ü stratifying lung cancer molecular subtypes ( Landi L. et al) q prognostic micro. RNAS in tumors ü mi. RNA expression profiles to predict clinical outcomes of resectable SCLC patients (Nan Bi et al) ü Micro. RNAs associated with survival in malignant pleural mesothelioma patients (Kirschner et al) q diagnostic micro. RNAs in biological fluids ü sputum mi. RNA expression profiles for the detection of non-small cell lung cancer (Razzak et al) ü plasma mi. RNA test for lung cancer screening (Sozzi et al) q Biomarker-Driven Programs for Lung Cancer Screening General considerations (Massion P. )
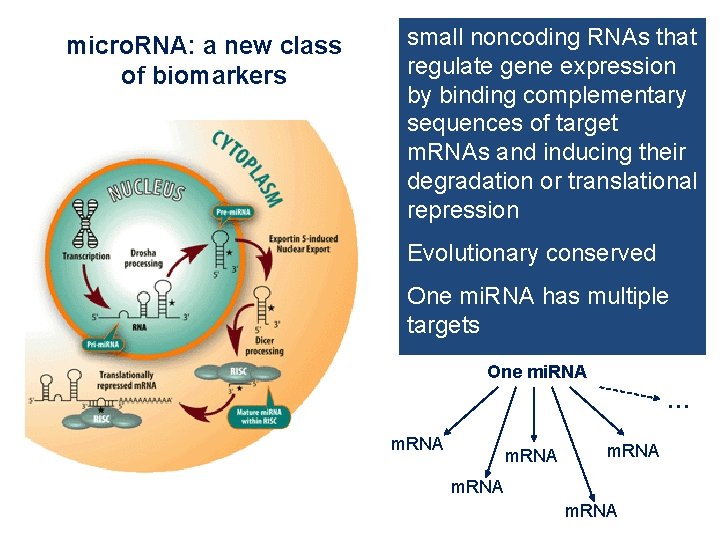
micro. RNA: a new class of biomarkers small noncoding RNAs that regulate gene expression by binding complementary sequences of target m. RNAs and inducing their degradation or translational repression Evolutionary conserved One mi. RNA has multiple targets One mi. RNA … m. RNA
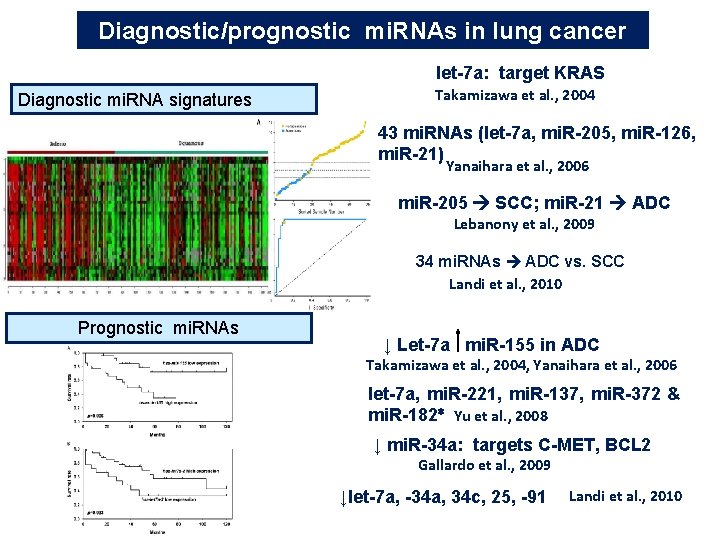
Diagnostic/prognostic mi. RNAs in lung cancer let-7 a: target KRAS Diagnostic mi. RNA signatures Takamizawa et al. , 2004 43 mi. RNAs (let-7 a, mi. R-205, mi. R-126, mi. R-21) Yanaihara et al. , 2006 mi. R-205 SCC; mi. R-21 ADC Lebanony et al. , 2009 34 mi. RNAs ADC vs. SCC Landi et al. , 2010 Prognostic mi. RNAs ↓ Let-7 a mi. R-155 in ADC Takamizawa et al. , 2004, Yanaihara et al. , 2006 let-7 a, mi. R-221, mi. R-137, mi. R-372 & mi. R-182∗ Yu et al. , 2008 ↓ mi. R-34 a: targets C-MET, BCL 2 Gallardo et al. , 2009 ↓let-7 a, -34 a, 34 c, 25, -91 Landi et al. , 2010
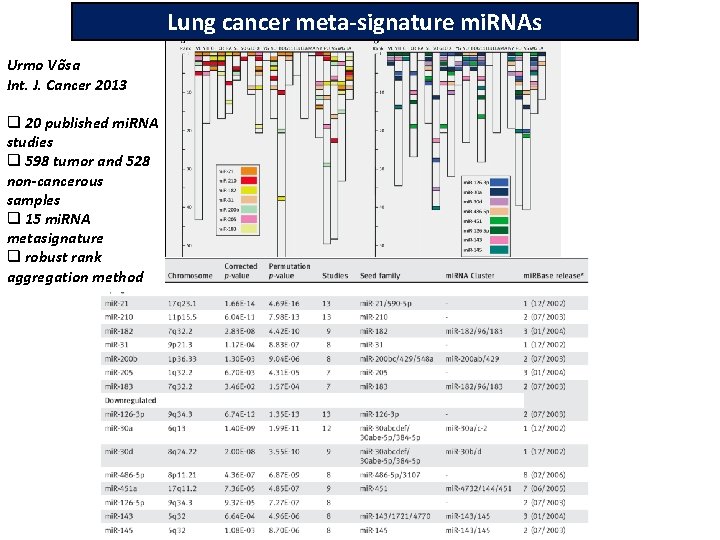
Lung cancer meta-signature mi. RNAs Vo˜sa Urmo Võsa Int. J. Cancer 2013 q 20 published mi. RNA studies q 598 tumor and 528 non-cancerous samples q 15 mi. RNA metasignature q robust rank aggregation method
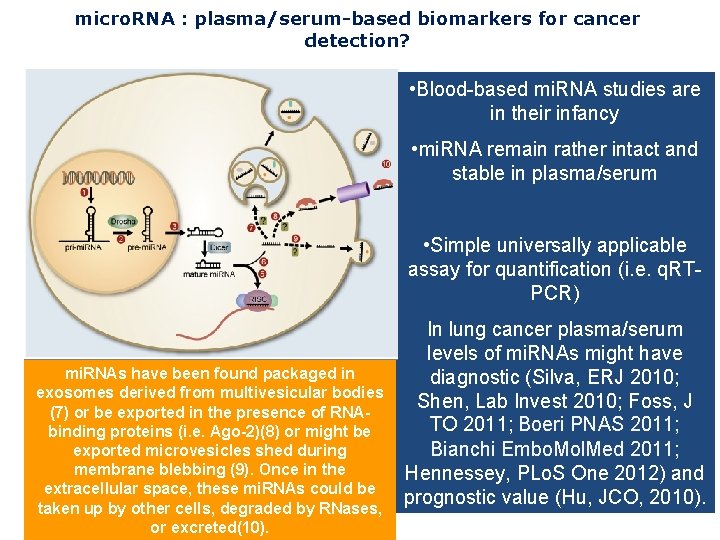
micro. RNA : plasma/serum-based biomarkers for cancer detection? • Blood-based mi. RNA studies are in their infancy • mi. RNA remain rather intact and stable in plasma/serum mi. RNAs have been found packaged in exosomes derived from multivesicular bodies (7) or be exported in the presence of RNAbinding proteins (i. e. Ago-2)(8) or might be exported microvesicles shed during membrane blebbing (9). Once in the extracellular space, these mi. RNAs could be taken up by other cells, degraded by RNases, or excreted(10). • Simple universally applicable assay for quantification (i. e. q. RTPCR) In lung cancer plasma/serum levels of mi. RNAs might have diagnostic (Silva, ERJ 2010; Shen, Lab Invest 2010; Foss, J TO 2011; Boeri PNAS 2011; Bianchi Embo. Mol. Med 2011; Hennessey, PLo. S One 2012) and prognostic value (Hu, JCO, 2010).

MO 16. 01: Different Micro-RNA Expression In Lung Adenocarcinoma With Molecular Driver Events Lorenza Landi 1 Pierluigi Gasparini 2, Stefania Carasi 2, Carmelo Tibaldi 1 , Luciano Cascione 2, Greta Alì 3, Armida D’Incecco 1, Jessica Salvini 1, Gabriele Minuti 1 , Antonio Chella 3 , Gabriella Fontanini 3, Federico Cappuzzo 1 and Carlo M. Croce 2 1 Istituto Toscano Tumori, Dipartimento di Oncologia, Livorno Italy 2 The Ohio State University, Comprehensive Cancer Center, Department of Molecular Virology, Immunology and Medical Genetics, Columbus, OH, USA 3 Azienda Ospedaliera Universitaria Pisana, Pisa, Italy

Trial Background: • Oncogenic driving mutations identify lung adenocarcinoma with different prognosis and sensitivity to targeted therapy • Recent studies have suggested that mi. RNAs could be useful for stratifying lung cancer subtypes, however mi. RNAs deregulation in NSCLC with ALK translocation, EGFR or KRAS mutations is largely unknown Aim: • Identify mi. RNA signature differences according to the presence of specific oncogenic driver Methods: • Retrospective analysis of a cohort of 67 NSCLCs matched with 17 normal lung tissues • RNA was isolated from FFPE using the Recover ALL kit (Ambion) and mi. RNA levels were analyzed using the Nano. String mi. RNA V 2 panel • Data were processed according to manufacture guidelines. We used Limma to test for differential expression analysis data • The mi. RNAs expression between tissues for all RT-q. PCR was analyzed using the parametric t-test (unpaired, 2 -tailed for validation) MO 16. 01: Different Micro-RNA Expression In Lung Adenocarcinoma With Molecular Driver Events - Landi L
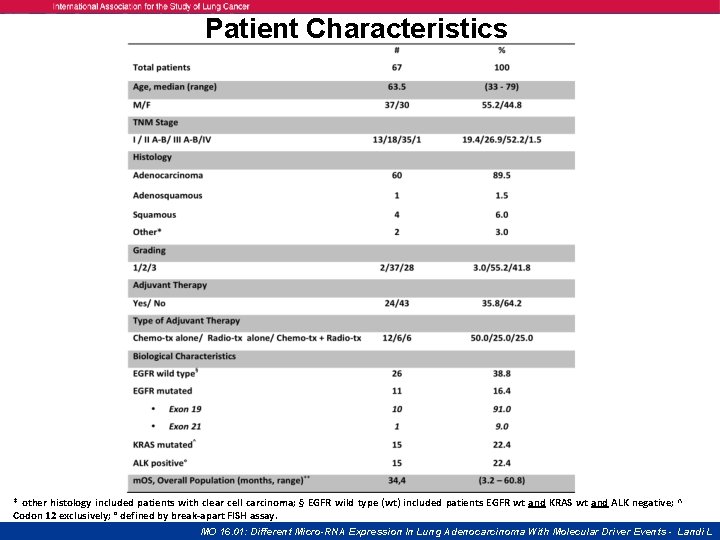
Patient Characteristics * other histology included patients with clear cell carcinoma; § EGFR wild type (wt) included patients EGFR wt and KRAS wt and ALK negative; ^ Codon 12 exclusively; ° defined by break-apart FISH assay. MO 16. 01: Different Micro-RNA Expression In Lung Adenocarcinoma With Molecular Driver Events - Landi L
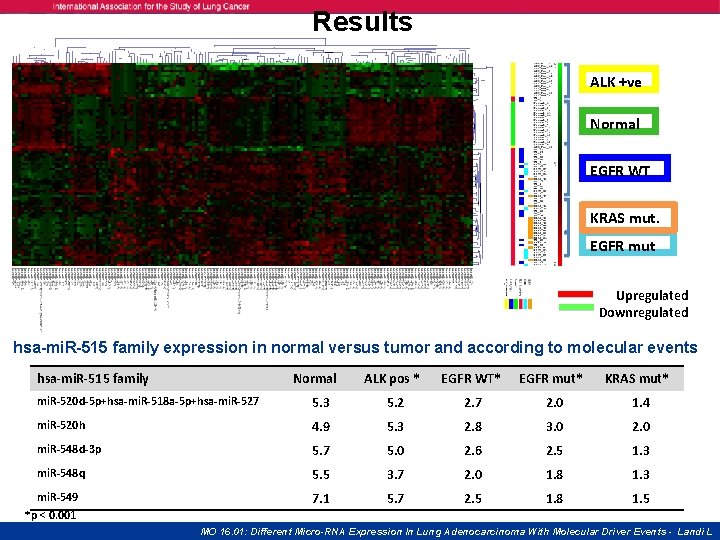
Results ALK +ve Normal EGFR WT KRAS mut. EGFR mut Upregulated Downregulated hsa-mi. R-515 family expression in normal versus tumor and according to molecular events hsa-mi. R-515 family Normal ALK pos * EGFR WT* EGFR mut* KRAS mut* mi. R-520 d-5 p+hsa-mi. R-518 a-5 p+hsa-mi. R-527 5. 3 5. 2 2. 7 2. 0 1. 4 mi. R-520 h 4. 9 5. 3 2. 8 3. 0 2. 0 mi. R-548 d-3 p 5. 7 5. 0 2. 6 2. 5 1. 3 mi. R-548 q 5. 5 3. 7 2. 0 1. 8 1. 3 7. 1 5. 7 2. 5 1. 8 1. 5 mi. R-549 *p < 0. 001 MO 16. 01: Different Micro-RNA Expression In Lung Adenocarcinoma With Molecular Driver Events - Landi L

Conclusions • mi. RNAs profile significantly differs in lung cancer patients with ALK translocation, EGFR mutations and KRAS mutations • Prognostic and predictive role of several mi. RNAs are currently under investigation • mi. RNAs expression could represent an useful tool to refine diagnosis of oncogene addicted NSCLC • Targeting mi. RNAs could represent a potential strategy to modulate sensitivity to biological agents MO 16. 01: Different Micro-RNA Expression In Lung Adenocarcinoma With Molecular Driver Events - Landi L

Micro. RNA Signature Predicts Survival in Resectable Small Cell Lung Cancer Nan Bi, Jianzhong Cao, Yongmei Song, Jie Shen, Wenyang Liu, Jing Fan, Guogui Sun, Tong, Jie He, Yuankai Shi, Xun Zhang, Ning Lu, Qimin Zhan, and Luhua Wang Cancer Hospital and Cancer Institute, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing 100021, China
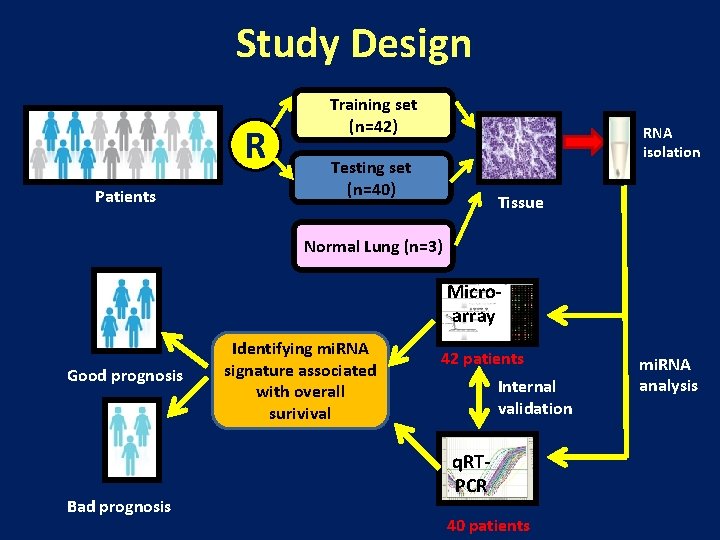
Study Design R Patients Training set (n=42) RNA isolation Testing set (n=40) Tissue Normal Lung (n=3) Microarray Good prognosis Bad prognosis Identifying mi. RNA signature associated with overall surivival 42 patients Internal validation q. RTPCR 40 patients mi. RNA analysis
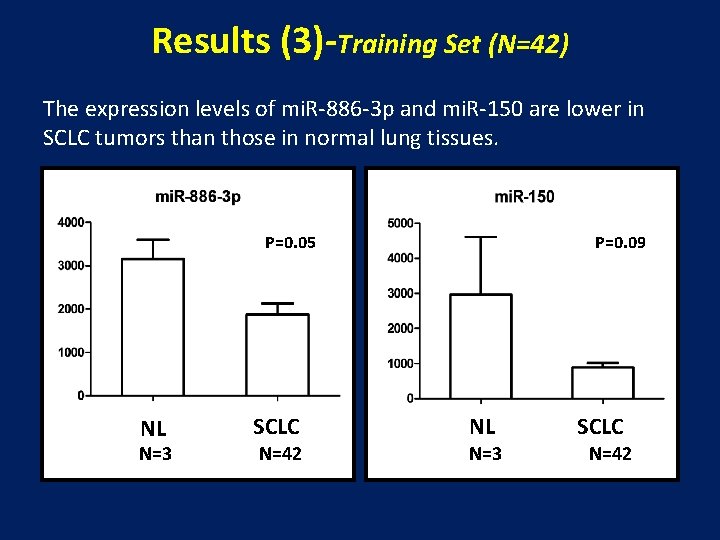
Results (3)-Training Set (N=42) The expression levels of mi. R-886 -3 p and mi. R-150 are lower in SCLC tumors than those in normal lung tissues. P=0. 05 NL N=3 P=0. 09 SCLC NL N=42 N=3 SCLC N=42
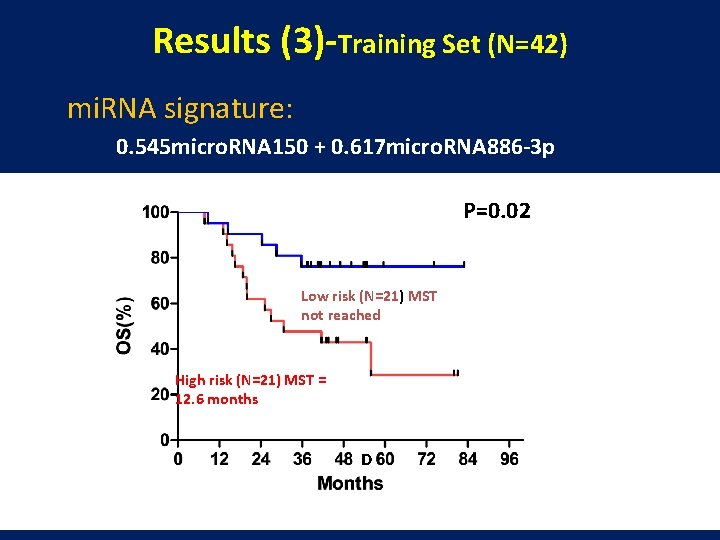
Results (3)-Training Set (N=42) mi. RNA signature: 0. 545 micro. RNA 150 + 0. 617 micro. RNA 886 -3 p P=0. 02 Low risk (N=21) MST not reached High risk (N=21) MST = 12. 6 months D
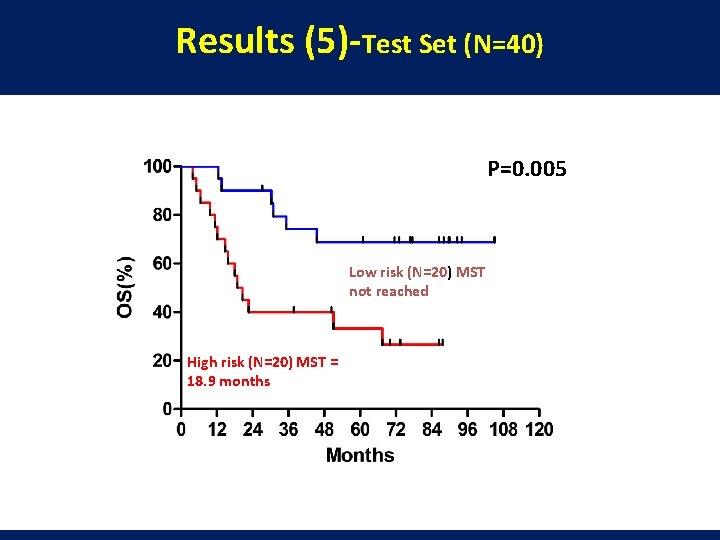
Results (5)-Test Set (N=40) P=0. 005 Low risk (N=20) MST not reached High risk (N=20) MST = 18. 9 months
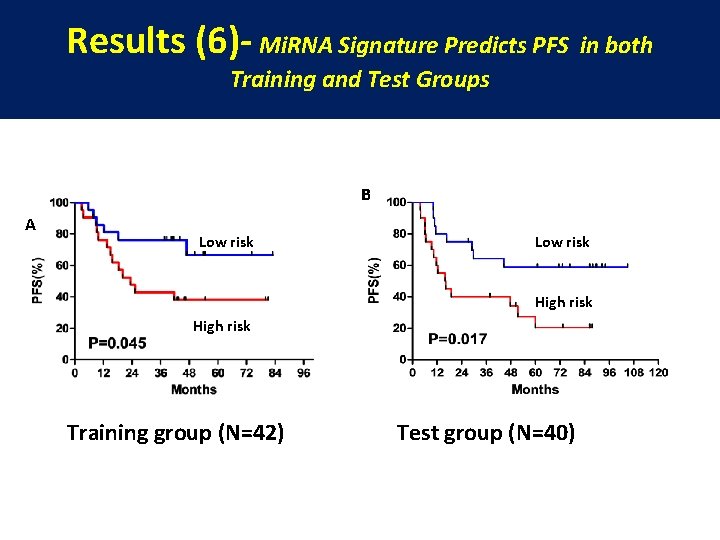
Results (6)- Mi. RNA Signature Predicts PFS in both Training and Test Groups B A Low risk High risk Training group (N=42) Test group (N=40)
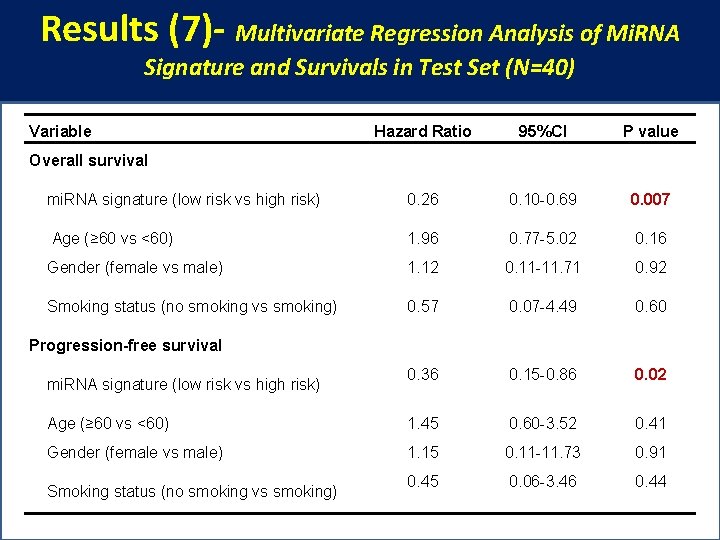
Results (7)- Multivariate Regression Analysis of Mi. RNA Signature and Survivals in Test Set (N=40) Variable Hazard Ratio 95%CI P value mi. RNA signature (low risk vs high risk) 0. 26 0. 10 -0. 69 0. 007 Age (≥ 60 vs <60) 1. 96 0. 77 -5. 02 0. 16 Gender (female vs male) 1. 12 0. 11 -11. 71 0. 92 Smoking status (no smoking vs smoking) 0. 57 0. 07 -4. 49 0. 60 0. 36 0. 15 -0. 86 0. 02 Age (≥ 60 vs <60) 1. 45 0. 60 -3. 52 0. 41 Gender (female vs male) 1. 15 0. 11 -11. 73 0. 91 0. 45 0. 06 -3. 46 0. 44 Overall survival Progression-free survival mi. RNA signature (low risk vs high risk) Smoking status (no smoking vs smoking)

Conclusion l A mi. R-150/mi. R-886 -3 p signature was correlated with the survivals in 42 resectable SCLCs and validated independently with another 40 SCLC cases. l The expression levels of both mi. R-150 and mi. R-886 -3 p were lower in tumors than in normal lung tissues, indicating both of them could serve as tumor suppressor genes in SCLC. l Micro. RNAs may serve as promising prognostic markers as well as noval therapeutic targets for SCLC. l Larger sample size and function studies are warranted to validate our findings.

Micro. RNAs mi. R-17 -5 p, mi. R-21 and mi. R-210 are associated with survival in malignant pleural mesothelioma patients undergoing extrapleural pneumonectomy Michaela B Kirschner 1, Yuen Yee Cheng 1, Steven C Kao 1, 2, Brian C Mc. Caughan 3, 4, Nico van Zandwijk 1, Glen Reid 1 1 Asbestos Diseases Research Institute, University of Sydney of Medical Oncology, Sydney Cancer Centre 3 Cardiothoracic Surgical Unit, Royal Prince Alfred Hospital Sydney 4 The Baird Institute and Sydney Medical School, University of Sydney 2 Department
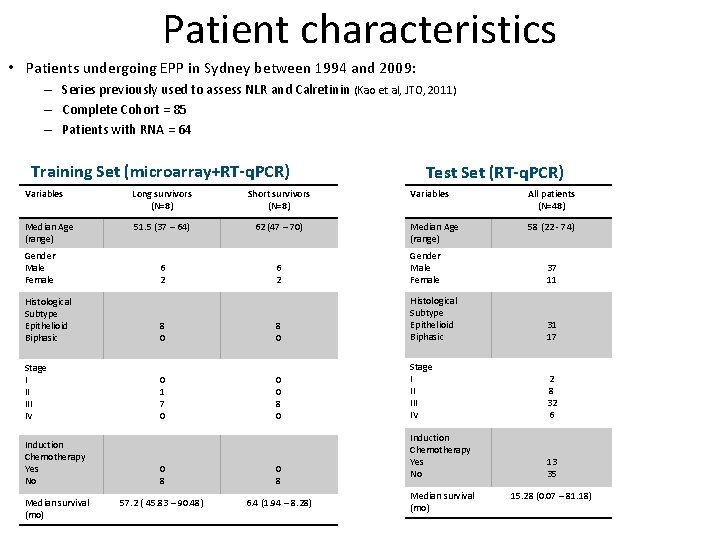
Patient characteristics • Patients undergoing EPP in Sydney between 1994 and 2009: – Series previously used to assess NLR and Calretinin (Kao et al, JTO, 2011) – Complete Cohort = 85 – Patients with RNA = 64 Training Set (microarray+RT-q. PCR) Test Set (RT-q. PCR) Variables All patients (N=48) Median Age (range) 58 (22 - 74) Variables Long survivors (N=8) Short survivors (N=8) Median Age (range) 51. 5 (37 – 64) 62 (47 – 70) 6 2 Gender Male Female 37 11 31 17 Gender Male Female Histological Subtype Epithelioid Biphasic 8 0 Histological Subtype Epithelioid Biphasic Stage I II IV 0 1 7 0 0 0 8 0 Stage I II IV 2 8 32 6 Induction Chemotherapy Yes No 13 35 Induction Chemotherapy Yes No Median survival (mo) 0 8 57. 2 ( 45. 83 – 90. 48) 6. 4 (1. 94 – 8. 28) Median survival (mo) 15. 28 (0. 07 – 81. 18)
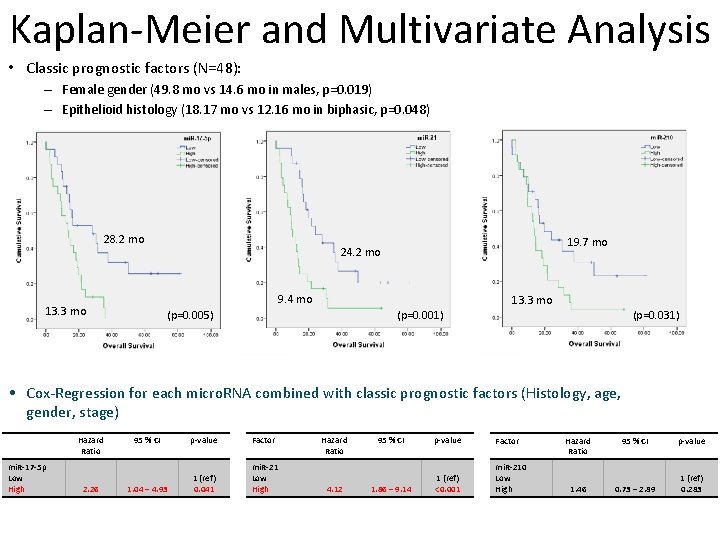
Kaplan-Meier and Multivariate Analysis • Classic prognostic factors (N=48): – Female gender (49. 8 mo vs 14. 6 mo in males, p=0. 019) – Epithelioid histology (18. 17 mo vs 12. 16 mo in biphasic, p=0. 048) 28. 2 mo 19. 7 mo 24. 2 mo 9. 4 mo 13. 3 mo (p=0. 001) (p=0. 005) (p=0. 031) • Cox-Regression for each micro. RNA combined with classic prognostic factors (Histology, age, gender, stage) Hazard Ratio mi. R-17 -5 p Low High 2. 26 95 % CI p-value Factor 1. 04 – 4. 93 1 (ref) 0. 041 mi. R-21 Low High Hazard Ratio 4. 12 95 % CI p-value Factor 1. 86 – 9. 14 1 (ref) <0. 001 mi. R-210 Low High Hazard Ratio 95 % CI p-value 1. 46 0. 73 – 2. 89 1 (ref) 0. 283

Conclusions and Future Directions • Lower expression levels of three micro. RNAs in tumour tissue are associated with longer survival of patients undergoing EPP • mi. R-17 -5 p and mi. R-21 remain significant in a multivariate model including classic prognostic factors Those micro. RNAs have the potential to assist in better selection of patients considered for EPP • Validation in independent samples sets is required • Combination of several micro. RNAs as potential prognostic signature

mi. RNAs in biological fluids • P 2. 20 -011 | A prospective clinical study evaluating stage dependent sputum micro-RNA expression profiles for the detection of nonsmall cell lung cancer • Authors: Rene Razzak 1, Eric L. R. Bédard 1, Julian O. Kim 2, Sayf Gazala 1, Linghong Guo 2, Sunita Ghosh 2, Anil A. Joy 2, Tirath Nijjar 2, Eric Wong 1, Wilson H. Roa 2 1 University of Alberta, Edmonton, AB/CANADA, 2 Cross Cancer Institute, Edmonton, AB/CANADA
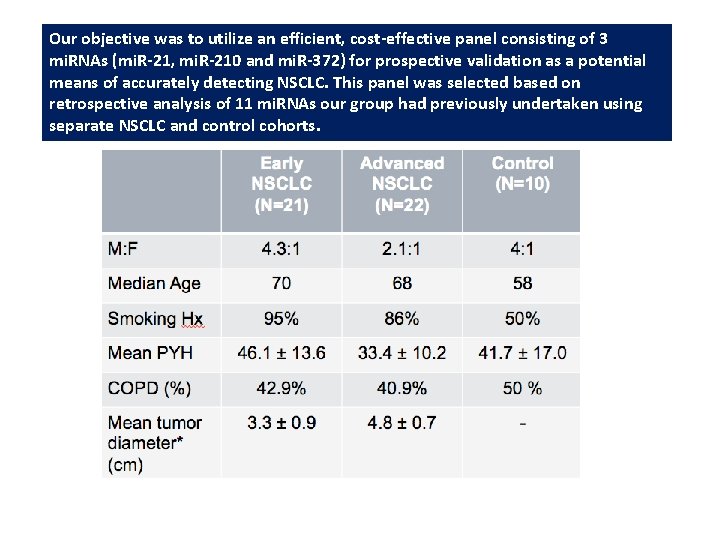
Our objective was to utilize an efficient, cost-effective panel consisting of 3 mi. RNAs (mi. R-21, mi. R-210 and mi. R-372) for prospective validation as a potential means of accurately detecting NSCLC. This panel was selected based on retrospective analysis of 11 mi. RNAs our group had previously undertaken using separate NSCLC and control cohorts.

• 21 early NSCLC (≤ Stage II) patients, 22 advanced NSCLC (≥ Stage III) patients and 10 control subjects were prospectively accrued. A single sputum sample was obtained through spontaneous expectoration from each study participant. • mi. R-21, mi. R-210 and mi. R-372 expression was conducted on each sputum sample and normalized to an endongenous control (U 6) relative to a MRC-5 reference sample, using RNA reverse transcription and Quantitative real-time Polymerase Chain Reaction (RT-q. PCR). • Statistical evaluation consisted of unsupervised hierarchical cluster analysis of the experimentalnormalized mi. RNA expression profiles using within-group linkage.

Comparing early NSCLC to controls, the use of mi. R-21, mi. R-210 and mi. R 372 expression yielded a diagnostic sensitivity of 66. 7% and a specificity of 90. 0%. Advanced NSCLC patients had an improved sensitivity of 81. 8% with the same specificity of 90. 0%. The utilization of mi. R-21, mi. R-210 and mi. R 372 sputum expression might provide a sensitive and specific means of detecting NSCLC. The potential linkage between their expression and NSCLC stage may account for the higher sensitivity observed in the advanced NSCLC group. Future use of this promising panel on a larger population will be required to establish its potential application as a screening tool.

Plasma mi. RNA test for lung cancer screening Gabriella Sozzi

2005 - 2011 4, 000 Smokers ≥ R 50 years Smoking cessation Lung function assessment blood sampling > 100, 000 biological samples + LDCT R LDCT every year LDCT every 2 years Pastorino U. et al. , Eur J Cancer Prev. 2012
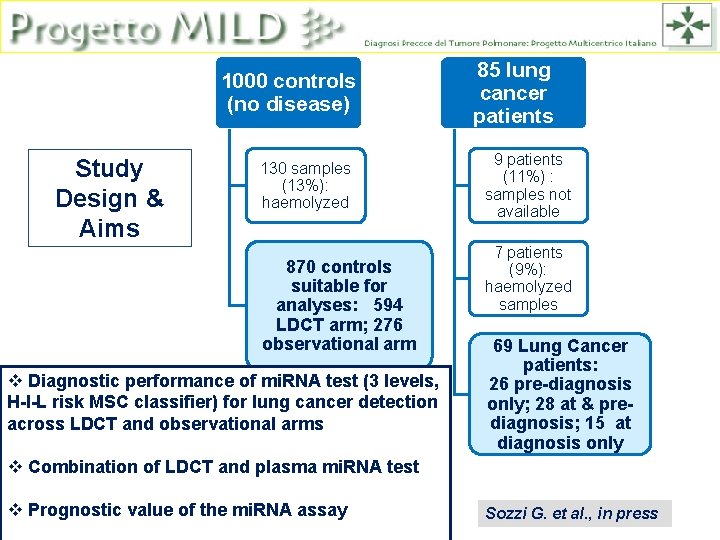
1000 controls (no disease) Study Design & Aims 130 samples (13%): haemolyzed 870 controls suitable for analyses: 594 LDCT arm; 276 observational arm v Diagnostic performance of mi. RNA test (3 levels, H-I-L risk MSC classifier) for lung cancer detection across LDCT and observational arms 85 lung cancer patients 9 patients (11%) : samples not available 7 patients (9%): haemolyzed samples 69 Lung Cancer patients: 26 pre-diagnosis only; 28 at & prediagnosis; 15 at diagnosis only v Combination of LDCT and plasma mi. RNA test v Prognostic value of the mi. RNA assay Sozzi G. et al. , in press
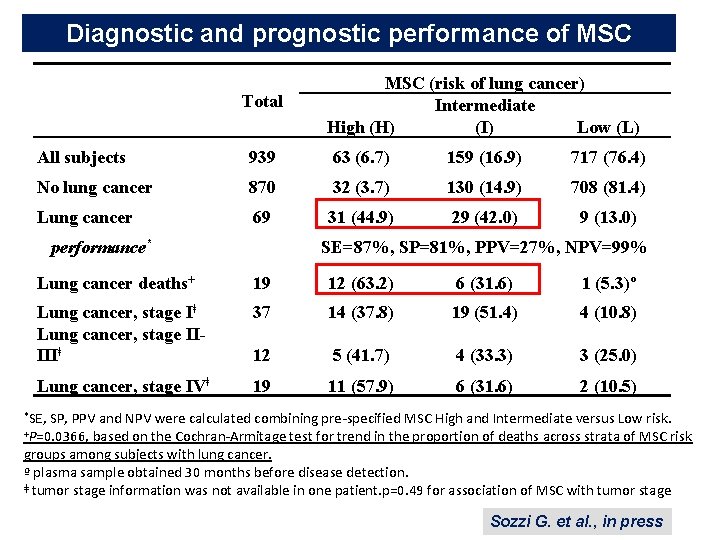
Diagnostic and prognostic performance of MSC Total MSC (risk of lung cancer) Intermediate High (H) (I) Low (L) All subjects 939 63 (6. 7) 159 (16. 9) 717 (76. 4) No lung cancer 870 32 (3. 7) 130 (14. 9) 708 (81. 4) Lung cancer 69 31 (44. 9) 29 (42. 0) 9 (13. 0) performance* SE=87%, SP=81%, PPV=27%, NPV=99% Lung cancer deaths+ 19 12 (63. 2) 6 (31. 6) 1 (5. 3)º Lung cancer, stage Iǂ Lung cancer, stage IIIIIǂ 37 14 (37. 8) 19 (51. 4) 4 (10. 8) 12 5 (41. 7) 4 (33. 3) 3 (25. 0) Lung cancer, stage IVǂ 19 11 (57. 9) 6 (31. 6) 2 (10. 5) *SE, SP, PPV and NPV were calculated combining pre-specified MSC High and Intermediate versus Low risk. +P=0. 0366, based on the Cochran-Armitage test for trend in the proportion of deaths across strata of MSC risk groups among subjects with lung cancer. º plasma sample obtained 30 months before disease detection. ǂ tumor stage information was not available in one patient. p=0. 49 for association of MSC with tumor stage Sozzi G. et al. , in press
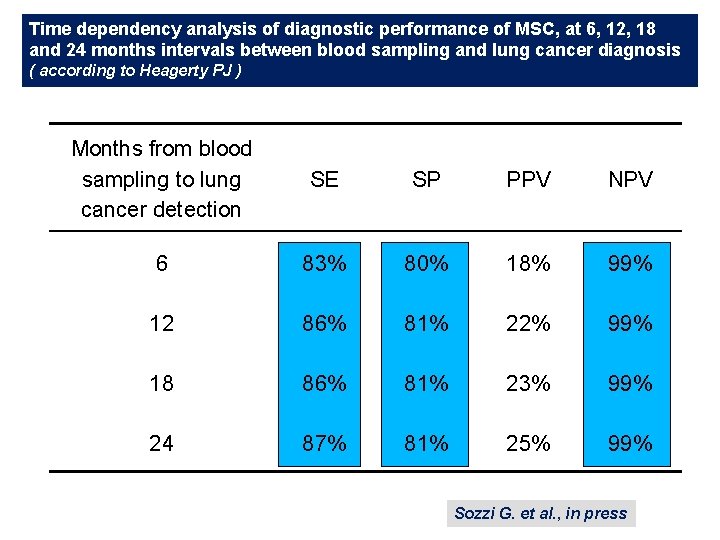
Time dependency analysis of diagnostic performance of MSC, at 6, 12, 18 and 24 months intervals between blood sampling and lung cancer diagnosis ( according to Heagerty PJ ) Months from blood sampling to lung cancer detection SE SP PPV NPV 6 83% 80% 18% 99% 12 86% 81% 22% 99% 18 86% 81% 23% 99% 24 87% 81% 25% 99% Sozzi G. et al. , in press
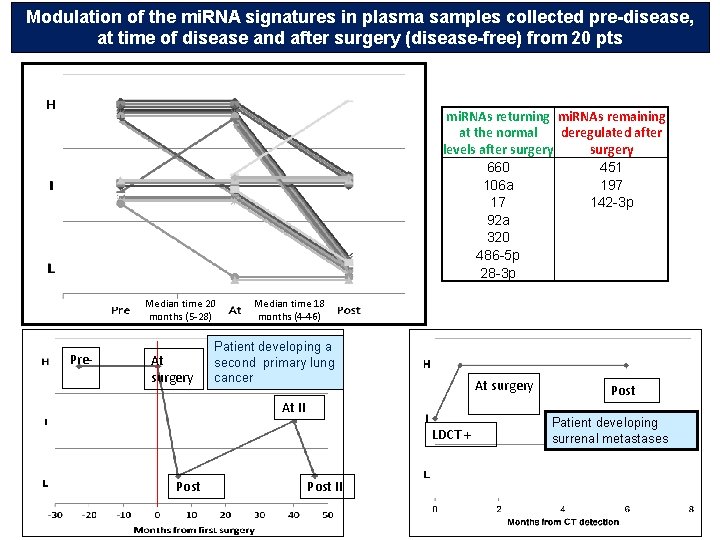
Modulation of the mi. RNA signatures in plasma samples collected pre-disease, at time of disease and after surgery (disease-free) from 20 pts mi. RNAs returning mi. RNAs remaining at the normal deregulated after levels after surgery 660 451 106 a 197 17 142 -3 p 92 a 320 486 -5 p 28 -3 p Median time 20 months (5 -28) Pre- At surgery Median time 18 months (4 -46) Patient developing a second primary lung cancer At surgery At II LDCT + Post II Post Patient developing surrenal metastases

Biomarker-Driven Programs for Lung Cancer Screening Pierre P. Massion, MD Thoracic Program Vanderbilt University Nashville, TN WCLC Oct 30 th, 2013
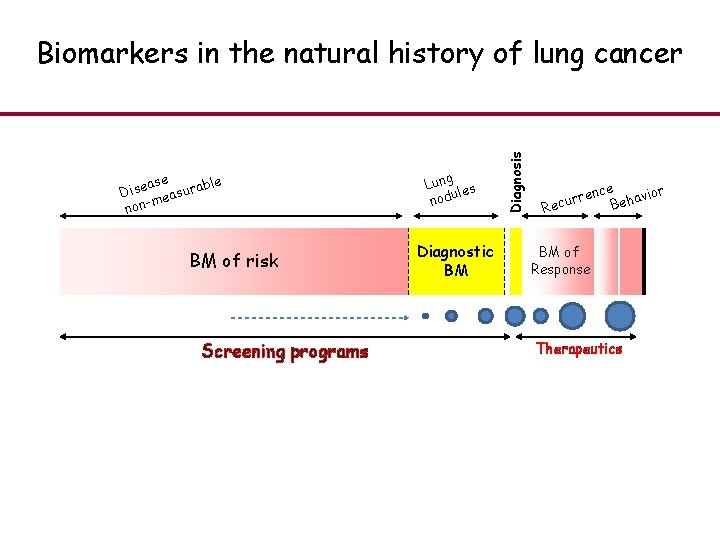
ase ble Dise easura m non- BM of risk Screening programs Lung es l nodu Diagnostic BM Diagnosis Biomarkers in the natural history of lung cancer r nce avio h e B rre Recu BM of Response Therapeutics
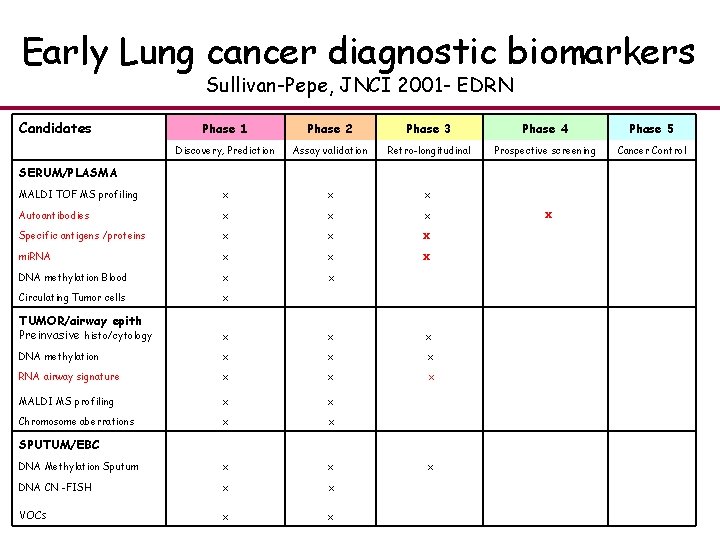
Early Lung cancer diagnostic biomarkers Sullivan-Pepe, JNCI 2001 - EDRN Candidates SERUM/PLASMA Phase 1 Phase 2 Phase 3 Phase 4 Phase 5 Discovery, Prediction Assay validation Retro-longitudinal Prospective screening Cancer Control MALDI TOF MS profiling x x x Autoantibodies x x x Specific antigens /proteins x x x mi. RNA x x x DNA methylation Blood x x Circulating Tumor cells x TUMOR/airway epith Preinvasive histo/cytology x x x DNA methylation x x x RNA airway signature x x x MALDI MS profiling x x Chromosome aberrations x x SPUTUM/EBC DNA Methylation Sputum x x DNA CN -FISH x x VOCs x x
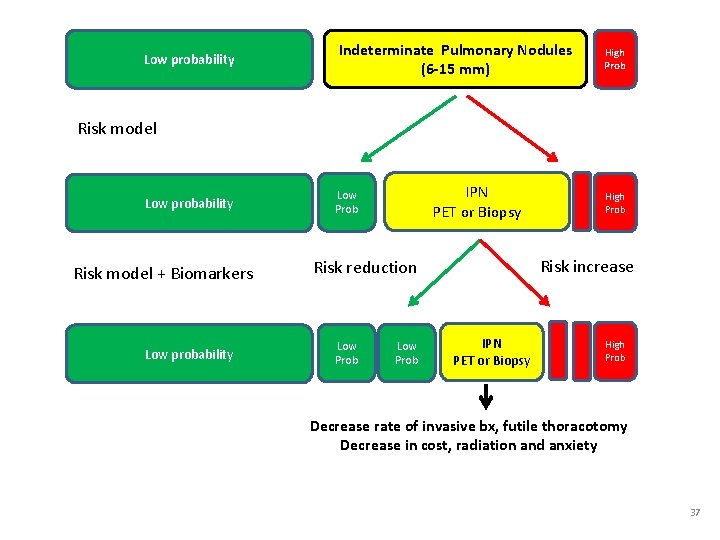
Low probability Indeterminate Pulmonary Nodules (6 -15 mm) High Prob Risk model Low probability Risk model + Biomarkers Low probability IPN PET or Biopsy Low Prob Risk increase Risk reduction Low Prob High Prob IPN PET or Biopsy High Prob Decrease rate of invasive bx, futile thoracotomy Decrease in cost, radiation and anxiety 37

How good should the biomarker be? • Better than standard of care. • What are the metrics? - Performance of the test: PPV & NPV ROC curves (TPR vs FPR). C index comparison Net reclassification Improvement (NRI) index Change in decision making. • De-emphasize Sensitivity and Specificity – – – Pecot, CEBP 2012 Irrelevant (except in early phase of marker evaluation) Not stable across populations Require dichotomization of marker values (loss of information) No information on added value Not actionable metric; PPV or NPV are. Independent of the prevalence of the cancer.
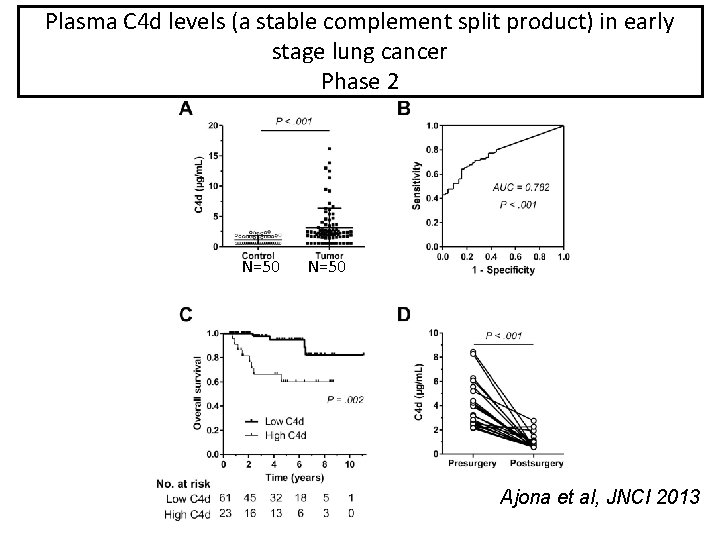
Plasma C 4 d levels (a stable complement split product) in early C 4 d in plasma samples from early lung cancer stage lung cancer Phase 2 N=50 Ajona et al, JNCI 2013
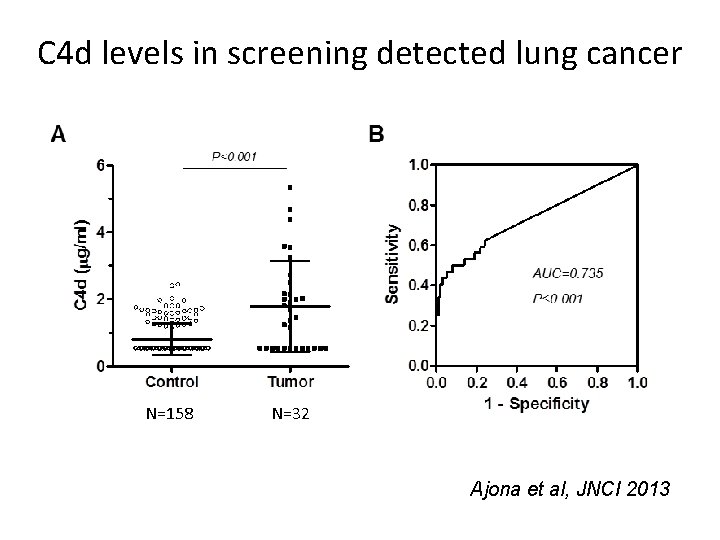
C 4 d levels in screening detected lung cancer N=158 N=32 Ajona et al, JNCI 2013

A Blood-Based Proteomic Classifier for the Molecular Characterization of Pulmonary Nodules Phase 3 • Shotgun Proteomic analysis of tumors. • Selected candidate proteins for testing in the blood • Developed 13 multiple reaction monitoring MRM assays. LRP 1, BGH 3, COIA 1, TETN, TSP 1, ALDOA, GRP 78, ISLR, FRIL, LG 3 BP, PRDX 1, FIBA, GSLG 1 • Training and testing algorithm. Li et al. Sci Transl Med. 2013 Oct
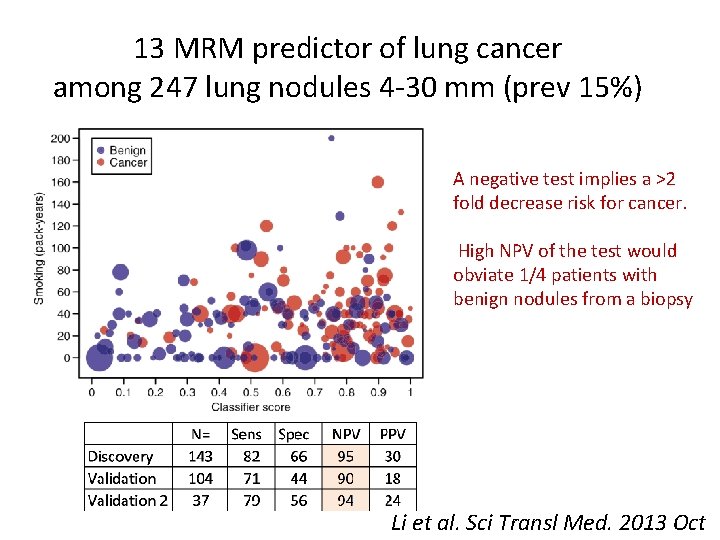
13 MRM predictor of lung cancer among 247 lung nodules 4 -30 mm (prev 15%) A negative test implies a >2 fold decrease risk for cancer. High NPV of the test would obviate 1/4 patients with benign nodules from a biopsy Li et al. Sci Transl Med. 2013 Oct
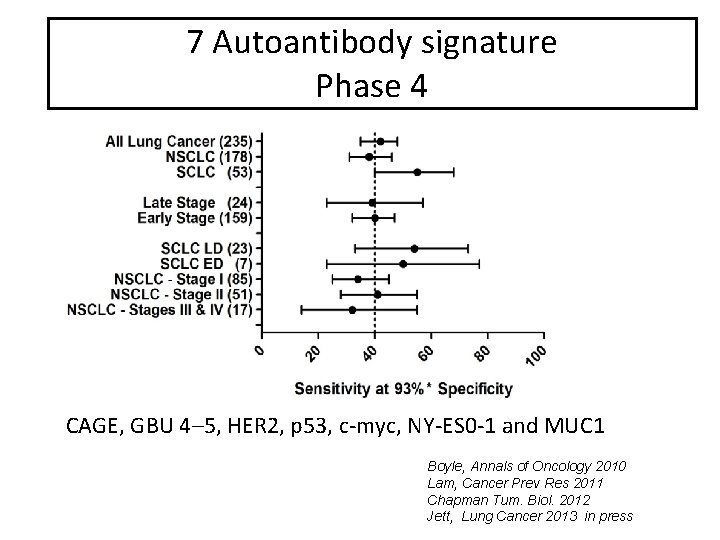
7 Autoantibody signature Phase 4 CAGE, GBU 4– 5, HER 2, p 53, c-myc, NY-ES 0 -1 and MUC 1 Boyle, Annals of Oncology 2010 Lam, Cancer Prev Res 2011 Chapman Tum. Biol. 2012 Jett, Lung Cancer 2013 in press

7 Autoantibody signature Early. CDT- Lung Oncimmune 189 nodules tested with the 7 AAB test Profile + Profile - Cases 19 24 43 Controls 17 129 146 36 153 Sensitivity Specificity PPV NPV Prevalence RR 44. 2 88. 4 52. 8 84. 3 0. 23 3. 36 In nodules 8 -20 mm, the RR is 4. 6 P 2. 20 -006 | Autoantibodies to a panel of lung cancer-associated antigens can provide significant discrimination between malignant and non-malignant lung nodules P. Massion 44
Personalizing the management of indeterminate pulmonary nodules 45
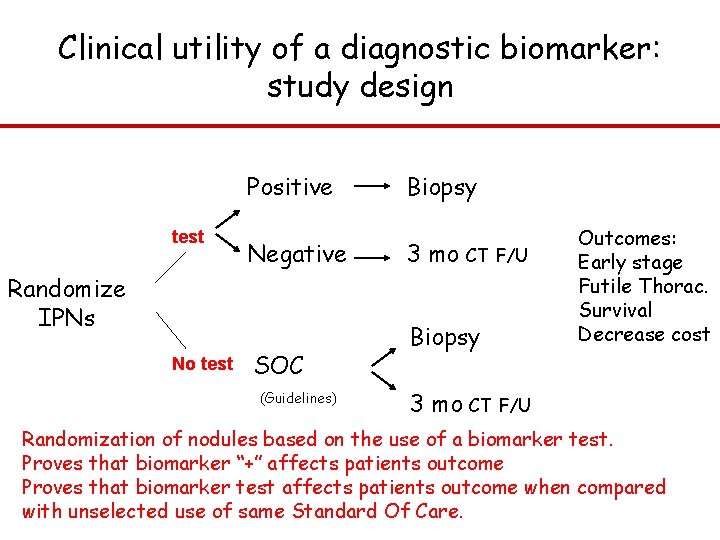
Clinical utility of a diagnostic biomarker: study design Positive test Negative Randomize IPNs No test SOC (Guidelines) Biopsy 3 mo CT F/U Biopsy 3 mo Outcomes: Early stage Futile Thorac. Survival Decrease cost CT F/U Randomization of nodules based on the use of a biomarker test. Proves that biomarker “+” affects patients outcome Proves that biomarker test affects patients outcome when compared with unselected use of same Standard Of Care.

Conclusions • Many early detection candidate biomarkers exist • Few are validated or tested in preclinical setting. Priority to validate existing candidates. • We need to de-emphasize Sensitivity and Specificity and emphasize NPV or PPV with change in decision making. • BM should provide knowledge about added value and therefore should be integrated to clinical, laboratory and imaging routine data. • To demonstrate clinical utility requires significant investment in effort and resources towards biomarkers driven clinical trial.
- Slides: 47