mi RNA targets Using undergraduate molecular biology labs

mi. RNA targets Using undergraduate molecular biology labs to discover targets of mi. RNAs in humans Adam Idica, Jordan Thompson, Irene Munk Pedersen, Pavan Kadandale

mi. RNAs Which of the following is NOT true about mi. RNAs? A. Processed from longer precursors B. Regulate developmental programs C. mi. RNAs are transcriptionally regulated D. mi. RNAs are conserved E. Different mi. RNAs can be separated on a gel

Doing real science! mi. R-NNN is involved in cancers What might we want to know? How to identify mi. R-NNN targets? - Does target need to be completely complementary? - Are all complementary sequences targets? How to identify mi. R-128 targets?

Using computer algorithms for mi. RNA targets Different algorithms to identify targets (Why? ) Which one is best? Common targets better? So, what use are they?
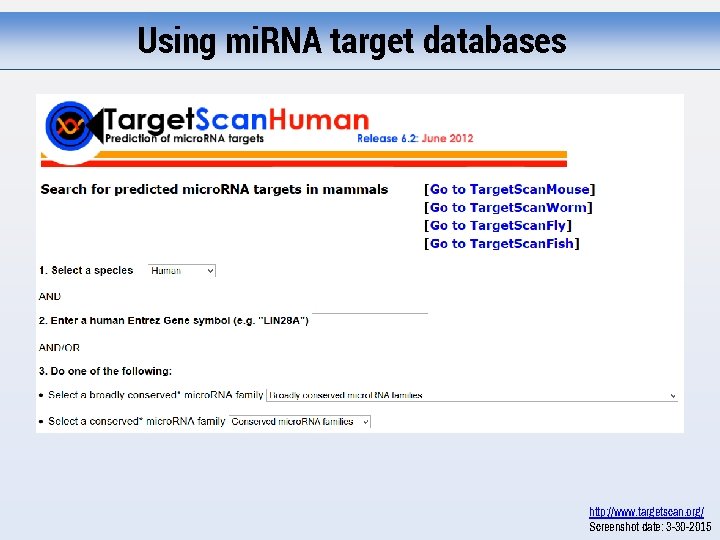
Using mi. RNA target databases http: //www. targetscan. org/ Screenshot date: 3 -30 -2015
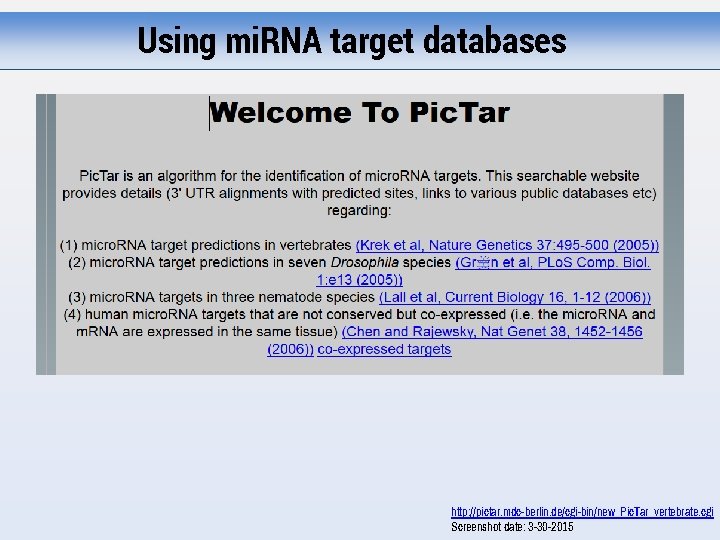
Using mi. RNA target databases http: //pictar. mdc-berlin. de/cgi-bin/new_Pic. Tar_vertebrate. cgi Screenshot date: 3 -30 -2015

Using mi. RNA target databases Database will give you list How will you pick target?
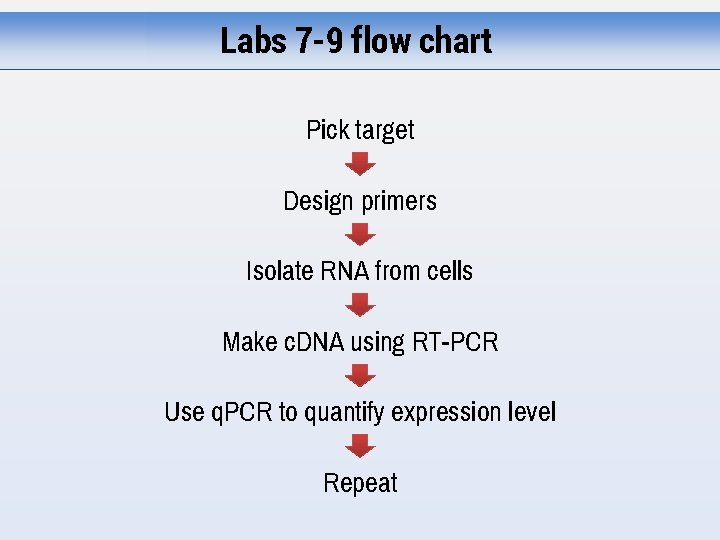
Labs 7 -9 flow chart Pick target Design primers Isolate RNA from cells Make c. DNA using RT-PCR Use q. PCR to quantify expression level Repeat

After picking a target… Pick a target – then what? How will you “validate” target? What controls do you need to include? DNA contamination? Amount of sample? Change in levels due to mi. RNA targeting?

Finding targets Which of the following is NOT required to find mi. R targets? A. RT-PCR B. q. PCR C. Clone target gene D. Isolate total m. RNA E. Make c. DNA

How to design primers? What do you need to know? - Sequence of transcript What are important considerations? - Length - Tm - Primer dimer - Primer self-complementarity - Primer specificity - Sequence runs & GC- content

Primer design If true, which of the following would make you discard a primer A. Primer has GC content of ~62% B. Primer sequence is AGGCGAGTGTTCGCCT C. Predicted Tm is 62°C D. Primer has AT content of 66% E. Primer binds one location on the genome

Primer design You design primers for a gene; using these primers in a PCR yields 1 band. Using the primer in an RT-PCR gives no band. Which of the following is not a valid reason: A. F-primer has GC content of ~78% B. Predicted Tm is 72°C for R-primer C. F-primer binds unique sequence in intron D. R-primers binds unique sequence in exon E. F-primer binds 4 different sites in genome
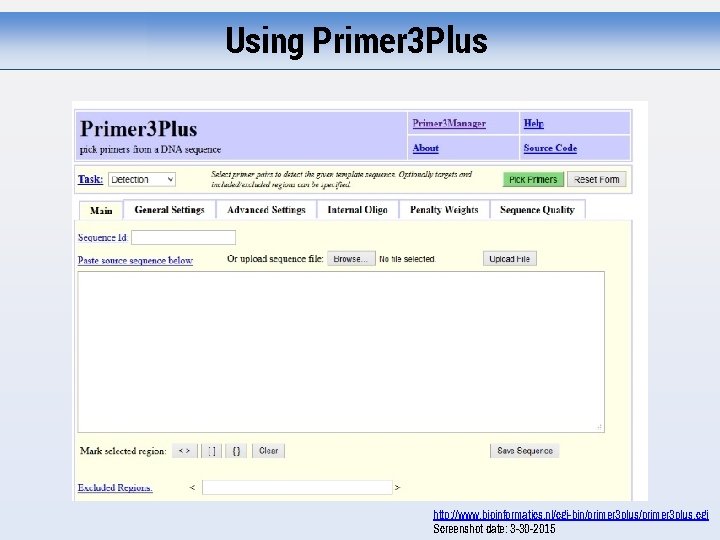
Using Primer 3 Plus http: //www. bioinformatics. nl/cgi-bin/primer 3 plus. cgi Screenshot date: 3 -30 -2015
Oligo. Calc http: //www. basic. northwestern. edu/biotools/Oligo. Calc. html Screenshot date: 3 -30 -2015

Primer design What important parameter have we still not checked? A. Tm B. Primer-primer dimers C. GC-Content D. Primer self-complementarity E. Primer specificity
Primer. BLAST http: //www. ncbi. nlm. nih. gov/tools/primer-blast/ Screenshot date: 3 -30 -2015

Primers designed: Now what? Enter in Google Drive Spreadsheet Lab Section Group number Group members Transcript Gene Reason DO NOT EDIT someone else’s already existing data!
- Slides: 18