Mi DRe G Mining Developmentally Regulated Genes Debashis

Mi. DRe. G: Mining Developmentally Regulated Genes Debashis Sahoo Ph. D, Electrical Engineering, Stanford University Joint work with The Weissman Lab ICBP, Stanford Integrative Cancer Biology Program, Stanford University
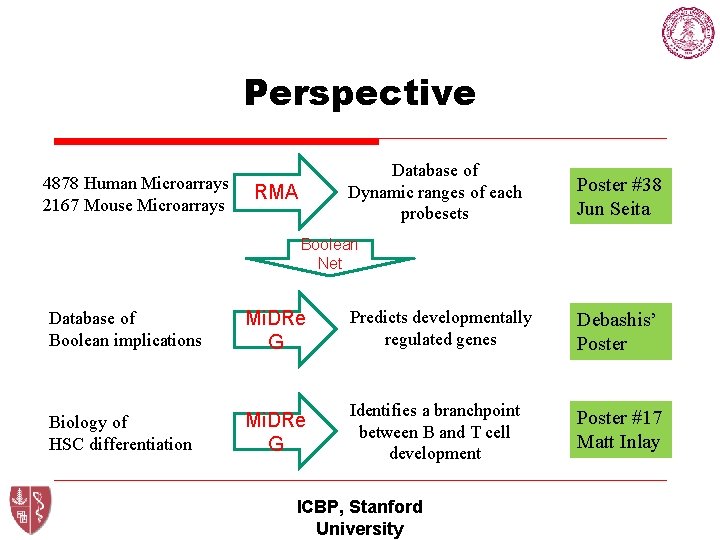
Perspective 4878 Human Microarrays 2167 Mouse Microarrays Database of Dynamic ranges of each probesets RMA Poster #38 Jun Seita Boolean Net Database of Boolean implications Mi. DRe G Predicts developmentally regulated genes Debashis’ Poster Biology of HSC differentiation Mi. DRe G Identifies a branchpoint between B and T cell development Poster #17 Matt Inlay ICBP, Stanford University

Motivation Hard to discover using other approaches Genetics Biochemistry These genes carry out important functions Development and differentiation Surface markers are easy to study ICBP, Stanford University
![Boolean. Net Get data GEO [Edgar et al. 02] Normalize RMA [Irizarry et al. Boolean. Net Get data GEO [Edgar et al. 02] Normalize RMA [Irizarry et al.](http://slidetodoc.com/presentation_image_h2/dc08a7dcdb14676af63ae1ba608067df/image-4.jpg)
Boolean. Net Get data GEO [Edgar et al. 02] Normalize RMA [Irizarry et al. 03] Determine thresholds Discover Boolean relationships Biological interpretation Sahoo et al. Genome Biology 2008 ICBP, Stanford University
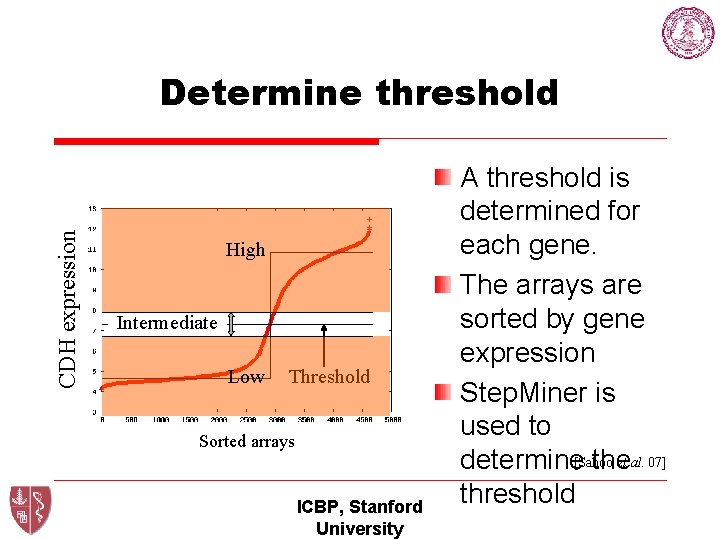
CDH expression Determine threshold High Intermediate Low Threshold Sorted arrays ICBP, Stanford University A threshold is determined for each gene. The arrays are sorted by gene expression Step. Miner is used to et al. 07] determine[Sahoo the threshold
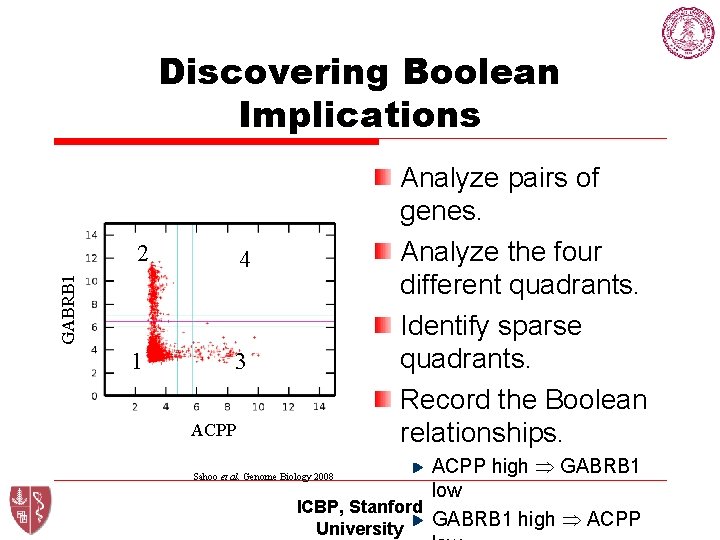
Discovering Boolean Implications 4 1 3 GABRB 1 2 Analyze pairs of genes. Analyze the four different quadrants. Identify sparse quadrants. Record the Boolean relationships. ACPP high GABRB 1 low ICBP, Stanford GABRB 1 high ACPP University Sahoo et al. Genome Biology 2008
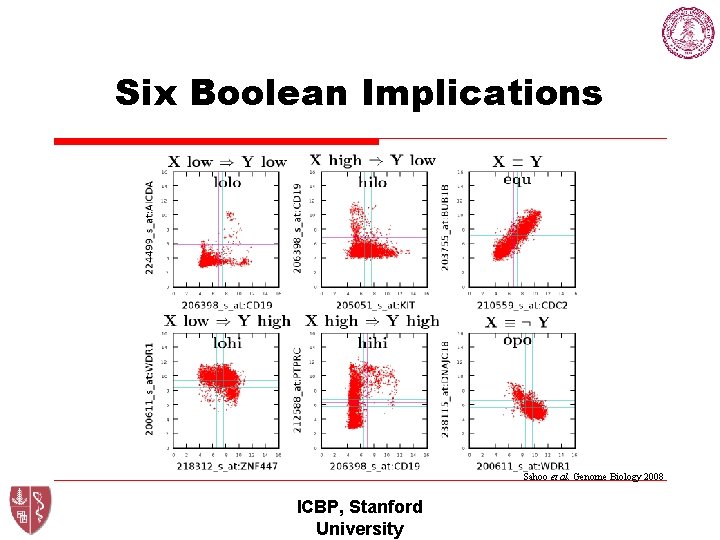
Six Boolean Implications Sahoo et al. Genome Biology 2008 ICBP, Stanford University
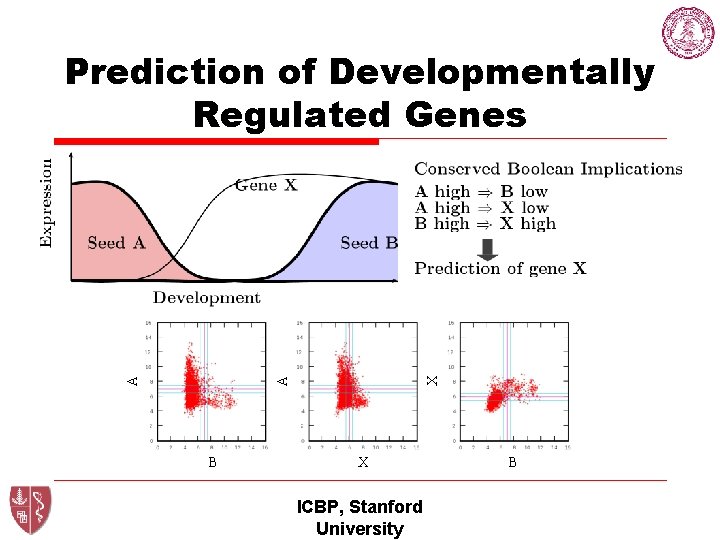
X A A Prediction of Developmentally Regulated Genes B X ICBP, Stanford University B
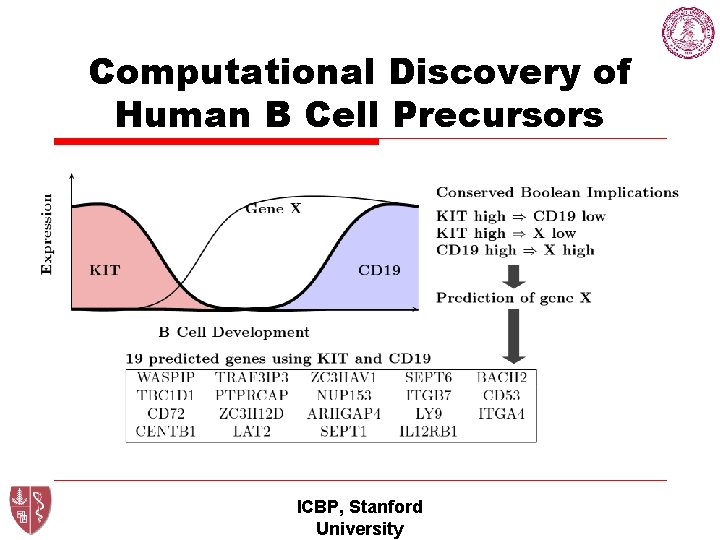
Computational Discovery of Human B Cell Precursors ICBP, Stanford University
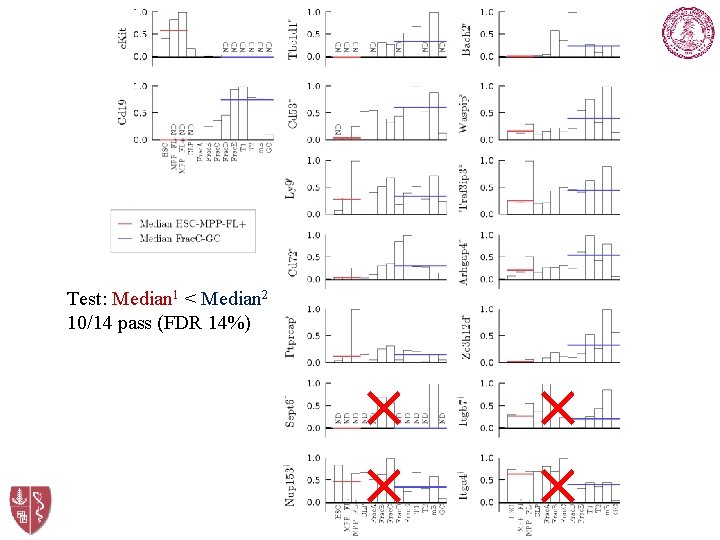
q. PCR Results Test: Median 1 < Median 2 10/14 pass (FDR 14%) × × ICBP, Stanford University × ×
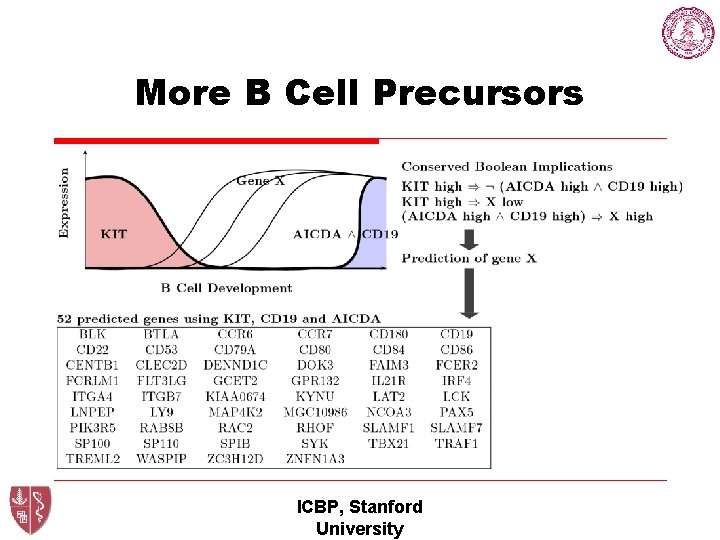
More B Cell Precursors ICBP, Stanford University
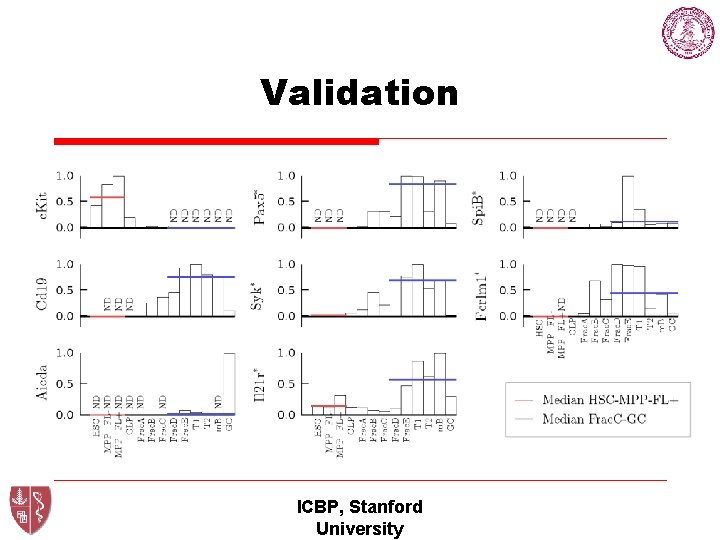
Validation ICBP, Stanford University

Analysis of Predicted Genes Total number of genes predicted: 62 33 genes have been knocked out in mice. [Literature] 18 genes have defects in B cell function and B cell differentiation. 2 genes are known prognostic markers of B cell lymphomas: WASPIP and GCET 2. ICBP, Stanford University

Conclusion Mi. DRe. G uses Boolean implications to predict genes related to B cell development Knockouts of the predicted genes have defects in B cell function and differentiation Mi. DRe. G can be directly applied to other less well-characterized developmental pathway ICBP, Stanford University

Acknowledgements David L. Dill Sylvia K. Plevritis Robert Tibshirani Irving L. Weissman The Weissman Lab: Deepta, Jun, Matt Funding: ICBP Program (NIH grant: 5 U 56 CA 112973 -02) ICBP, Stanford University

The END ICBP, Stanford University

Statistical Tests Compute the expected number of points under the independence model n. Alow = (a 00+ a 01), n. Blow = (a 00+ a 10) total = a 00+ a 01+ a 10+ a 11, observed = a 00 expected = (n. Alow/ total * n. Blow/ total) * total B a 01 a 11 a 00 a 10 A Compute maximum likelihood estimate of the error rate = 1 2 a 00 ( (a + a ) 00 01 + a 00 (a 00+ a 10) ICBP, Stanford University )
- Slides: 17