Methods for Dealing with Time Dependent Confounding Daniel







![Forcing the regimen A 0=0, A 1=0 E[Y 0, 0]= (2. 5*900+2. 5*300+1. 5*760+1. Forcing the regimen A 0=0, A 1=0 E[Y 0, 0]= (2. 5*900+2. 5*300+1. 5*760+1.](https://slidetodoc.com/presentation_image_h/93d328c2b3f0911347c008f8ef843fce/image-13.jpg)





![After IPW E[Y 00]= (2. 5*900+2. 5*300+1. 5*760+1. 5*40)/ (900+300+760+40) = 2. 1 After IPW E[Y 00]= (2. 5*900+2. 5*300+1. 5*760+1. 5*40)/ (900+300+760+40) = 2. 1](https://slidetodoc.com/presentation_image_h/93d328c2b3f0911347c008f8ef843fce/image-20.jpg)







- Slides: 28

Methods for Dealing with Time. Dependent Confounding Daniel, Sousens, De Stavola, Kenward, Sterne (2013)

This article is a tutorial that goes through much of the classic papers on the subject by Jamie Robins and colleagues • Robins JM (1986). A new approach to causal inference in mortality studies with a sustained exposure period – application to control of the healthy worker survivor effect. Mathematical Modelling 7: 1393 -1512. • Robins JM, Hernan MA, Brumback B (2000). Marginal structural models and causal inference in epidemiology. Epidemiology 11: 550 -560. • Robins JM, Blevins D, Ritter G, Wulfsohn M (1992). G-estimation of the effect of prophylaxis therapy for Pneumocystis carinii pneumonia on the survival of AIDS patients. Epidemiology 3: 319336. • Robins JM (1992). Estimation of the time-dependent accelerated failure time model in the presence of confounding factors. Biometrika 79: 321 -334. • …

Classic Confounding

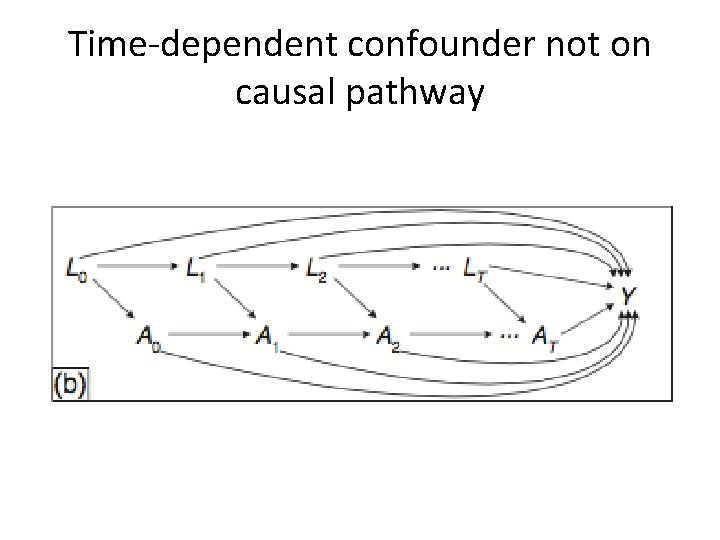
Time-dependent confounder not on causal pathway

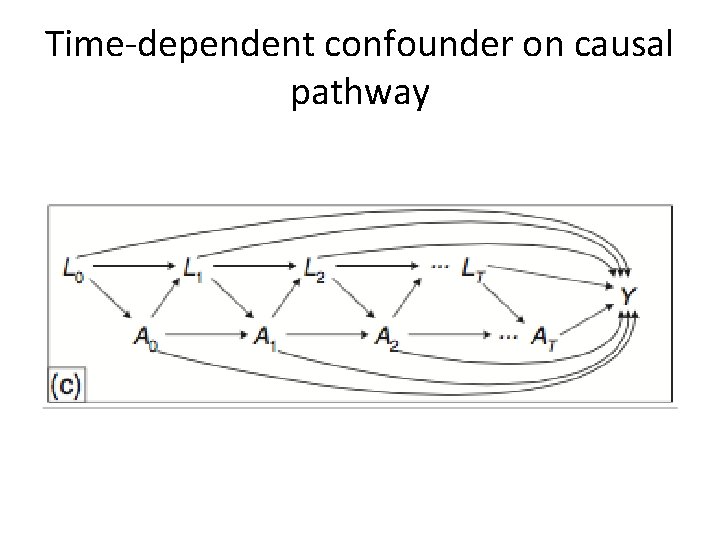
Time-dependent confounder on causal pathway

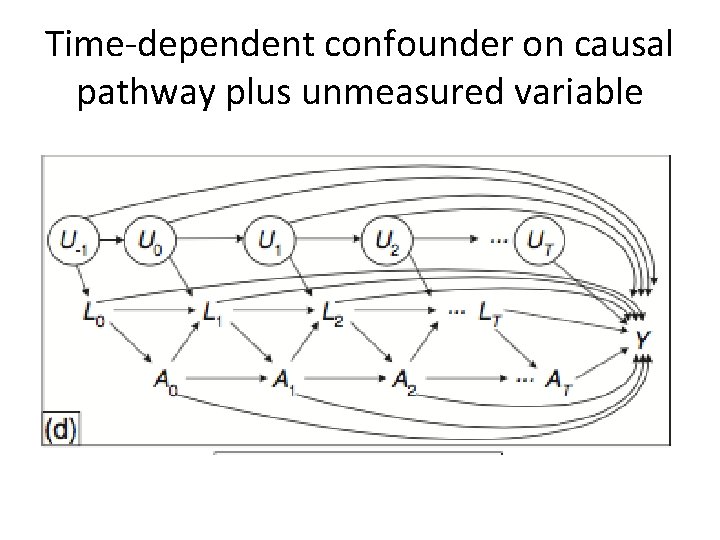
Time-dependent confounder on causal pathway plus unmeasured variable

Causal Effects • • 2. 4. 1. The existence of a causal effect 2. 4. 2. Characterizing the causal effect 2. 4. 3. Joint, total and direct causal effects 2. 4. 4. More parsimonious characterizations – Marginal structural models

Identification • No unmeasured confounding – i. e. , Sequentially ignorable treatment assignment – i. e. , Conditional exchangeability

Static and Dynamic Regimes • Static – Interventions being considered do not depend on the values of a time-varying covariate. – Impact of being on ART. • Dynamic – Interventions being considered depend on the values of a time-varying covariate. – Impact of waiting to start ART until CD 4 drops below a certain level. – We’ll talk more about dynamic regimes later in the course.

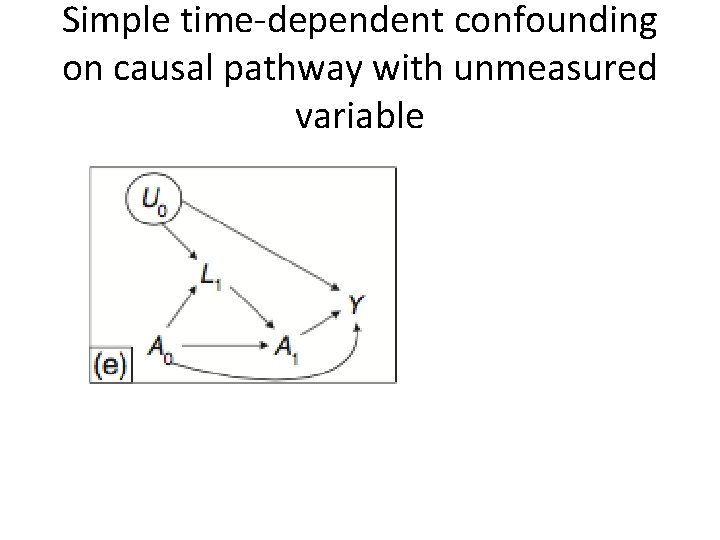
Simple time-dependent confounding on causal pathway with unmeasured variable

G-computation • Standardization • G-computation formula – Generalization of standardization

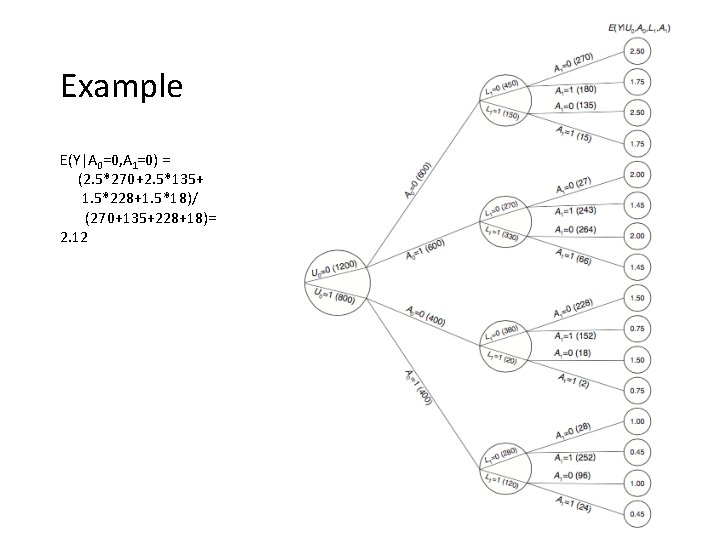
Example E(Y|A 0=0, A 1=0) = (2. 5*270+2. 5*135+ 1. 5*228+1. 5*18)/ (270+135+228+18)= 2. 12
![Forcing the regimen A 00 A 10 EY 0 0 2 59002 53001 57601 Forcing the regimen A 0=0, A 1=0 E[Y 0, 0]= (2. 5*900+2. 5*300+1. 5*760+1.](https://slidetodoc.com/presentation_image_h/93d328c2b3f0911347c008f8ef843fce/image-13.jpg)
Forcing the regimen A 0=0, A 1=0 E[Y 0, 0]= (2. 5*900+2. 5*300+1. 5*760+1. 5*40)/ (900+300+760+40) = 2. 1

Some intuition • P(u 0, a 0, l 1, a 1, y) =P(u 0) P(a 0|u 0) P(l 1|u 0, a 0) P(a 1|u 0, a 0, l 1) x P(y|u 0, a 0, l 1, a 1) • Force everyone to have a 0=0 and a 1=0. – Pearl would say, do(a 0=0, a 1=0). • P(u 0, do(a 0=0), l 1, do(a 1=0), y) =P(u 0) P(l 1|u 0, do(a 0=0)) P(y|u 0, do{a 0=0}, l 1, do{a 1=0}) =P(u 0) P(l 1|u 0, a 0=0) P(y|u 0, a 0=0, l 1, a 1=0) by do-calculus • E[Y(0, 0)]=∑u 0 ∑l 1 E[y|u 0, a 0=0, l 1, a 1=0] P(l 1|u 0, a 0=0) P(u 0) = ∑l 1 E[y|a 0=0, l 1, a 1=0] P(l 1|a 0=0) (after some math based on no confounding assumptions implied by the graph)

Discuss students’ implementation of simple example

Other g-computation stuff • Estimating parameters of marginal structural model (done post-hoc) • Confidence intervals with bootstrap • Monte Carlo methods needed as number of time points increases • Addressing loss to follow-up under MAR assumption • Software

Inverse probability weighted (IPW) marginal structural models (MSM) • MSM: • Naïve model: • IP Weights: • MSM estimated by fitting the naïve model with IP Weights.

Experimental treatment assignment assumption • We usually call this the positivity assumption

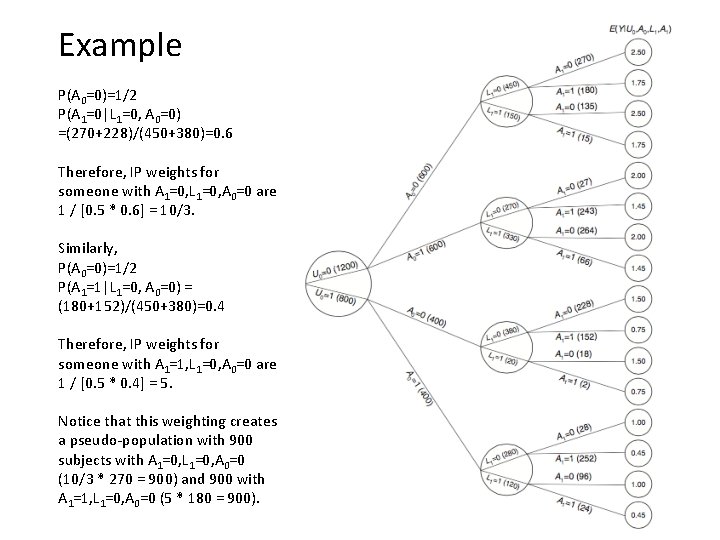
Example P(A 0=0)=1/2 P(A 1=0|L 1=0, A 0=0) =(270+228)/(450+380)=0. 6 Therefore, IP weights for someone with A 1=0, L 1=0, A 0=0 are 1 / [0. 5 * 0. 6] = 10/3. Similarly, P(A 0=0)=1/2 P(A 1=1|L 1=0, A 0=0) = (180+152)/(450+380)=0. 4 Therefore, IP weights for someone with A 1=1, L 1=0, A 0=0 are 1 / [0. 5 * 0. 4] = 5. Notice that this weighting creates a pseudo-population with 900 subjects with A 1=0, L 1=0, A 0=0 (10/3 * 270 = 900) and 900 with A 1=1, L 1=0, A 0=0 (5 * 180 = 900).
![After IPW EY 00 2 59002 53001 57601 540 90030076040 2 1 After IPW E[Y 00]= (2. 5*900+2. 5*300+1. 5*760+1. 5*40)/ (900+300+760+40) = 2. 1](https://slidetodoc.com/presentation_image_h/93d328c2b3f0911347c008f8ef843fce/image-20.jpg)
After IPW E[Y 00]= (2. 5*900+2. 5*300+1. 5*760+1. 5*40)/ (900+300+760+40) = 2. 1

Stabilized Weights • Stabilized weights with baseline covariates – MSM must include baseline covariates.

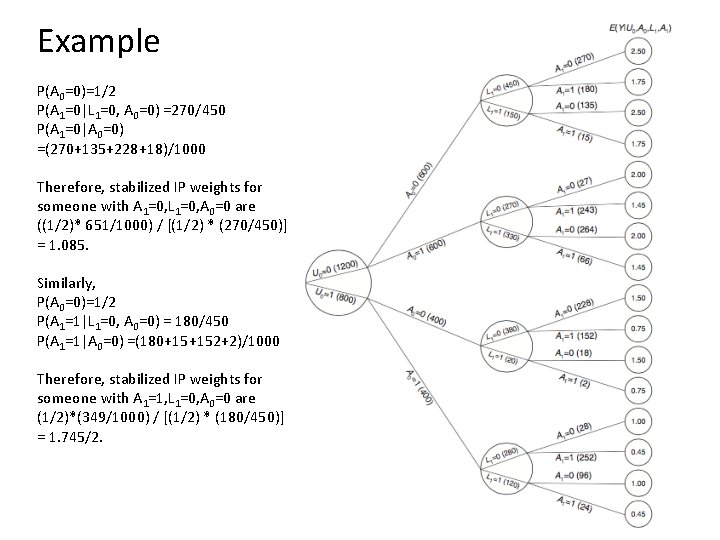
Example P(A 0=0)=1/2 P(A 1=0|L 1=0, A 0=0) =270/450 P(A 1=0|A 0=0) =(270+135+228+18)/1000 Therefore, stabilized IP weights for someone with A 1=0, L 1=0, A 0=0 are ((1/2)* 651/1000) / [(1/2) * (270/450)] = 1. 085. Similarly, P(A 0=0)=1/2 P(A 1=1|L 1=0, A 0=0) = 180/450 P(A 1=1|A 0=0) =(180+15+152+2)/1000 Therefore, stabilized IP weights for someone with A 1=1, L 1=0, A 0=0 are (1/2)*(349/1000) / [(1/2) * (180/450)] = 1. 745/2.

Discuss student’s implementation of simple example

Other IPW-MSM stuff • Loss to follow-up

Comparison of g-computation and IPW

Parametric Extensions • Everything is essentially the same except you can no longer use saturated models so smoothing assumptions are made. • Model misspecification can be an issue. • Collapsibility can also be an issue.

The g-null paradox • Given enough data, we would reject the causal null hypothesis even when it is true. • When standard parsimonious models are chosen for the conditional distributions of Y and L, then when combined using the gcomputation formula, the implied distribution for Ya-bar is a complex function of a-bar, so that Ya-bar is not independent of a-bar without additional assumptions.

Strengths and Limitations G-computation Strengths Limitations 1. Extension of standardization 2. Can handle complex interventions 3. Each hypothetical intervention can be compared with no intervention (i. e. , the status quo) 1. MSM parameters are post hoc estimated 2. Computationally intensive 3. G-null paradox 4. Model misspecification is a big concern because fitting lots of models. IPW Strengths Limitations 1. Easier to explain to subject-matter experts. (? ) 2. Analyses are more easily implemented. 3. Only models for treatment assignment and MSM need to be specified. 4. Clear set of parameters that are estimated that can be used for testing. 1. Unstable and inefficient if extreme weights. Generally less efficient than properly specified g-computation. 2. Continuous treatments hard to handle. 3. Requires positivity assumption. 4. Interactions with time-varying covariates cannot be studied.