METAGENOMICS SEQUENCING THE UNKNOWN Mariana de Britto Wilson

METAGENOMICS: SEQUENCING THE UNKNOWN Mariana de Britto Wilson Master in Advanced Genetics - UAB

• Introduction • Metagenomics methods: intrinsic versus extrinsic Overview • Functional profiling and community-level expression • Environmental studies – Sargasso Sea example • Human Microbiome studies • Metagenomics and Personalized medicine
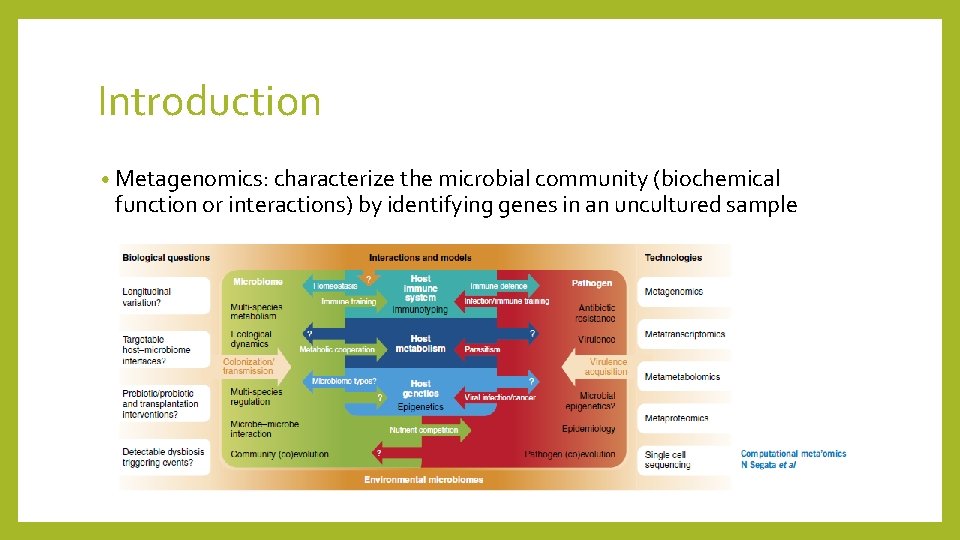
Introduction • Metagenomics: characterize the microbial community (biochemical function or interactions) by identifying genes in an uncultured sample

r. RNA gene sequencing (amplicon sequencing) Cultureindependent methods • 1 st culture-independent method • More efficient to assess overall phylogeny and diversity • Very popular • Illumina/ Roche 454 platforms Whole Metagenome Shotgun sequencing (WMS) • Specific assemblers (Meta. Velvet, Khmer, metamos, Meta-IDBA, Meta-Orfa) • Greater cost per sample • More complex bioinformatic analysis • Higher taxonomic resolution in identifying organisms

Metagenomics: extrinsic vs. Intrinsic approaches

Functional community profiling • Assembly-free read-based approaches Reference genome read-mapping • Translated protein database searches • • Genes in longer assembled contigs • • Protein-coding genes in contigs assigned functional categories by orthology relations with database sequences Automated pipelines

Community interactions and expression • Correlation and association approaches to determine co-occurrence and coexclusion interactions • • Gene and taxon-specific expression patterns • • Need for multi-organism metabolic models to explain interactions Hard to isolate m. RNA Single cell sequencing Predictive models for community responses (Human Intestinal Microbial Ecosystem model)
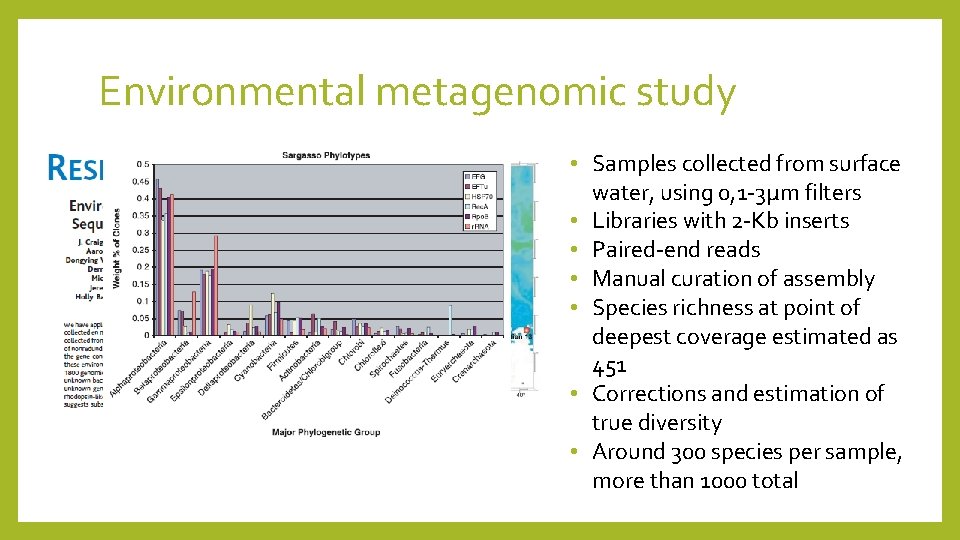
Environmental metagenomic study • Samples collected from surface water, using 0, 1 -3µm filters • Libraries with 2 -Kb inserts • Paired-end reads • Manual curation of assembly • Species richness at point of deepest coverage estimated as 451 • Corrections and estimation of true diversity • Around 300 species per sample, more than 1000 total

Conclusions • More phylogenetic markers than r. RNA studies • Same basic taxonomic groups identified, but different quantitative results for some • New insights into some biochemical processes Limitations • • Well conserved regions can cause scaffold breaks Related organisms can co-assemble Less sequence coverage of low abundance organisms Sequencing cost

Human gut microbiome • • • Random DNA libraries from fecal Conclusions specimens of 2 healthy humans 65059 and 74462 high quality sequence Functionally, many microbiome genes reads generated involved in various digestion processes • were with respect to the human 50164 enriched ORFs predicted Archaea and genome Bacteria • More studiesto needed to analyze Comparison B. longum and M. changes smithii toto microbiome and 1 determine biomarkers of assess diversity: archaeal phylotype and health 72 bacterial phylotypes (16 novel)
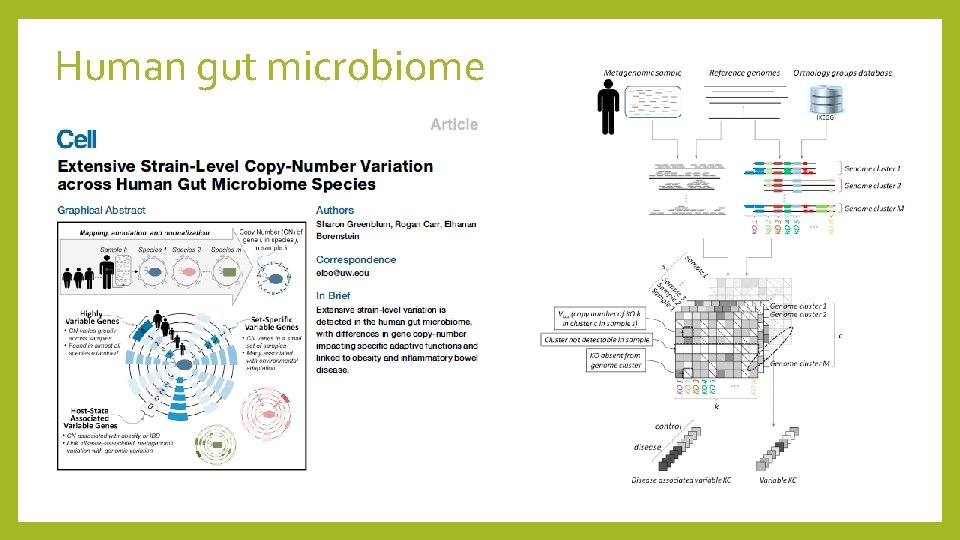
Human gut microbiome

Human gut microbiome Conclusions • Copy number variation is prevalent in the gut environment • More intraspecies variation for transport and signaling functions • Role of environment in bacterial evolution • Adaptability to host immune responses

Human Microbiome Project
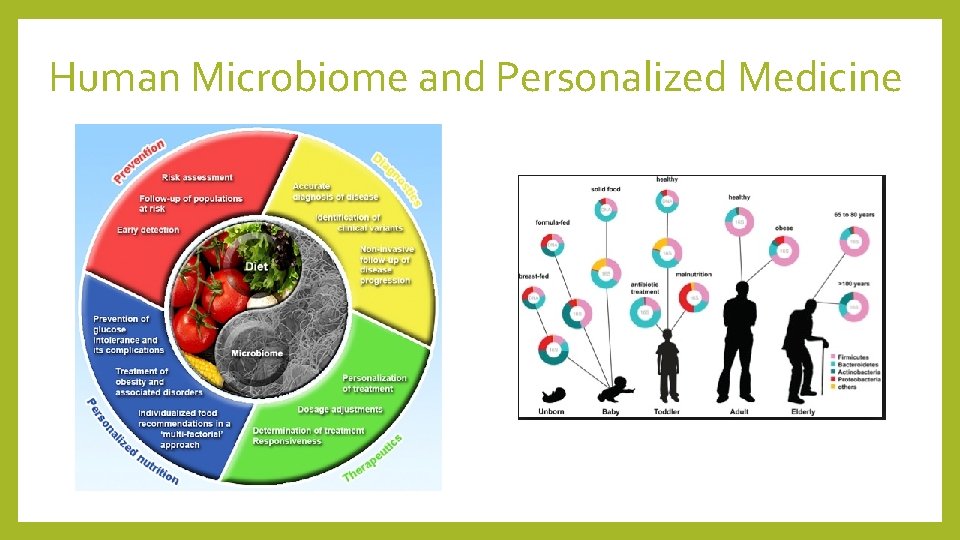
Human Microbiome and Personalized Medicine

Personalized medicine - limitations • Great interpersonal variability • Rapid changes to microbial composition • Experimental parameters & techniques can act as confounding factors • No unified protocol for data analysis or biomarkers • Clinical confounding factors • Influence of environment and human genetics

References • Segata N, Boernigen D, Tickle TL, Morgan XC, Garrett WS, Huttenhower C. Computational meta'omics for microbial community studies. Mol Syst Biol. 2013; 9: 666. • Venter J. C. , Remington K. , Heidelberg J. F. , Halpern A. L. , Rusch D. , Eisen J. A. , et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 2004; 304 (5667): 66– 74. • Greenblum S, Carr R, Borenstein E. Extensive strain-level copy-number variation across human gut microbiome species. Cell. 2015; 160(4): 583 -594. • Gill SR, Pop M, Deboy RT, et al. Metagenomic analysis of the human distal gut microbiome. Science. 2006; 312(5778): 1355 -9. • N. Zmora, D. Zeevi, T. Korem, E. Segal, E. Elinav. Taking it personally: personalized utilization of the human microbiome in health and disease. Cell Host & Microbe. 2016; Vol. 19(1): 12 -20.

QUESTIONS?
- Slides: 17