Metagenomics of Microbial Communities Scott Sproule Cam Mac

Metagenomics of Microbial Communities Scott Sproule Cam Mac. Millan Daniel Hann

Outline Context - A brief history of microbiology - A brief history of genomics - Defining metagenomics Metagenomics - Transcending genomics - Accurate diversity measurements Two approaches of metagenomics - Sequence based approach - Function based approach Environmental analysis - Marine - Soil Applications of Metagenomics - Industrial - Agriculture & Renewable Energy - Environmental remediation - Life sciences Conclusion

Context
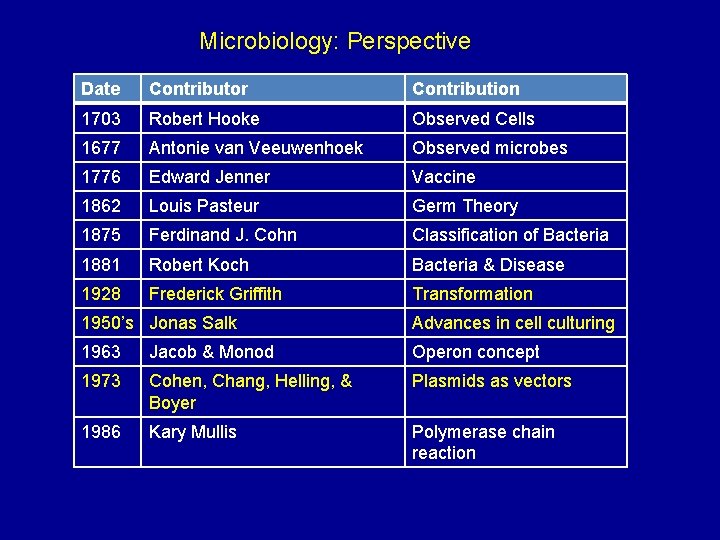
Microbiology: Perspective Date Contributor Contribution 1703 Robert Hooke Observed Cells 1677 Antonie van Veeuwenhoek Observed microbes 1776 Edward Jenner Vaccine 1862 Louis Pasteur Germ Theory 1875 Ferdinand J. Cohn Classification of Bacteria 1881 Robert Koch Bacteria & Disease 1928 Frederick Griffith Transformation 1950’s Jonas Salk Advances in cell culturing 1963 Jacob & Monod Operon concept 1973 Cohen, Chang, Helling, & Boyer Plasmids as vectors 1986 Kary Mullis Polymerase chain reaction

Main Techniques Microscopic Techniques - Direct observation - Combine with - Staining - Isotopes - Flourescence Culturing Techniques - Growing - Isolation - Examination - Manipulation - experimentation Inconsistent estimates of diversity and organisms numbers

Accounting for Inconsistencies “The Great Plate Count Anomaly” - How much were they missing? - What were they missing? - Why were they missing it? It was clear that there were many viable cells that could not be cultured

“Unculturability” Environmental - Nutritional factors - Signaling factors - Essential factors - Artificial reproduction difficult Bacteria - Specific requirements - Competition - Community structure - Defense mechanisms “Can we culture the unculturable?

Genome - Common all living things on earth - A universal language - An instruction manual - A toolbox - A record of history
Genomics the study of whole genomes Nucleotides DNA Genes Genomes
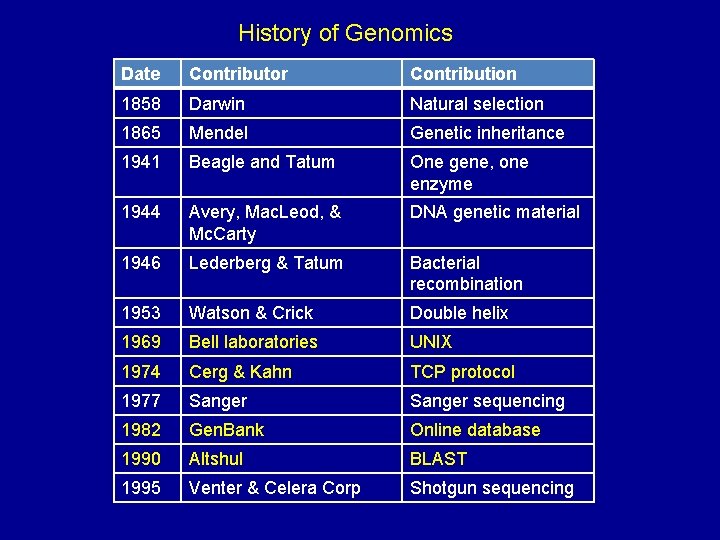
History of Genomics Date Contributor Contribution 1858 Darwin Natural selection 1865 Mendel Genetic inheritance 1941 Beagle and Tatum One gene, one enzyme 1944 Avery, Mac. Leod, & Mc. Carty DNA genetic material 1946 Lederberg & Tatum Bacterial recombination 1953 Watson & Crick Double helix 1969 Bell laboratories UNIX 1974 Cerg & Kahn TCP protocol 1977 Sanger sequencing 1982 Gen. Bank Online database 1990 Altshul BLAST 1995 Venter & Celera Corp Shotgun sequencing

Genomics Today Rapid sequencing of whole genomes The pinnacle of genomics? Metagenomics transcends genomics Multiple genome level

Metagenomics Meta-analysis – combination of separate analysis Genomics – analysis of an organisms genetic material

What is metagenomics? Study of the collection of microbial genomes or genome fragments through direct extraction A culture independent technique that can provide metaanalytic level information about: - Population structure - Genetic Diversity - Functional elements - Novel genetic material A synthesis of a number of fields: - Molecular Genetics - Microbiology - Bioinformatics - Population Genetics - Computer science

Applying Metagenomics to Microbial Communities Traditional methods of quantifying microbial diversity - Sample environments independently - Isolate each species through culturing techniques - Characterize through biochemical & sequencing techniques Realistic? - Very labour intensive - Time estimate: 100’s of years - Incredibly expensive $$$$$$ - Most organisms can not currently be be isolated through culture

Microorganisms Are Everywhere!

Scratching the surface

Tools of Metagenomics - Cloning techniques PCR Cutting edge sequencing techniques Bioinformatics Open source databases - Genbank - Protein Database - Tree. BASE
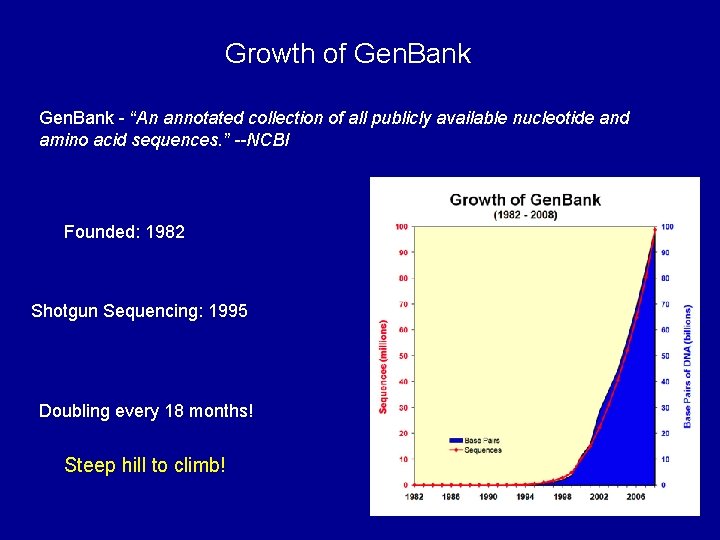
Growth of Gen. Bank - “An annotated collection of all publicly available nucleotide and amino acid sequences. ” --NCBI Founded: 1982 Shotgun Sequencing: 1995 Doubling every 18 months! Steep hill to climb!
Approaches to metagenomics?
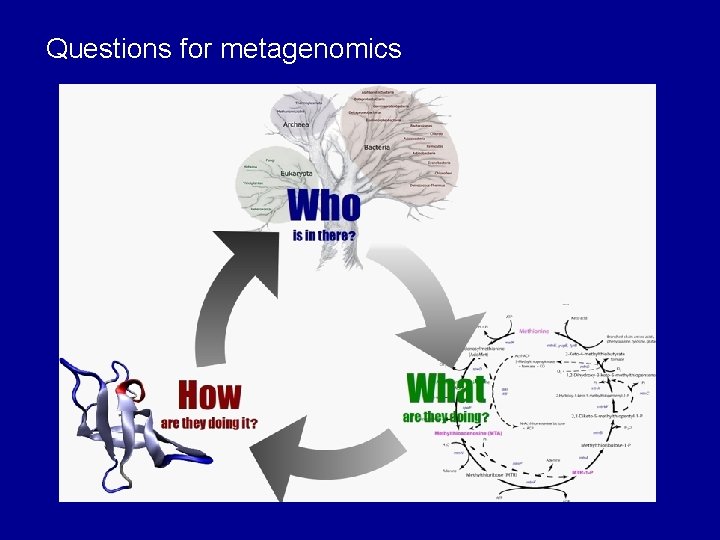
Questions for metagenomics
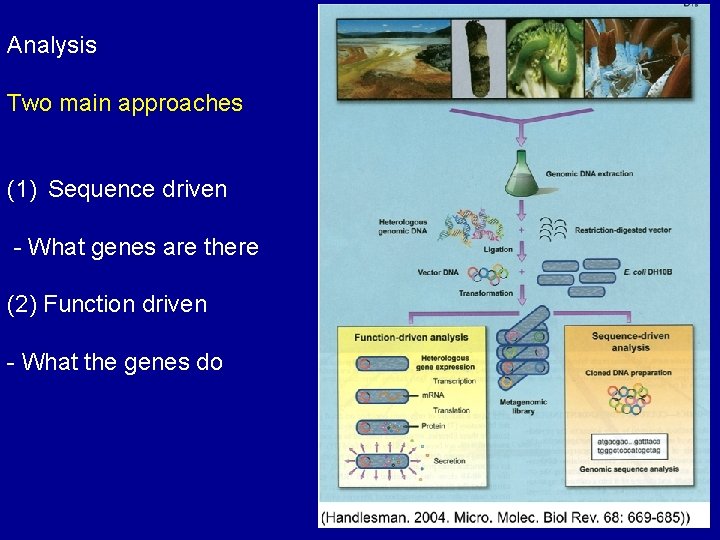
Analysis Two main approaches (1) Sequence driven - What genes are there (2) Function driven - What the genes do

Metagenomic Process Determine what the genes are Sequence driven approaches Extract DNA - Data collection Relies on conserved DNA Phylogenic analysis Used to measure biological diversity Determine what the genes do Functional driven approaches - Functional screening - Can identify novel genes - Relies on gene expression - Proteins

Sequence-Based Approach

Sequencing • One way to classify metagenomic fragments • Relies on nucleotide diversity analysis – Discriminate between species Seq. A GACTACGATCCGTATACGCACA--GGTTCAGAC || |||||||| Seq. B. GAATACGAGCCGTATACGCACACAGGTTCAGA • Requires use of online databases – Ex: BLAST in Gen. Bank Compares “unknown to known”

Restrictions Genomics Metagenomics Whole Genome Sequenced ✔ ✖ Know Species of Origination ✔ ✖ ✖ Many DNA elements Identified ✔

Sequence Metagenomics • Not necessary to determine species of origin • Obtain large volume of data ~ Less redundant • Fragment’s = 20 bp – 700 bp • Assembled sequence reads don’t exceed 5000 bp
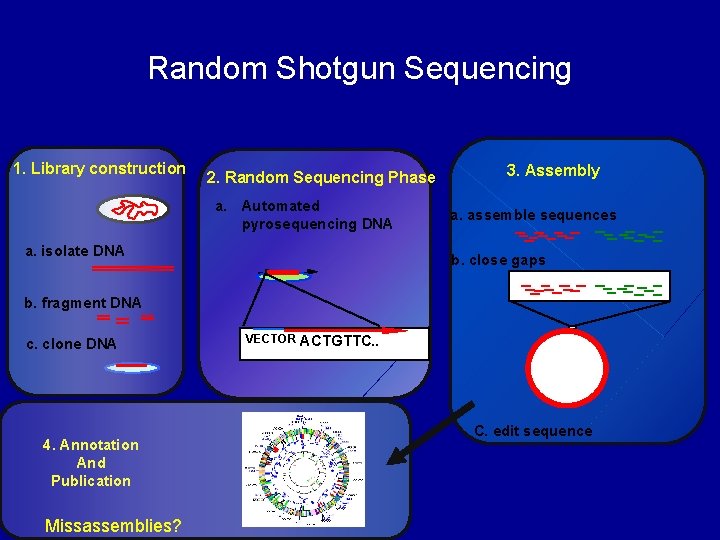
Random Shotgun Sequencing 1. Library construction 2. Random Sequencing Phase a. Automated pyrosequencing DNA a. isolate DNA 3. Assembly a. assemble sequences b. close gaps b. fragment DNA c. clone DNA 4. Annotation And Publication Missassemblies? VECTOR ACTGTTC. . . C. edit sequence

Random Sequencing Objective – – – To estimate bacterial biodiversity ~ Species Richness Identify 1000’s of prokaryotic, viral & eukaryotic species Mass amounts of genomic data obtained Does not depend on PCR Put sequence in computer BLAST – Studies in: • • • Sea water Soil microbial mats Dead whale carcass Feces etc. Organism level (microbiome) – Microorganisms are Everywhere!

Sequence Specific: Phylogenic • Look at evolutionary relationships

Key Challenge What to look for? • Analysis based on evolutionarily conserved marker sequences Want – High conservation across species – Slight & measureable changes over millions of years 16 S r. RNA

16 S r. RNA Value • Vital for translation – Essential • Short • Conserved within a species • Different between different species • Very slow mutation rate Species Concept - Sequence based arbitrary - ~ Consensus: ~97% identity = species - Changing all the time
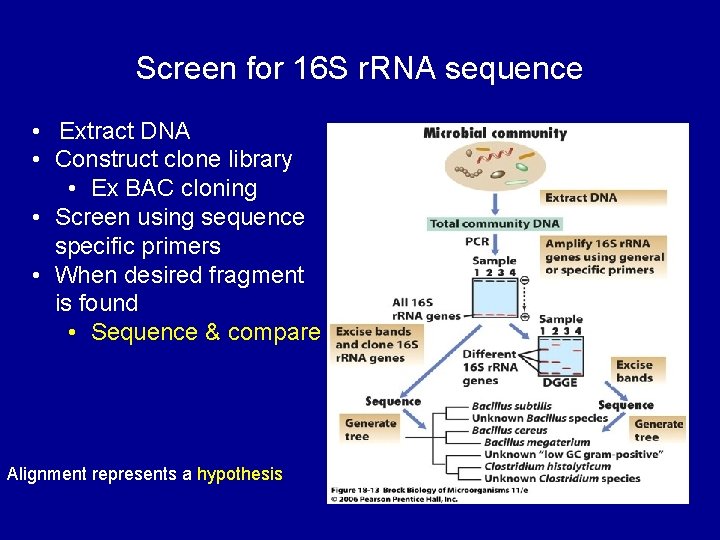
Screen for 16 S r. RNA sequence • Extract DNA • Construct clone library • Ex BAC cloning • Screen using sequence specific primers • When desired fragment is found • Sequence & compare Alignment represents a hypothesis

Function Based Approach
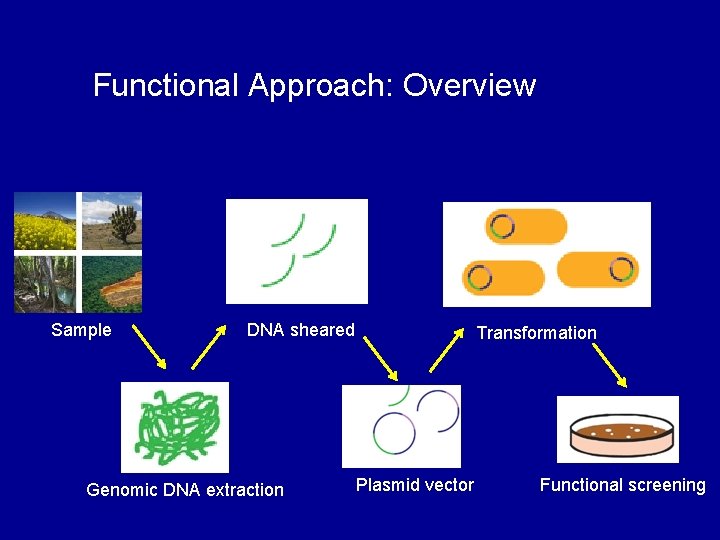
Functional Approach: Overview Sample DNA sheared Genomic DNA extraction Transformation Plasmid vector Functional screening
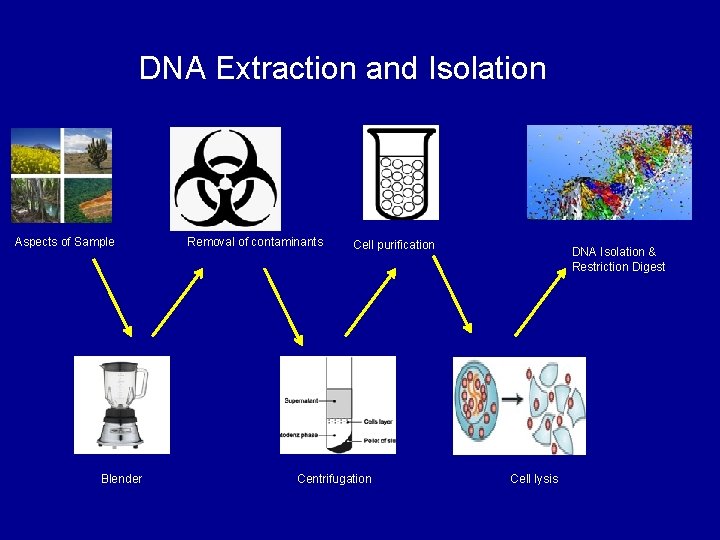
DNA Extraction and Isolation Aspects of Sample Blender Removal of contaminants Cell purification Centrifugation DNA Isolation & Restriction Digest Cell lysis
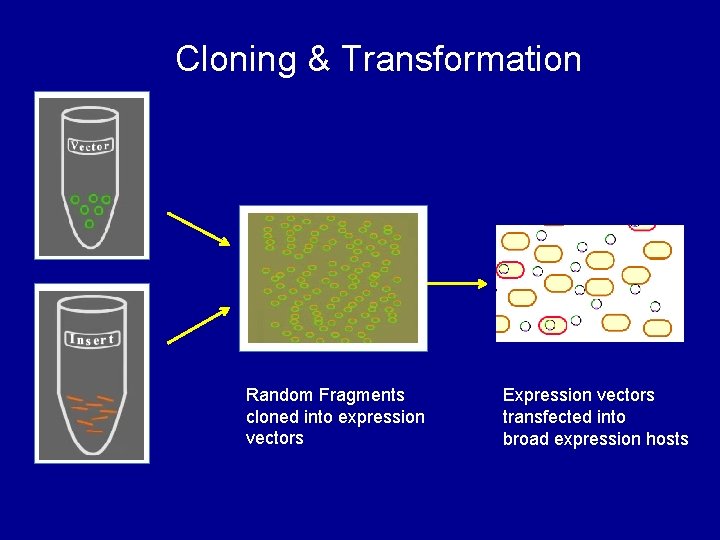
Cloning & Transformation Random Fragments cloned into expression vectors Expression vectors transfected into broad expression hosts

Plasmid Expression vectors
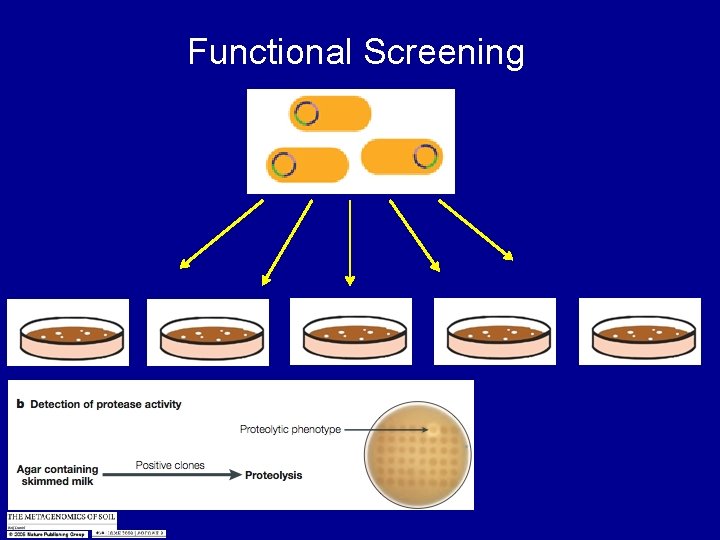
Functional Screening

Treasure Hunting Looking for novel genes Biological tool boxes Detergents - Proteases - Lipases - Esterase’s Antibiotics - Novel antibiotics - Not synthetic - Mutate the antibiotics

Marine Metagenomics - Microbes = 90% of marine biomass - 98% of Primary Producers in Sea -. 001 -. 1% are cultivable

Ocean Exploration Craig Venter
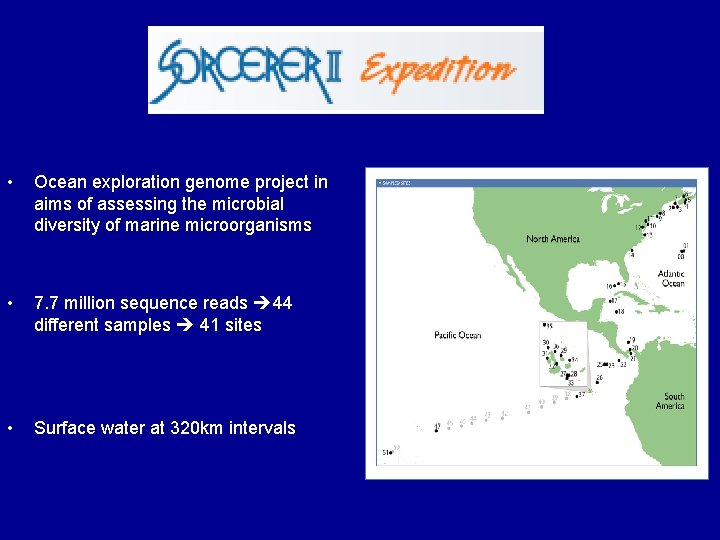
• Ocean exploration genome project in aims of assessing the microbial diversity of marine microorganisms • 7. 7 million sequence reads 44 different samples 41 sites • Surface water at 320 km intervals

Sample Collection • Determine physical characteristics of sample site – Salinity, p. H, depth, dissolved O 2, temperature – Filtered & storage • Characterize Genetic Material – DNA Isolation – Constructing a Library – Automated Sequencing • Metagenomic Analysis

Discoveries • First twelve hours in Sargasso Sea – Tripled the number of known prokaryotes on earth • Six million new genes – 1. 3 million new genes + 50 000 species from single site • Tens of thousands of new protein families • 782 rhodopsin-like photoreceptors – Previously only found in Archaea • Unexpected links between genetics & environment – Different rhodopsin proteins in open ocean vs coast line Better understanding of key biological processes? – New ideas for alternative energy production? – Solutions to deal with climate change?
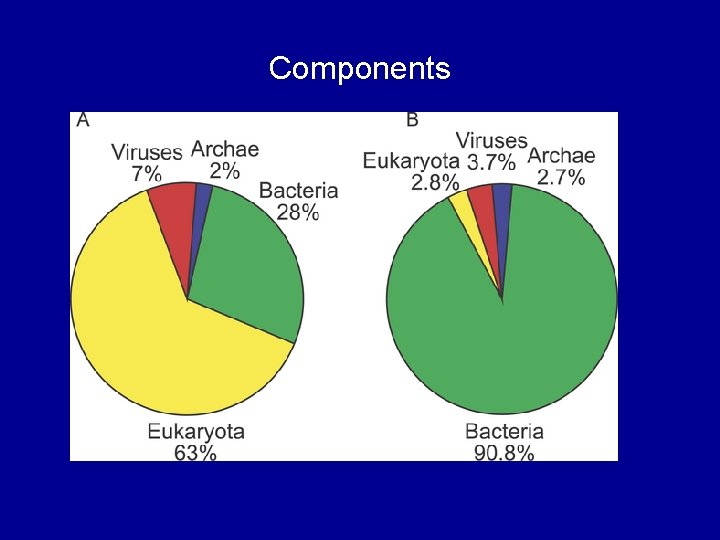
Components

Soil Metagenomics Soil is very diverse - Nutrients - Moisture - p. H - Organic - Oxygen - Temperature - Surface vs. Subsurface Elusive - Poor recovery rate - Nuclease - Cell bias to lysis techniques - Different methods yield dramatically different estimates of diversity and organism number

Soil Diversity Soil is very diverse - Nutrients - Moisture - p. H - Organic - Oxygen - temperature 40 X more diverse than marine The number of prokaryotic species found in a single soil sample exceeds the number of known cultured prokaryotes Estimates:

Soil Requires more Complicated Approach Variability in Extraction Methods Accounting for different extraction methods How much are we still missing?

Other Metagenomic Hot Spots… Wastewater Whale Carcass Human Gut Feces

Applications of Metagenomics “The metagenome provides a potentially inexhaustible genetic resource for biomolecules of potential utility in a variety of industries”

Applications of Metagenomics - Metagenomics offers potential solutions to some of the most complex medical, environmental, agricultural and economic challenges of today - Biotechnological potential of uncultivated bacteria might be accessible by directly cloning DNA sequences retrieved from the environment Discover novel Pathways Information on why certain organisms are unculturable Culture these organisms
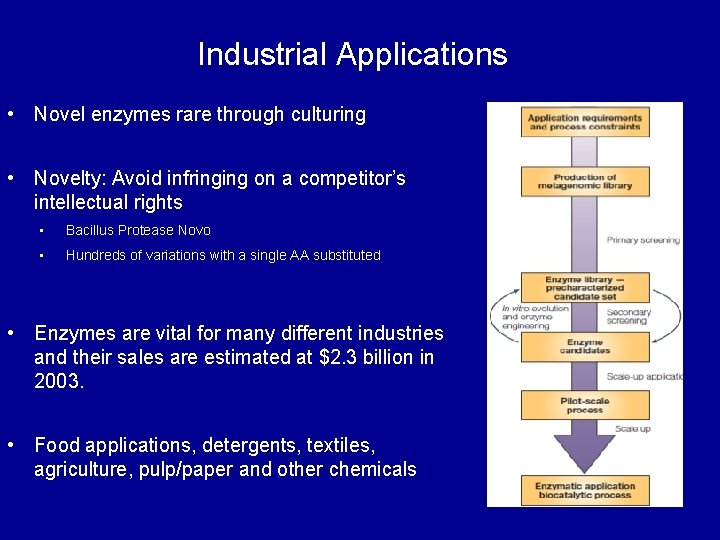
Industrial Applications • Novel enzymes rare through culturing • Novelty: Avoid infringing on a competitor’s intellectual rights • Bacillus Protease Novo • Hundreds of variations with a single AA substituted • Enzymes are vital for many different industries and their sales are estimated at $2. 3 billion in 2003. • Food applications, detergents, textiles, agriculture, pulp/paper and other chemicals
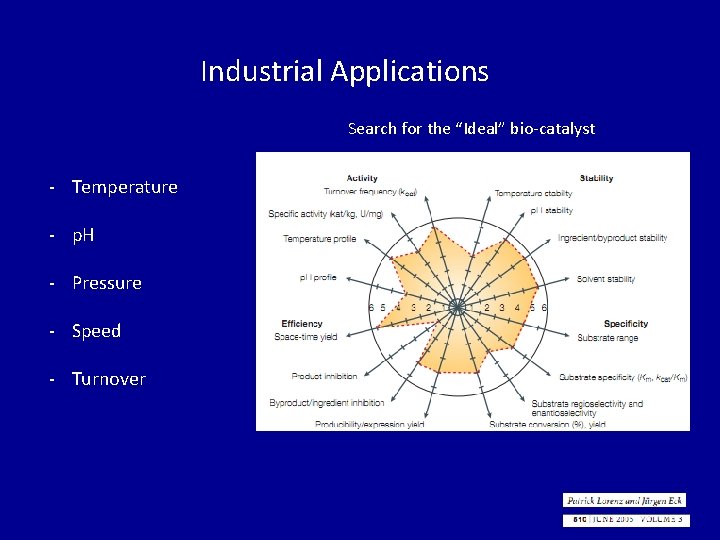
Industrial Applications Search for the “Ideal” bio-catalyst - Temperature - p. H - Pressure - Speed - Turnover

Environmental Remediation Environmental Contamination • Toxic metals • Fossil fuels • Chemicals • Xenobiotics Microorganisms can interact with contaminants • Oxidize • Bind • Transform • Immobilize Metagenomic searches for genes & proteins involved Picking on Alberta: Tar Sands

Agriculture – Detecting diseases in livestock, crops and other products – Soils rich in microbial communities. – Communities very complex, poorly understood and their intimacy with crops means they are of economic importance • nutrient cycling, nitrogen fixation, sequestering metals – Understanding soil composition Enhanced farming

Renewable Energy • Typically derived from biomass sources • Cellulose and other non-edible parts of plants transformed into biofuels • Transform cellulose into usable ethanol, methanol • Ex: Cellulosic ethanol • Produce energy sources such as hydrogen and methane • Capture and store these by-products • Metagenomics approaches for new, efficient ways of producing energy sources

Renewable Energy Searching mircobial communities for biomolecules that can be used as energy Looking in unlikely places Cow Rumen: - Cellulose digestion Methane - Cleaner methane - Metagenomic analysis for compounds involved in this reaction Ex: Bio- Alcohols, Bio-Diseases, Oils, etc….


Human Health – The microbiome: The relationship between the human body and the microbial communities will lead to new methods for diagnosing, treating and preventing diseases – Being used to sequence the microbial communities from ~18 body sites from 250 individuals to determine if changes to the human microbiome can be correlated with human health. – Drugs from microbe-derived compounds: Look for function • metagenomics searches

Metagenomic Approach to Microbiome - Microbiome very influential to human health - What do know about the microbiome? - Metagenomic approach tells us not very much - Comparing microbiome of healty and non. Healthy - Microbiome transplants?

Future Directions • New enzymes, antibiotics, and other reagents identified • More exotic habitats can be intently studied • Can only progress as library technology progresses, technology • Improved bioinformatics will quicken library profile analysis • Investigating ancient DNA remnants • Discoveries such as phylogenic tags (r. RNA genes, etc) • Learning novel pathways will lead to knowledge about the current nonculturable bacteria to then learning to culture these systems including sequencing

Conclusion
- Slides: 63