Metabolic Network Plasticity in Ethyleneproducing Cyanobacteria Algae Biomass

Metabolic Network Plasticity in Ethylene-producing Cyanobacteria Algae Biomass Summit Jianping Yu October 2, 2014 NREL is a national laboratory of the U. S. Department of Energy, Office of Energy Efficiency and Renewable Energy, operated by the Alliance for Sustainable Energy, LLC.
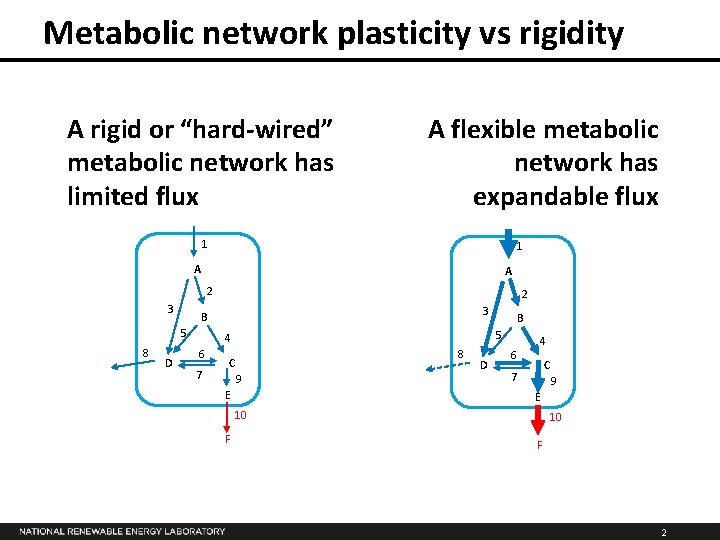
Metabolic network plasticity vs rigidity A rigid or “hard-wired” metabolic network has limited flux A flexible metabolic network has expandable flux 1 1 A A 2 3 D 3 B 5 8 2 5 4 6 7 B 8 C 9 E D 4 6 C 7 9 E 10 F 2
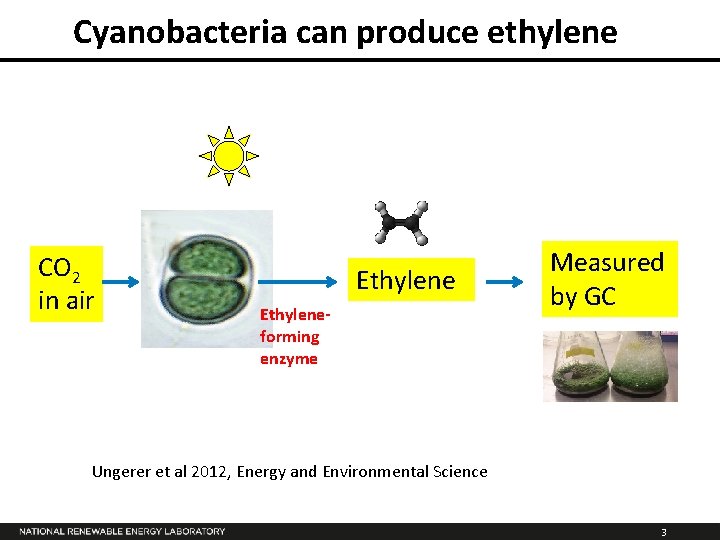
Cyanobacteria can produce ethylene CO 2 in air Ethyleneforming enzyme Measured by GC Ungerer et al 2012, Energy and Environmental Science 3
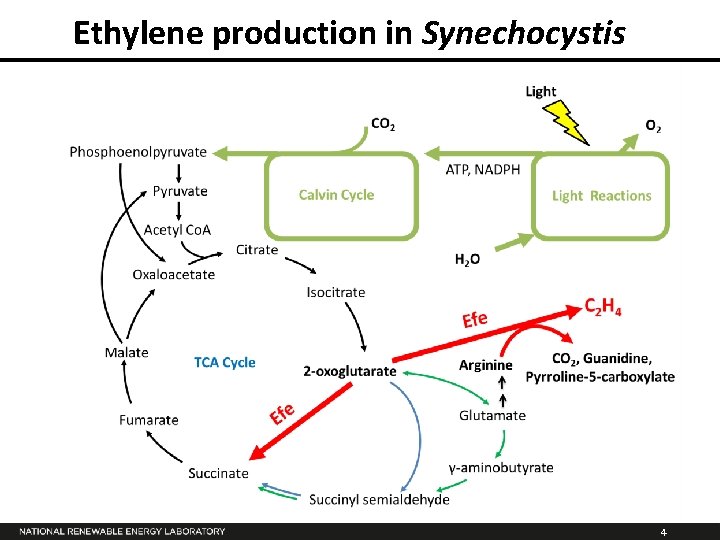
Ethylene production in Synechocystis 4
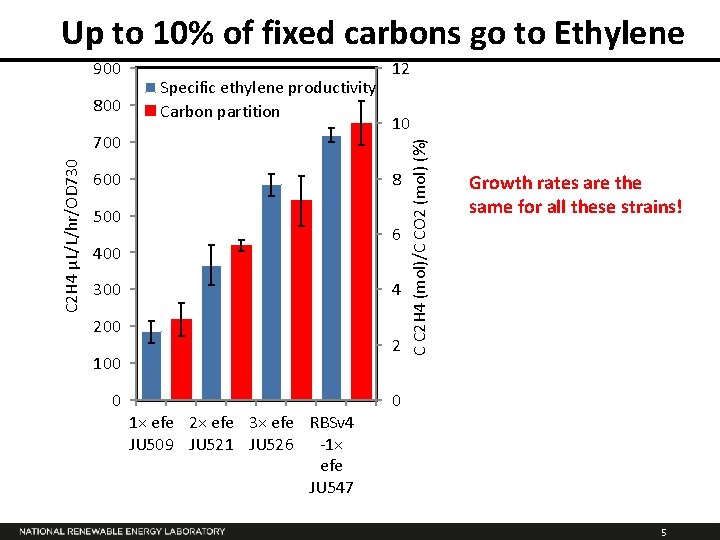
Up to 10% of fixed carbons go to Ethylene 800 Specific ethylene productivity Carbon partition C 2 H 4 µL/L/hr/OD 730 700 600 10 8 500 6 400 300 4 200 2 100 0 12 1× efe 2× efe 3× efe RBSv 4 JU 509 JU 521 JU 526 -1× efe JU 547 C C 2 H 4 (mol)/C CO 2 (mol) (%) 900 Growth rates are the same for all these strains! 0 5
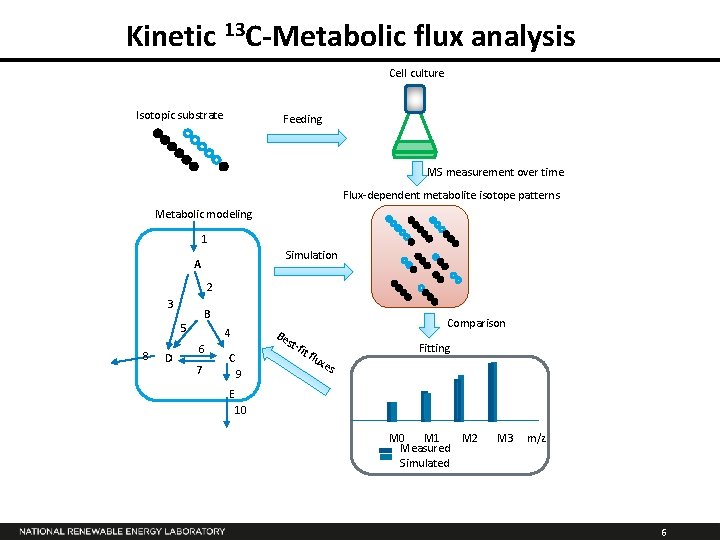
Kinetic 13 C-Metabolic flux analysis Cell culture Isotopic substrate Feeding MS measurement over time Flux-dependent metabolite isotope patterns Metabolic modeling 1 Simulation A 2 3 B 5 8 D 4 6 7 Comparison Be st- C 9 fit flu xes Fitting E 10 M 1 Measured Simulated M 2 M 3 m/z 6

Kinetic labeling Strains: Synechocystis OD 730 ~1. 5 Medium: liquid BG 11 20 ml Light Intensity: 80 µE Culture conditions: shaking at 200 rpm at 30 o. C in 5% CO 2 growth chamber Cell Harvest: filtering with 0. 8µM nylon membrane Quenching: -80 o. C methanol Extraction: 50% cold methanol Post processing: evaporated with speed vacuum and re-dissolve in 200 µl 50% cold methanol; the supernatant is ready for LC-MS analysis 7
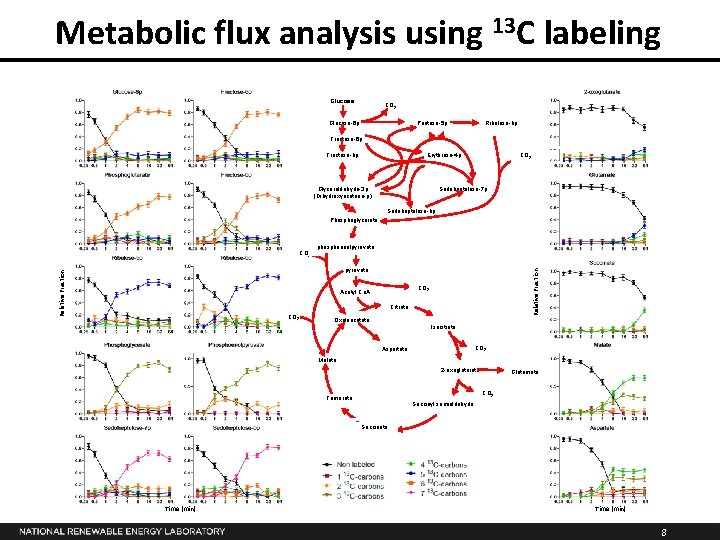
Metabolic flux analysis using 13 C labeling Glucose CO 2 Pentose-5 p Glucose-6 p Ribulose-bp Fructose-6 p Fructose-bp Erythrose-4 p Glyceraldehyde-3 p (Dihydroxyacetone-p) CO 2 Sedoheptulose-7 p Sedoheptulose-bp Phosphoglycerate CO 2 phosphoenolpyruvate Relative Fraction pyruvate CO 2 Acetyl Co. A Citrate CO 2 Oxaloacetate Isocitrate CO 2 Aspartate Malate 2 -oxoglutarate Glutamate CO 2 Fumarate Succinyl semialdehyde Succinate Time (min) 8
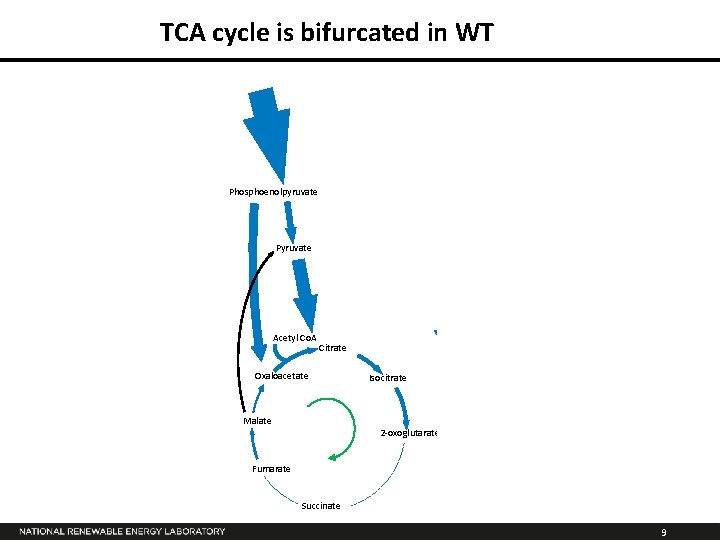
TCA cycle is bifurcated in WT Phosphoenolpyruvate Pyruvate Acetyl Co. A Citrate Oxaloacetate Isocitrate Oxaloacetate Ethylene Isocitrate Malate 2 -oxoglutarate Fumarate Succinate 9
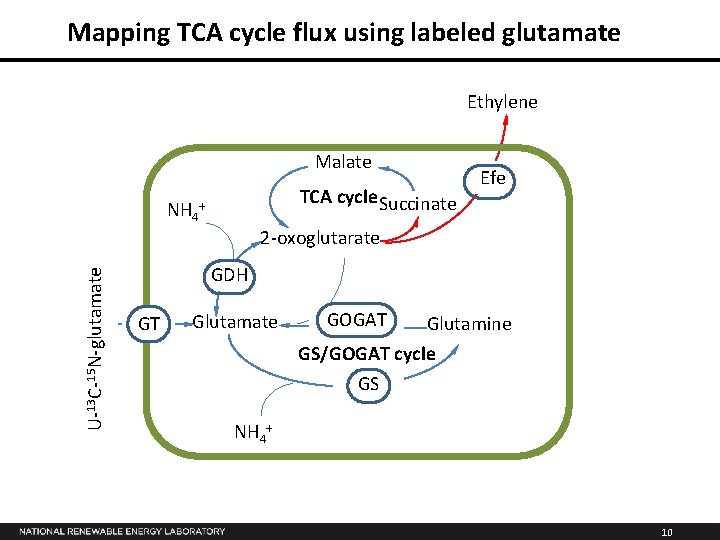
Mapping TCA cycle flux using labeled glutamate Ethylene Malate NH 4 TCA cycle Succinate + Efe U-13 C-15 N-glutamate 2 -oxoglutarate GDH GT Glutamate GOGAT Glutamine GS/GOGAT cycle GS NH 4+ 10
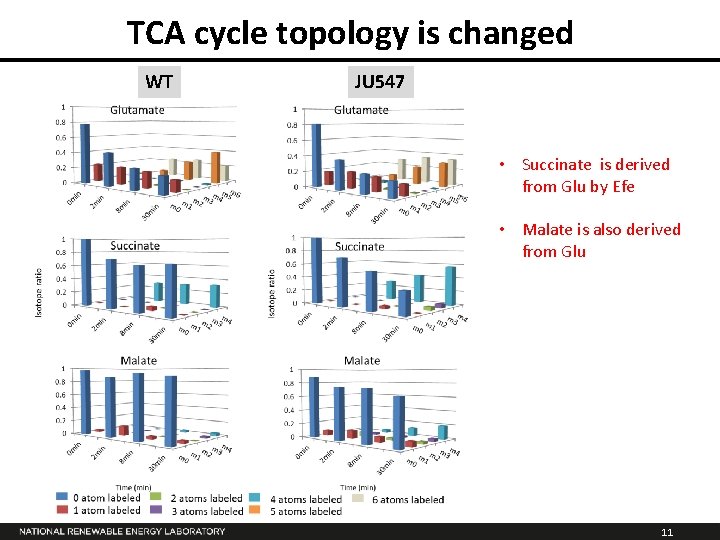
TCA cycle topology is changed WT JU 547 • Succinate is derived from Glu by Efe • Malate is also derived from Glu 11
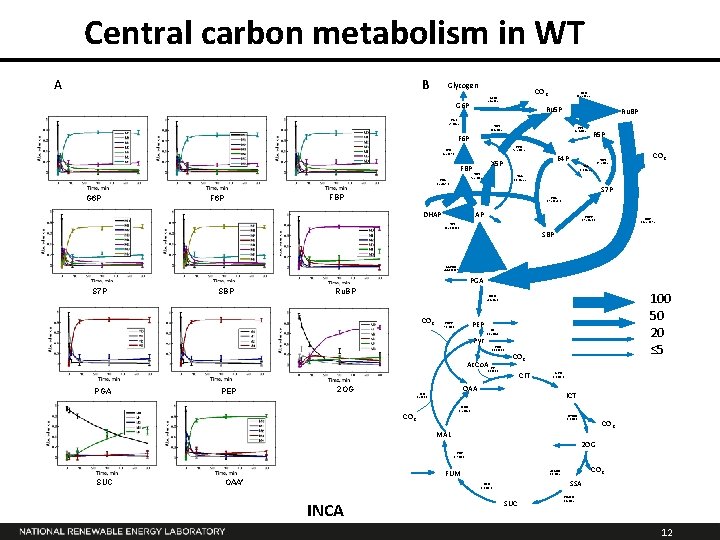
Central carbon metabolism in WT A B Glycogen CO 2 G 6 PD 24. 2± 5. 5 G 6 P PRK 135. 2 ± 5. 5 Ru 5 P PGI 27. 1± 5. 5 Ru. BP TKT 38. 4 ± 0. 2 PPI 35. 4 ± 0. 2 R 5 P F 6 P PPE 75. 5 ± 0. 5 PFK 65. 5± 29. 3 TKT 75. 5 ± 0. 5 FBA 65. 5± 29. 3 G 6 P E 4 P X 5 P FBP TAL 0. 0 ± 25. 5 CO 2 TAL 0. 0 ± 25. 5 S 7 P FBP F 6 P TKT 37. 2 ± 0. 2 SBA 37. 2 ± 26. 8 DHAP GAP SBPS 37. 2 ± 26. 8 RBC 135. 2 ± 17. 2 TPI 102. 6 ± 11. 3 SBP GAPDH 244. 0 ± 10. 9 PGA S 7 P SBP Ru. BP CO 2 PEPC 7. 8 ± 0. 3 PK 13. 2 ± 4. 1 Pyr PDH 11. 8 ± 0. 3 Ac. Co. A CO 2 CS 3. 0 ± 0. 3 PGA PEP 100 50 20 ≤ 5 ENO 23. 4 ± 2. 1 2 OG CIT ACO 3. 0 ± 0. 3 OAA ME 1. 5 ± 0. 3 ICT MDH 0. 2 ± 4. 6 CO 2 ICTDH 3. 0 ± 0. 3 CO 2 MAL 2 OG FUS 1. 7 ± 0. 3 SUC FUM OAA* SSA SDH 0. 0 ± 0. 3 INCA CO 2 2 OGDH 0. 0 ± 1. 5 SUC SSADH -0. 4 ± 0. 2 12
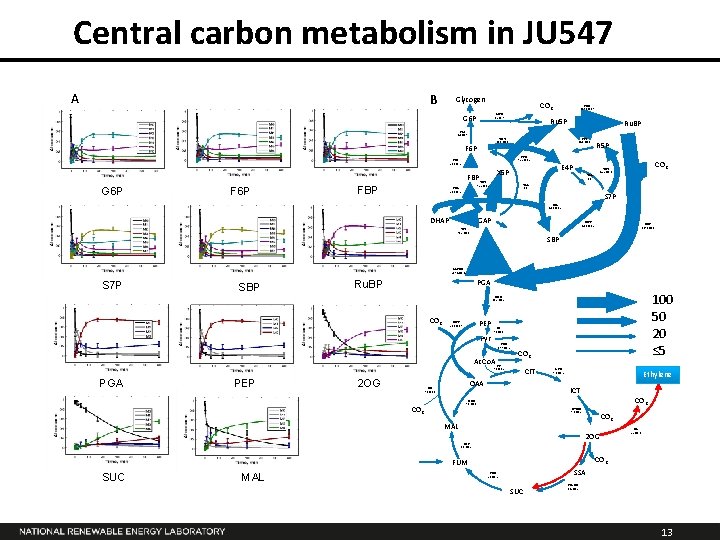
Central carbon metabolism in JU 547 A B Glycogen CO 2 G 6 PD 1. 5± 0. 7 G 6 P PGI 4. 1± 0. 7 R 5 P TKT 92. 2 ± 0. 2 FBA 50. 0± 1. 2 E 4 P X 5 P FBP Ru. BP PPI 44. 0 ± 0. 1 PPE 92. 2 ± 0. 2 PFK 50. 0± 1. 2 F 6 P Ru 5 P TKT 46. 6 ± 0. 1 F 6 P G 6 P PRK 137. 6 ± 0. 7 CO 2 TKT 45. 5 ± 0. 1 TAL 0. 6 S 7 P SBA 46. 1 ± 0. 2 DHAP GAP SBPS 46. 1 ± 0. 2 TPI 96. 2 ± 0. 6 RBC 137. 6± 0. 6 SBP GAPDH 239. 4 ± 1. 4 S 7 P SBP PGA Ru. BP 100 50 20 ≤ 5 ENO 33. 2 ± 0. 2 CO 2 PEPC 21. 8 ± 0. 7 Pyr PK 9. 1 ± 1. 3 PDH 17. 7 ± 0. 2 Ac. Co. A CO 2 CS 9. 8 ± 0. 2 PGA PEP 2 OG CIT OAA ME 7. 6 ± 1. 3 ACO 9. 8 ± 0. 2 Ethylene ICT CO 2 MDH 7. 6 ± 1. 3 CO 2 ICTDH 9. 8 ± 0. 2 MAL CO 2 2 OG Efe 2. 5 ± 0. 0 FUS 3. 6 ± 0. 2 CO 2 FUM SUC MAL SSA SDH 2. 1 ± 0. 2 SUC SSADH -0. 4 ± 0. 2 13
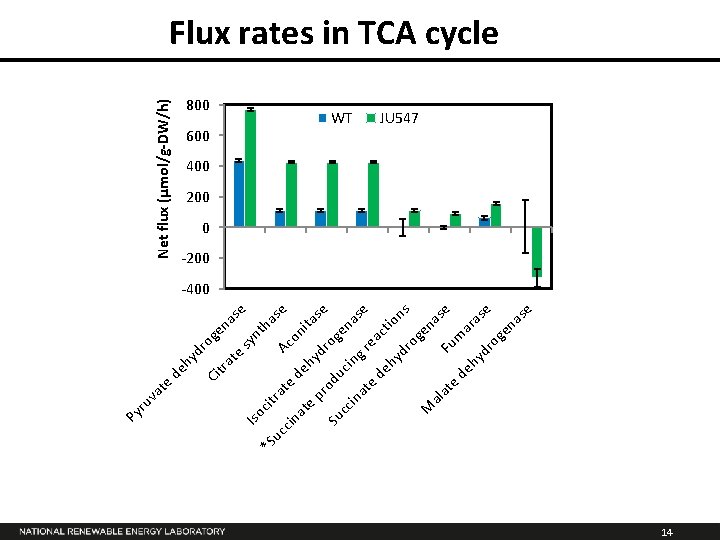
hy d de ro Ci tra gen as te e sy Iso nt cit ha *S ra uc Ac se te cin on de at ita hy e se pr dr o o Su du ge cc c i ng nas in at e re e ac de tio hy ns dr og en M as al at F um e e de hy ara se dr og en as e te va ru Py Net flux (µmol/g-DW/h) Flux rates in TCA cycle 800 600 WT JU 547 400 200 0 -200 -400 14
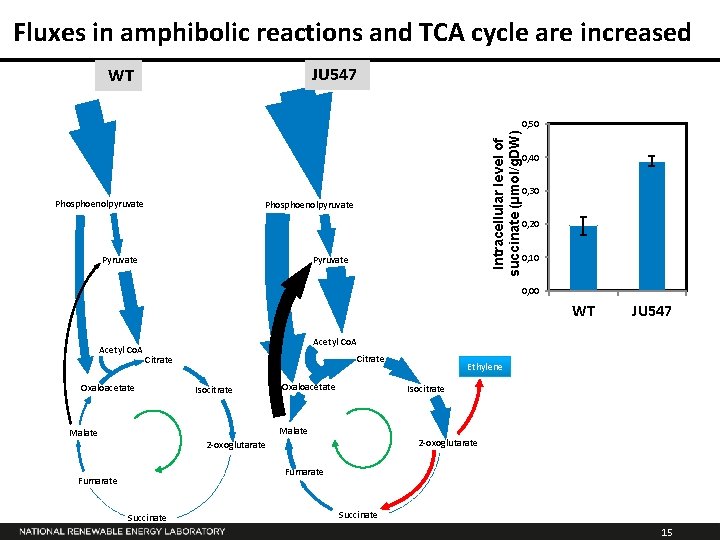
Fluxes in amphibolic reactions and TCA cycle are increased JU 547 WT Intracellular level of succinate (µmol/g. DW) 0, 50 0, 40 0, 30 Phosphoenolpyruvate 0, 20 Pyruvate 0, 10 Pyruvate 0, 00 WT Acetyl Co. A JU 547 Acetyl Co. A Citrate Oxaloacetate Isocitrate Oxaloacetate Ethylene Isocitrate Malate 2 -oxoglutarate Fumarate Succinate 15
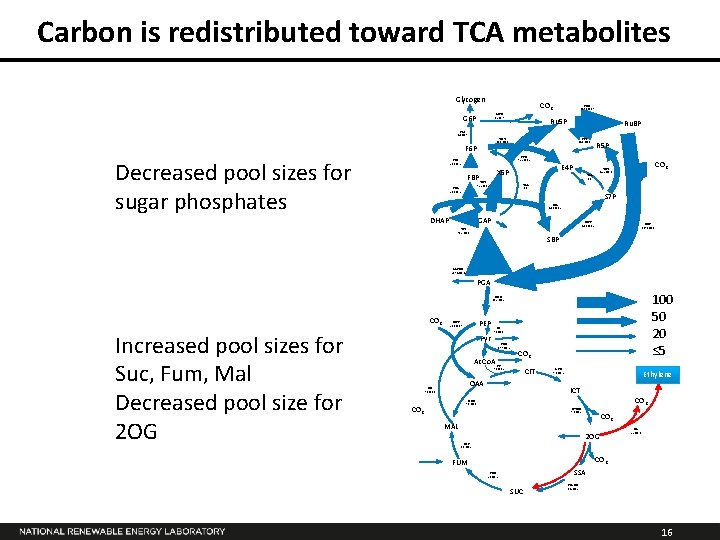
Carbon is redistributed toward TCA metabolites Glycogen CO 2 G 6 PD 1. 5± 0. 7 G 6 P PGI 4. 1± 0. 7 Ru 5 P Ru. BP TKT 46. 6 ± 0. 1 F 6 P Decreased pool sizes for sugar phosphates PRK 137. 6 ± 0. 7 PPI 44. 0 ± 0. 1 R 5 P PPE 92. 2 ± 0. 2 PFK 50. 0± 1. 2 TKT 92. 2 ± 0. 2 FBA 50. 0± 1. 2 E 4 P X 5 P FBP CO 2 TKT 45. 5 ± 0. 1 TAL 0. 6 S 7 P SBA 46. 1 ± 0. 2 DHAP GAP SBPS 46. 1 ± 0. 2 TPI 96. 2 ± 0. 6 RBC 137. 6± 0. 6 SBP GAPDH 239. 4 ± 1. 4 PGA 100 50 20 ≤ 5 ENO 33. 2 ± 0. 2 CO 2 Increased pool sizes for Suc, Fum, Mal Decreased pool size for 2 OG PEPC 21. 8 ± 0. 7 Pyr PK 9. 1 ± 1. 3 PDH 17. 7 ± 0. 2 Ac. Co. A CO 2 CS 9. 8 ± 0. 2 CIT OAA ME 7. 6 ± 1. 3 ACO 9. 8 ± 0. 2 Ethylene ICT CO 2 MDH 7. 6 ± 1. 3 CO 2 ICTDH 9. 8 ± 0. 2 MAL CO 2 2 OG Efe 2. 5 ± 0. 0 FUS 3. 6 ± 0. 2 CO 2 FUM SSA SDH 2. 1 ± 0. 2 SUC SSADH -0. 4 ± 0. 2 16
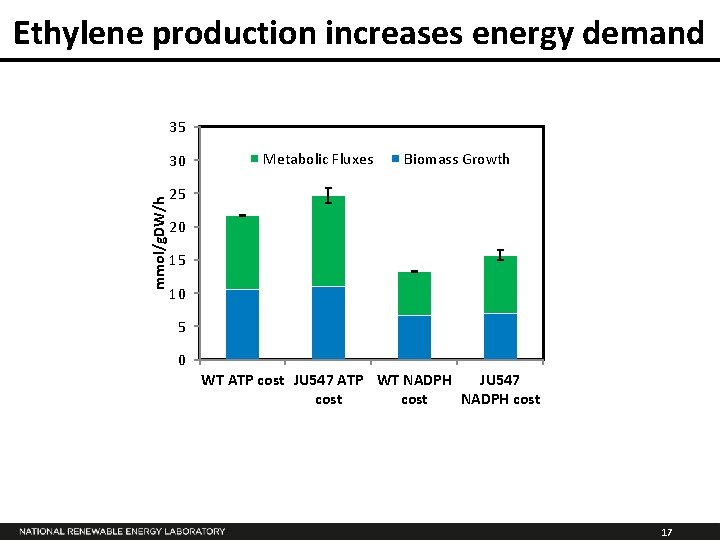
Ethylene production increases energy demand 35 mmol/g. DW/h 30 Metabolic Fluxes Biomass Growth 25 20 15 10 5 0 WT ATP cost JU 547 ATP WT NADPH JU 547 cost NADPH cost 17
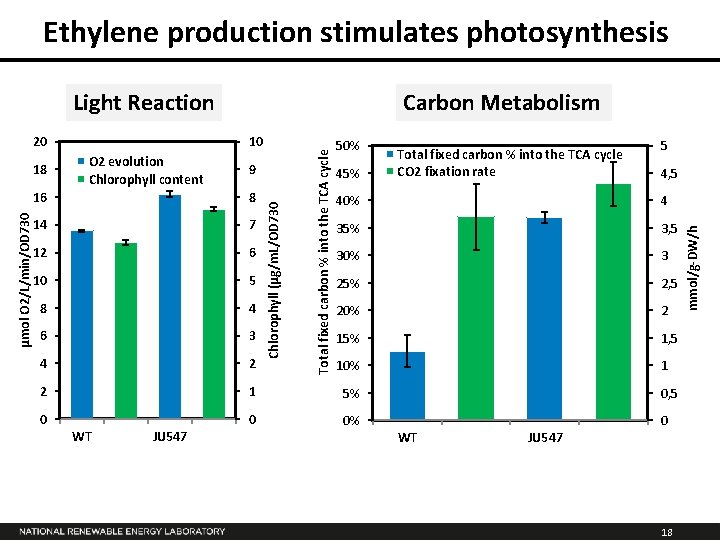
Ethylene production stimulates photosynthesis O 2 evolution Chlorophyll content 9 50% 45% Total fixed carbon % into the TCA cycle CO 2 fixation rate 5 4, 5 16 8 14 7 12 6 10 5 8 4 6 3 4 2 2 1 5% 0, 5 0 0 0% 0 WT JU 547 Chlorophyll (µg/m. L/OD 730 µmol O 2/L/min/OD 730 18 10 40% 4 35% 3, 5 30% 3 25% 2, 5 20% 2 15% 1, 5 10% 1 WT JU 547 18 mmol/g-DW/h 20 Carbon Metabolism Total fixed carbon % into the TCA cycle Light Reaction

Conclusions Metabolic network in a cyanobacterium is very flexible • TCA cycle topology can change in response to an engineered ethylene pathway • Amphibolic reactions and TCA cycle fluxes can be increased to accommodate this engineered pathway • Carbon can be pulled from elsewhere to accommodate an engineered pathway • Photosynthesis can run faster to meet the increased demand 19

Conclusions Metabolic network in a cyanobacterium is very flexible • TCA cycle topology can change in response to an engineered ethylene pathway • Amphibolic reactions and TCA cycle fluxes can be increased to accommodate this engineered pathway • Carbon can be pulled from elsewhere to accommodate an engineered pathway • Photosynthesis can run faster to meet the increased demand Source-sink relationship in photosynthesis • Sink expansion can stimulate photosynthesis, so that loss of carbon from an engineered pathway is fully compensated without slowing down cell growth. 20

Conclusions Metabolic network in a cyanobacterium is very flexible • TCA cycle topology can change in response to an engineered ethylene pathway • Amphibolic reactions and TCA cycle fluxes can be increased to accommodate this engineered pathway • Carbon can be pulled from elsewhere to accommodate an engineered pathway • Photosynthesis can run faster to meet the increased demand Source-sink relationship in photosynthesis • Sink expansion can stimulate photosynthesis, so that loss of carbon from an engineered pathway is fully compensated without slowing down cell growth. • How much is the unrealized potential in photosynthesis? • Can we find the trigger to unleash that potential and increase algal productivity? 21

Acknowledgements Wei Xiong Justin Ungerer Pin-Ching Maness John Morgan Funding from DOE BETO DOE BER NREL 22
- Slides: 22