MEMBRANE PROTEINS Anna Adell Mariona Bartrol Marina Bras

MEMBRANE PROTEINS Anna Adell, Mariona Bartrolí, Marina Brasó i Abel Eraso
Index 1. Introduction • What is cell membrane? • Integral Proteins • Crystallization 2. Transmembrane Proteins • β - Barrels • α-Helical 3. GPCRs • What are G protein – coupled receptors? • Classification • Sequence conservation • Class A • SCOP • Chronology • Alignments • β 2 -Adrenergic Receptor • Extracellular Loop 2 (ECL 2) • Ligand binding site • Ionic Lock • Activation 4. 5. Conclusions Bibliography
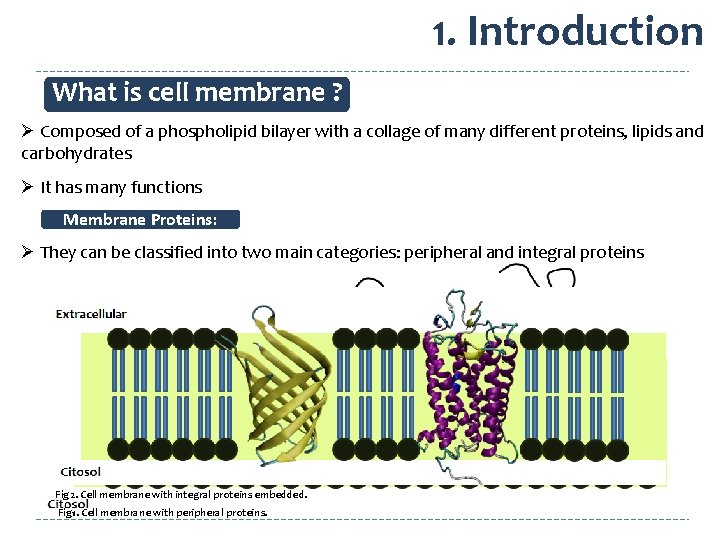
1. Introduction What is cell membrane ? Ø Composed of a phospholipid bilayer with a collage of many different proteins, lipids and carbohydrates Ø It has many functions Membrane Proteins: Ø They can be classified into two main categories: peripheral and integral proteins Fig 2. Cell membrane with integral proteins embedded. Fig 1. Cell membrane with peripheral proteins.
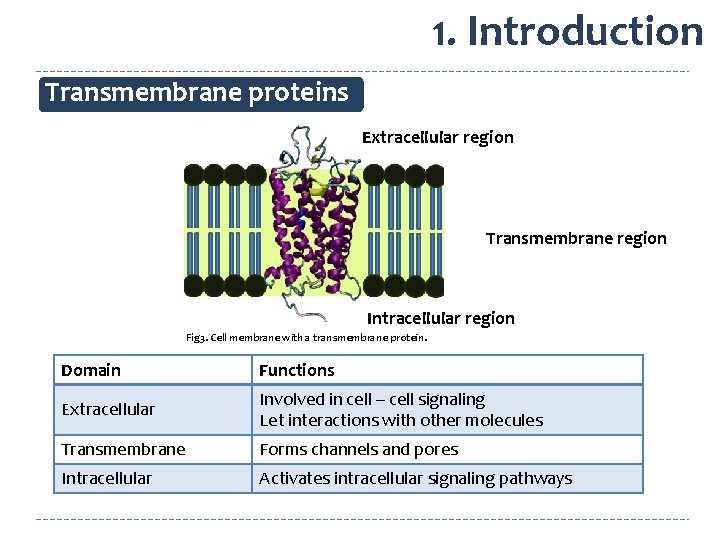
1. Introduction Transmembrane proteins Extracellular region Transmembrane region Intracellular region Fig 3. Cell membrane with a transmembrane protein. Domain Functions Extracellular Involved in cell – cell signaling Let interactions with other molecules Transmembrane Forms channels and pores Intracellular Activates intracellular signaling pathways

1. Introduction CRYSTALLIZATION. Why is it so complicated? Ø Difficulties in handling and crystallization of integral membrane proteins reside in the amphiphatic nature of their surface ØThey have hydrophobic transmembrane regions and two hydrophillic regions, one of each side of the membrane ØAs a result, membrane proteins are not soluble in aqueous buffers or in organic solvents Membrane protein structural biology: The most challenging targets in structural biology
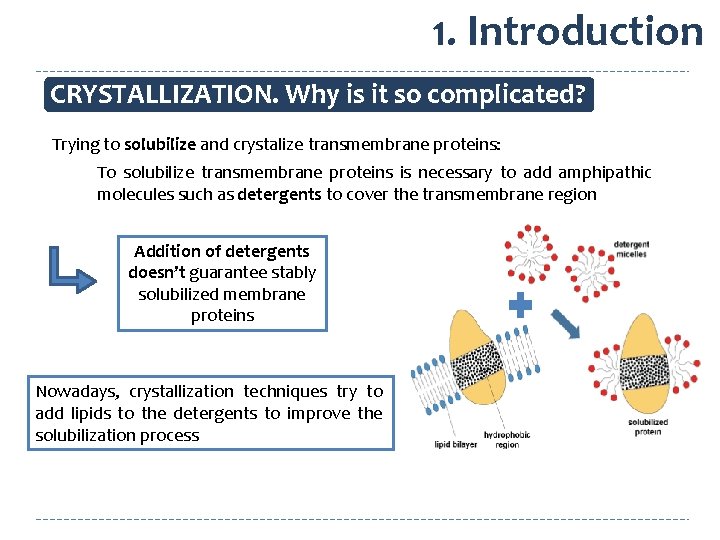
1. Introduction CRYSTALLIZATION. Why is it so complicated? Trying to solubilize and crystalize transmembrane proteins: To solubilize transmembrane proteins is necessary to add amphipathic molecules such as detergents to cover the transmembrane region Addition of detergents doesn’t guarantee stably solubilized membrane proteins Nowadays, crystallization techniques try to add lipids to the detergents to improve the solubilization process
Index 1. Introduction • What is cell membrane? • Integral Proteins • Crystallization 2. Transmembrane Proteins • β - Barrels • α-Helical 3. GPCRs • What are G protein – coupled receptors? • Classification • Sequence conservation • Class A • SCOP • Chronology • Alignments • β 2 -Adrenergic Receptor • Extracellular Loop 2 (ECL 2) • Ligand binding site • Ionic Lock • Activation 4. 5. Conclusions Bibliography
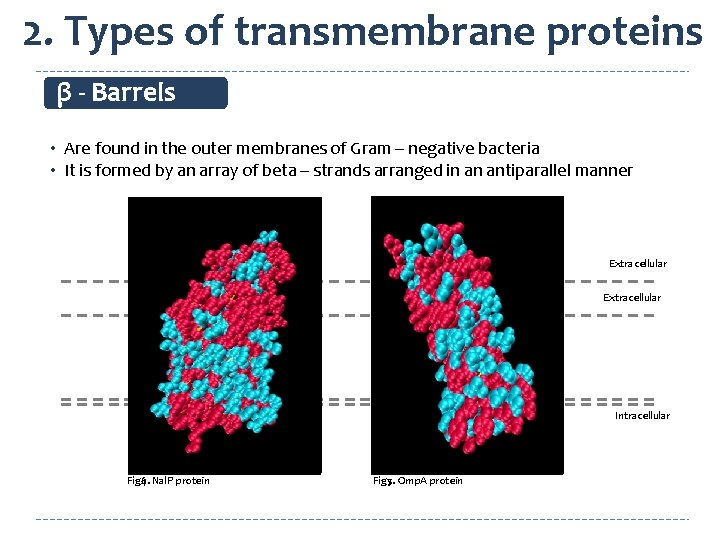
2. Types of transmembrane proteins β - Barrels • Are found in the outer membranes of Gram – negative bacteria • It is formed by an array of beta – strands arranged in an antiparallel manner Extracellular Intracellular Fig 4. Nal. P protein Fig 6. Fig 5. Omp. A protein Fig 7.
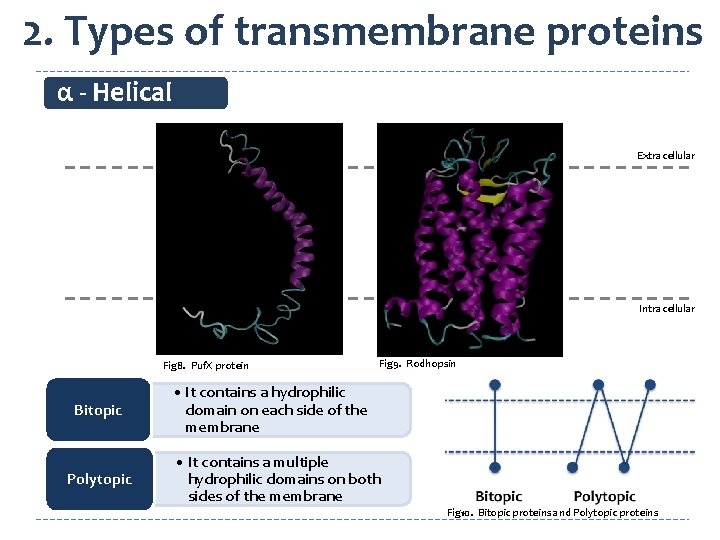
2. Types of transmembrane proteins α - Helical Extracellular Intracellular Fig 8. Puf. X protein Bitopic Polytopic Fig 9. Rodhopsin • It contains a hydrophilic domain on each side of the membrane • It contains a multiple hydrophilic domains on both sides of the membrane Fig 10. Bitopic proteins and Polytopic proteins
Index 1. Introduction • What is cell membrane? • Integral Proteins • Crystallization 2. Transmembrane Proteins • β - Barrels • α-Helical 3. GPCRs • What are G protein – coupled receptors? • Classification • Sequence conservation • Class A • SCOP • Chronology • Alignments • β 2 -Adrenergic Receptor • Extracellular Loop 2 (ECL 2) • Ligand binding site • Ionic Lock • Activation 4. 5. Conclusions Bibliography
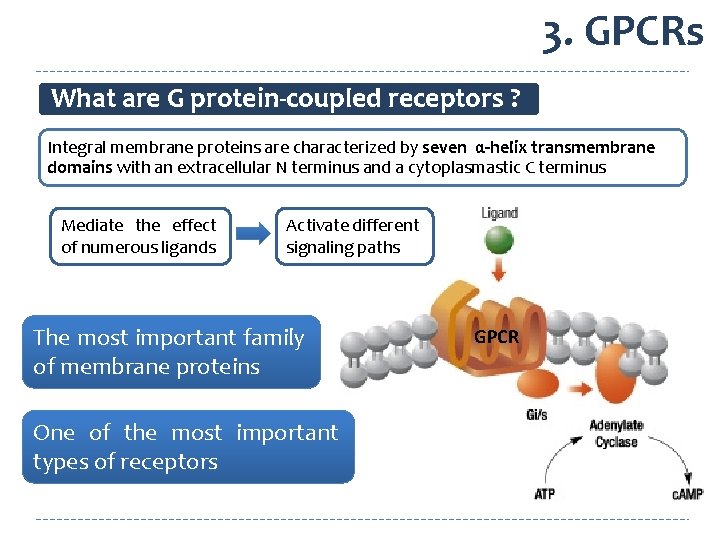
3. GPCRs What are G protein-coupled receptors ? Integral membrane proteins are characterized by seven α-helix transmembrane domains with an extracellular N terminus and a cytoplasmastic C terminus Mediate the effect of numerous ligands Activate different signaling paths The most important family of membrane proteins One of the most important types of receptors GPCR
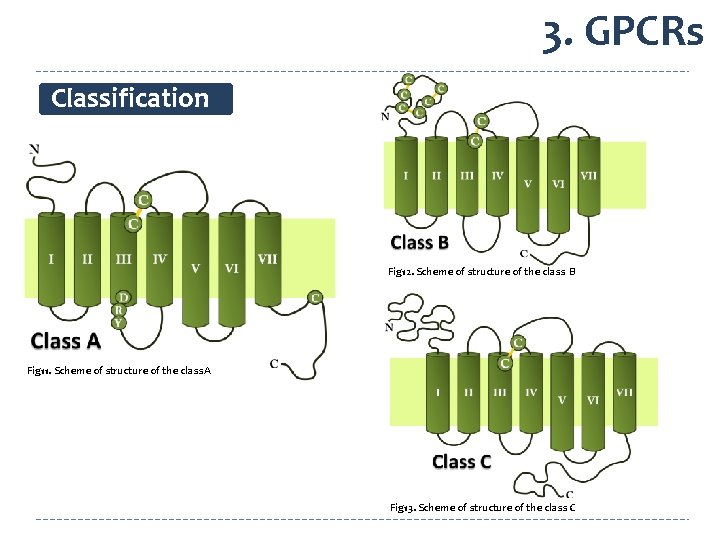
3. GPCRs Classification Fig 12. Scheme of structure of the class B Fig 11. Scheme of structure of the class A Fig 13. Scheme of structure of the class C
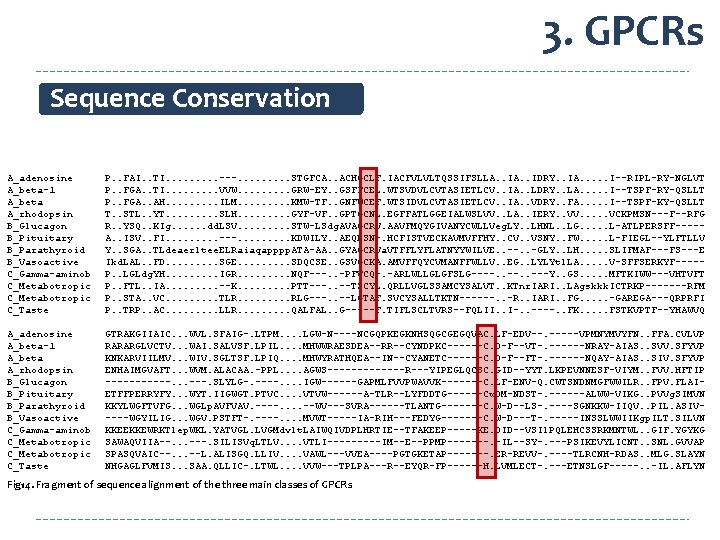
3. GPCRs Sequence Conservation A_adenosine A_beta-1 A_beta A_rhodopsin B_Glucagon B_Pituitary B_Parathyroid B_Vasoactive C_Gamma-aminob C_Metabotropic C_Taste P. . FAI. . TI. . ---. . STGFCA. . ACHGCLF. IACFVLVLTQSSIFSLLA. . IDRY. . IA. . . I--RIPL-RY-NGLVT P. . FGA. . TI. . VVW. . GRW-EY. . GSFFCEL. WTSVDVLCVTASIETLCV. . IA. . LDRY. . LA. . . I--TSPF-RY-QSLLT P. . FGA. . AH. . ILM. . KMW-TF. . GNFWCEF. WTSIDVLCVTASIETLCV. . IA. . VDRY. . FA. . . I--TSPF-KY-QSLLT T. . STL. . YT. . SLH. . GYF-VF. . GPTGCNL. EGFFATLGGEIALWSLVV. . LA. . IERY. . VV. . . VCKPMSN---F--RFG R. . YSQ. . KIg. . . dd. LSV. . STW-LSdg. AVAGCRV. AAVFMQYGIVANYCWLLVeg. LY. . LHNL. . LG. . . L-ATLPERSFF----A. . ISV. . FI. . ---. . KDWILY. . AEQDSN-. HCFISTVECKAVMVFFHY. . CV. . VSNY. . FW. . . L-FIEGL--YLFTLLV Y. . SGA. . TLdeaerltee. ELRaiaqapppp. ATA-AA. . GYAGCRVa. VTFFLYFLATNYYWILVE. . --. . -GLY. . LH. . . SLIFMAF---FS---E Ikd. LAL. . FD. . SGE. . SDQCSE. . GSVGCKA. AMVFFQYCVMANFFWLLV. . EG. . LYLYtl. LA. . . V-SFFSERKYF----P. . LGLdg. YH. . IGR. . NQF---. . -PFVCQ-. -ARLWLLGLGFSLG----. . ---Y. . GS. . . MFTKIWW---VHTVFT P. . FTL. . IA. . --K. . PTT---. . --TSCYL. QRLLVGLSSAMCYSALVT. . KTnr. IARI. . LAgskkk. ICTRKP-------RFM P. . STA. . VC. . TLR. . RLG---. . --LGTAF. SVCYSALLTKTN------. . -R. . IARI. . FG. . . -GAREGA---QRPRFI P. . TRP. . AC. . LLR. . QALFAL. . G-----F. TIFLSCLTVRS--FQLII. . I-. . ----. . FK. . . FSTKVPTF--YHAWVQ A_adenosine A_beta-1 A_beta A_rhodopsin B_Glucagon B_Pituitary B_Parathyroid B_Vasoactive C_Gamma-aminob C_Metabotropic C_Taste GTRAKGIIAIC. . . WVL. SFAIG-. LTPM. . LGW-N----NCGQPKEGKNHSQGCGEGQVAC. LF-EDV--. -----VPMNYMVYFN. . FFA. CVLVP RARARGLVCTV. . . WAI. SALVSF. LPIL. . MHWWRAESDEA--RR--CYNDPKC------C. D-F--VT-. ------NRAY-AIAS. . SVV. SFYVP KNKARVIILMV. . . WIV. SGLTSF. LPIQ. . MHWYRATHQEA--IN--CYANETC------C. D-F--FT-. ------NQAY-AIAS. . SIV. SFYVP ENHAIMGVAFT. . . WVM. ALACAA. -PPL. . AGWS-------R---YIPEGLQCSC. GID--YYT. LKPEVNNESF-VIYM. . FVV. HFTIP ------. . . ---. SLYLG-. ----. . IGW------GAPMLFVVPWAVVK-------C. LF-ENV-Q. CWTSNDNMGFWWILR. . FPV. FLAIETFFPERRYFY. . . WYT. IIGWGT. PTVC. . VTVW------A-TLR--LYFDDTG------Cw. DM-NDST-. ------ALWW-VIKG. . PVVg. SIMVN KKYLWGFTVFG. . . WGLp. AVFVAV. ----. . --WV---SVRA------TLANTG-------C. W-D--LS-. ----SGNKKW-IIQV. . PIL. ASIV----WGYILIG. . . WGV. PSTFT-. ----. . MVWT-----IA-RIH---FEDYG-------C. W-D---T-. -----INSSLWWIIKgp. ILT. SILVN KKEEKKEWRKTlep. WKL. YATVGL. LVGMdvlt. LAIWQIVDPLHRTIE--TFAKEEP-----KE. DID--VSIl. PQLEHCSSRKMNTWL. . GIF. YGYKG SAWAQVIIA--. . . ---. SILISVq. LTLV. . VTLI-----IM--E--PPMP-------. -IL--SY-. ---PSIKEVYLICNT. . SNL. GVVAP SPASQVAIC--. . . --L. ALISGQ. LLIV. . VAWL---VVEA----PGTGKETAP-------. ER-REVV-. ----TLRCNH-RDAS. . MLG. SLAYN NHGAGLFVMIS. . . SAA. QLLIC-. LTWL. . VVW---TPLPA---R--EYQR-FP------H. LVMLECT-. ---ETNSLGF-----. . -IL. AFLYN Fig 14. Fragment of sequence alignment of the three main classes of GPCRs
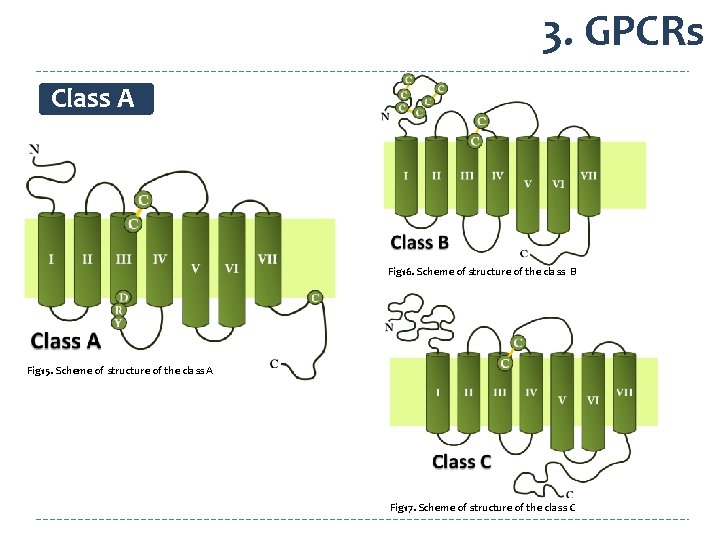
3. GPCRs Class A Fig 16. Scheme of structure of the class B Fig 15. Scheme of structure of the class A Fig 17. Scheme of structure of the class C

3. GPCRs Class A. SCOP Fig 18. Class A classification from SCOP (Structural Classification of Proteins)
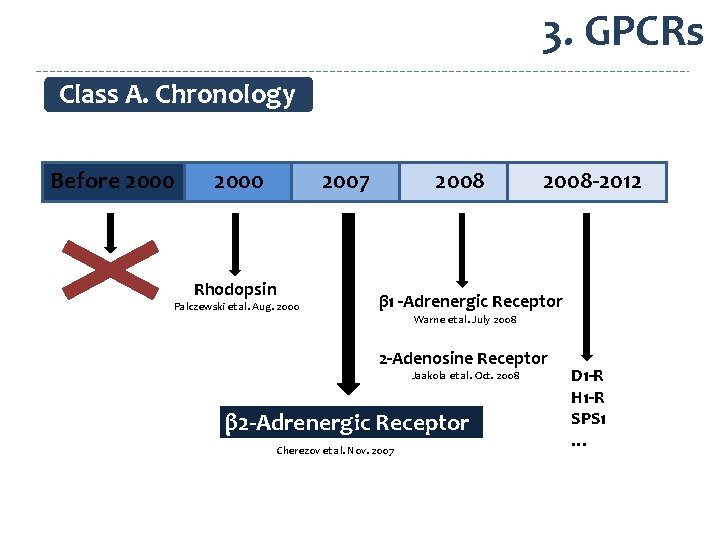
3. GPCRs Class A. Chronology Before 2000 Rhodopsin 2007 Palczewski et al. Aug. 2000 2008 -2012 β 1 -Adrenergic Receptor Warne et al. July 2008 2 -Adenosine Receptor Jaakola et al. Oct. 2008 β 2 -Adrenergic Receptor Cherezov et al. Nov. 2007 D 1 -R H 1 -R SPS 1 …

3. GPCRs adenosine AVLAILGNVLVCWAVWL. . NSNLQNV. . . TNYFVVSLAAADIAVGVLAIP. FAITI---STGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAI--RIPL-RY-NGLV. TGTR. A. KGIIAICWVLSFAIG-LTPMLGW-N---beta-1 VLLIVAGNVLVIVAIAK. . TPRLQTL. . . TNLFIMSLASADLVMGLLVVP. FGATIVVWGRW-EYGSFFCELWTSVDVLCVTASIETLCVIALDRYLAI--TSPF-RY-QSLL. TRAR. A. RGLVCTVWAISALVSFLPILMHWWRAESD beta VLAIVFGNVLVITAIAK. . FERLQTV. . . TNYFITSLACADLVMGLAVVP. FGAAHILMKMW-TFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAI--TSPF-KY-QSLL. TKNK. A. RVIILMVWIVSGLTSFLPIQMHWYRATHQ chemokine_r FLTGIVGNGLVILVMGY. . QKKLRSM. . . TDKYRLHLSVADLLFVIT-LP. FWAVDAVANWY--FGNFLCKAVHVIYTVNLYSSVLILAFISLDRYLAI-VHATN-S-Q--RP. RKLL. Ae. KVVYVGVWIPALLLT-IPDFI--F-ANVS D 2 IAVIVFGNVLVCMAVSR. . EKALQTT. . . TNYLIVSLAVADLLVATLVMP. WVVYLEVVGEW-KFSRIHCDIFVTLDVMMCTASILNLCAISIDRYTAV--AMPML--YNTRYs. SKRR. V. TVMISIVWVLSFTISC-PLLFGL-N---galanin FLLGTVGNGLVLAVLLQpg. PSAWQEPgst. TDLFILNLAVADLCFILCCVP. FQATIYTLDAW-LFGALVCKAVHLLIYLTMYASSFTLAAVSVDRYLAV--RHPL-R-SRALR. TPRN. A. RAAVGLVWLLAALFSA--PYLSYY----histamine ILITVAGNVVVCLAVGL. . NRRLRNL. . . TNCFIVSLAITDLLLGLLVLP. FSAIYQLSCKW-SFGKVFCNIYTSLDVMLCTASILNLFMISLDRYCAV--MDPL-RY-PVLV. TPVR. V. AISLVLIWVISITLSFLSIHLGW-NSRNE melatonin TAVDVVGNLLVILSVLR. . NRKLRNA. . . GNLFLVSLALADLVVAFYPYP. LILVAIFYDGW-ALGEEHCKASAFVMGLSVIGSVFNITAIAINRYCYICHSMAYHRIYR---. -RWH. T. PLHICLIWLLTV-VALLPNFFVGSL---muscarinic SLVTIIGNILVMVSIKV. . NRHLQTV. . . NNYFLFSLACADLIIGVFSMNl. YTLYT-VIGYW-PLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCV--TKPL-TY-PVKR. TTKM. A. GMMIAAAWVLSFILWA-PAILFW-----delta CAVGLLGNVLVMFGIVR. . YTKMKTA. . . TNIYIFNLALAD-ALATSTLP. FQSAKYLMETW-PFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALD-FR---. TPAK. A. KLINICIWVLASGVGV-PIMVMAV--TRP kappa FVVGLVGNSLVMFVIIR. . YTKMKTA. . . TNIYIFNLALAD-ALVTTTMP. FQSTVYLMNSW-PFGDVLCKIVISIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALD-FR---. TPLK. A. KIINICIWLLSSSVGISAIVLGG-TKVRE Mu-type CVVGLFGNFLVMYVIVR. . YTKMKTA. . . TNIYIFNLALAD-ALATSTLP. FQSVNYLMGTW-PFGTILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALD-FR---. TPRN. A. KIINVCNWILSSAIG-LPVMFMAT----orexin FVVALVGNTLVCLAVWR. . NHHMRTV. . . TNYFIVNLSLADVLVTAICLP. ASLLVDITESW-LFGHALCKVIPYLQAVSVSVAVLTLSFIALDRWYAICHPLLF-K--S---. TARR. A. RGSILGIWAVSLAIMV--PQAAVM----rhodopsin IVLGFPINFLTLYVTVQ. . HKKLRTP. . . LNYILLNLAVADLFMVLGGFT. STLYTSLHGYF-VFGPTGCNLEGFFATLGGEIALWSLVVLAIERYVVVCKPMSN---F--RF. GENH. A. IMGVAFTWVMALACAA-PPLAGWS----5 -hydroxytryp IFCAVLGNACVVAAIAL. . ERSLQNV. . . ANYLIGSLAVTDLMVSVLVLP. MAALYQVLNKW-TLGQVTCDLFIALDVLCCTSSILHLCAIALDRYWAI--TDPI-DYVN-KR. TPRR. A. AALISLTWLIGFLIS-IPPMLGW-R--TP somatostatin CAAGLGGNTLVIYVVLR. . FAKMKTV. . . TNIYILNLAVAD-VLYMLGLP. FLATQNAASFW-PFGPVLCRLVMTLDGVNQFTSVFCLTVMSVDRYLAV--VHPL---SSARW. RRPRv. A. KLASAAAWVLSLCMS-LPLLV--F-ADVQ TM 1 TM 2 TM 3 adenosine NCGQP. KEGKNHSQGCGEGQVACLF-EDV-------VPMNYMVYFNF. . . FACVLVPLLLMLGVYLR. . I. . . FLAAR. . . beta-1 EA--R. R--CYNDPKC------CD-F--VT-------NRAY-AIASS. . . VVSFYVPLCIMAFVYLR. . V. . . FREAQ. . . beta EA--I. N--CYANETC------CD-F--FT-------NQAY-AIASS. . . IVSFYVPLVIMAFVYSR. . V. . . FQEAK. . . chemokine_r EA--D. D--R---YI-------CDRF--YP-------NDLWVVVFQFqhi. MVGLILPGIVILSCYCI. . ISKL-. . . D 2 NA---. ---DQNE-----CIIA------NPAF-VVYSS. . . IVSFYVPFIVTLLVYIK. . I. . . YIVLR. . . galanin --GTV. R--YGALEL-------CV-P--AW---EDARRRAL-DVATF. . . AAGYLLPVAVVSLAYGRtlrf. L. . . WAAVG. . . histamine TS---. ---KGNHTTS-----KCKVQ--V----NEVY-GLVDG. . . LVTFYLPLLIMCITYYR. . I. . . FKVAR. . . melatonin -----. ---EYDPRIY-----SCT-F--IQ-----TASTQY-TAAVV. . . VIHFLLPIAVVSFCYLR. . Iwvl. VLQAR. . . muscarinic QF--I. VGVRTVEDGE------CYIQ--FF------SNAAV-TFGTA. . . IAAFYLPVIIMTVLYWH. . I. . . SRASK. . . delta RDGAV. VCML-QFPSP-----SWYWD-TVT------KICVF. . . LFAFVVPILIITVCYGL. . M. . . LLRLR. . . kappa DVDVI. ECSL---QFP-------D-D-DYS------WWDLFMKICVF. . . IFAFVIPVLIIIVCYTL. . M. . . ILRLK. . . Mu-type ----T. K--YRQGSID------CTLT--FS--HPTWYWENLLKICVF. . . IFAFIMPVLIITVCYGL. . M. . . ILRLK. . . orexin ECSSVl. P--ELANRTR--LFSVCD--ERWA---DDLYPKIY-HSCFF. . . IVTYLAPLGLMAMAYFQ. . I. . . FRKLW. . . rhodopsin -----. ---R---YIPEGLQCSCGID--YYTLKPEVNNESF-VIYMF. . . VVHFTIPMIIIFFCYGQ. . L. . . VFTVK. . . 5 -hydroxytryp ED---. ---R-SDPDA------CTISKDH-----GY-TIYST. . . FGAFYIPLLLMLVLYGR. . I. . . FRAARfri somatostatin EGGTC. NASW---PEP------VGLW------GAVF-IIYTA. . . VLGFFAPLLVICLCYLL. . I. . . VVKVR. . . TM 5 TM 4 HAA. . K. . . SLAIIVGLFALCWLPLHI. . . IN. . CFT. . . F. FC-P KAL. . K. . . TLGIIMGVFTLCWLPFFL. . . AN. . VVK. . . A. FH-KAL. . K. . . TLGIIMGTFTLCWLPFFI. . . VN. . IVH. . . V. IQ-KAL. . K. . . TTVILILAFFACWLPYYIgis. ID. . SFIlle. I. IKQG KAT. . Q. . . MLAIVLGVFIICWLPFFI. . . TH. . ILN. . . I. HC-RATgrag. R. . . AMLAVAALYALCWGPHHA. . . L-. . ILC. . . F. WY-G KAT. . V. . . TLAAVMGAFIICWFPYFT. . . AF. . VYR. . . G. LR-G RSF. . L. . . TMFVVFVIFAICWAPLNC. . . IG. . LAV. . . A. IN-P VAN. . Qdpv. SPSLVQG-----RI. . . VK. . PNN. . . N. NM-P RIT. . R. . . MVLVVVGAFVVCWAPIHI. . . FV. . IVW. . . T. LV-RIT. . R. . . LVLVVVAVFVVCWTPIHI. . . FI. . LVE. . . A. LGST RIT. . R. . . MVLVVVAVFIVCWTPIHI. . y. VI. . IKA. . . L. VTIP KTA. . K. . . MLMVVLLVFALCYLPISV. . . LNvlkr. VFG. . . M. FRQA EVT. . R. . . MVIIMVIAFLICWVPYAS. . . VA. . FYI. . . F. THQG KTV. . K. . . TLGIIMGTFILCWLPFFI. . . VA. . LVL. . . P. FCES KVT. . R. . . MVLVVVLVFAGCWLPFFT. . . VN. . IVN. . . La. VALP adenosine. . . DCSHAPLW. . . LM. . . Y. . LAIVLSHTNSVVNPFI-YAYRIREFRQTFRKIIRSHV-. LRQqepfkaagtsarvlaahgsdgeqvslrlnghppg beta-1. . . RE-LVPDR. . . LF. . . V. . FFNWLGYANSAFNPII-Y-CRSPDFRKAFQGLLC---C. ARRaarrrhathgdrprasgclarpgpppspgaasdd beta. . . DN-LIRKE. . . VY. . . I. . LLNWIGYVNSGFNPLI-Y-CRSPDFRIAFQELL----C. LRRsslkaygngyssngntgeqsgyhveqekenkllc chemokine_r cef. EN-TVHKW. . . I-. . . S. . ITEALAFFHCCLNPIL-YAFLGAKFKTSAQHALTSVS-. -RGsslkilskgkrgghssvstesesssfhss. . . D 2. . . DC-NIPPV. . . LY. . . S. . AFTWLGYVNSAVNPII-YTTFNIEFRKAFLKIL--HC-. ---. . . . galanin. . . RF-AFSPA. . . TYacr. L. . ASHCLAYANSCLNPLV-YALASRHFRARFRRLWP---C. GRRrrhrarralrrvrpassgppgcpgdarpsgrlla histamine. . . DD-AINEV. . . LE. . . A. . IVLWLGYANSALNPIL-YAALNRDFRTGYQQLFC---Cr. LANrnshktslrsnasqlsrtqsreprqqeekplklq melatonin. . . QE-MAPQIpeg. LF. . . V. . TSYLLAYFNSCLNAIV-YGLLNQNFRREYKRIL-----. LALwnprhciqdaskgshaeglqspappiigvqhqad muscarinic. . . S-SDDGLE. . . --. . . . -HNKIQNGKAPRDPVTENCVQGEEKESSNDSTSVSAV-. ASNmrddeitqdentvstslghskdenskqtcirigt delta. . . DI-DRRDP. . . LV. . . Vaalh. LCIALGYANSSLNPVL-YAFLDENFKRCFRQLCR-KPC. GRPdpssfsrareatarervtactpsdgpgggaaa. . kappa. . . SHSTAALS. . . SY. . . . FCIALGYTNSSLNPIL-YAFLDENFKRCFRDFCF---P. LKMrmerqstsrvrntvqdpaylrdidgmnkpv. . Mu-type. . . ET-TFQTV. . . SW. . . H. . FCIALGYTNSCLNPVL-YAFLDENFKRCFREFCIPTS-. -SNieqqnstrirqntrdhpstantvdrtnhqlenle orexin. . . SDREAVYA. . . CF. . . T. . FSHWLVYANSAANPII-YNFLSGKFREQFKAAF----S. CCLpglgpcgslkapsprssashkslslqsrcsiski rhodopsin. . . SN--FGPI. . . FM. . . T. . IPAFFAKSAAIYNPVIYI-MMNKQFRNCMLTTIC---C. GKNplgddeasatvsktetsqvapa. . . 5 -hydroxytryp. . . SC-HMPTL. . . LG. . . A. . IINWLGYSNSLLNPVI-YAYFNKDFQNAFKKIIKCKF-. CRQ. . . . somatostatin. . . QE-PASAG. . . LY. . . FVVILSYANSCANPVL-YGFLSDNFRQSFQKVL----C. LRKgsgakdadateprpdrirqqqeatrprtaaangl TM 7 Fig 19. Sequence aligment of many class A receptors based on hmm model TM 6
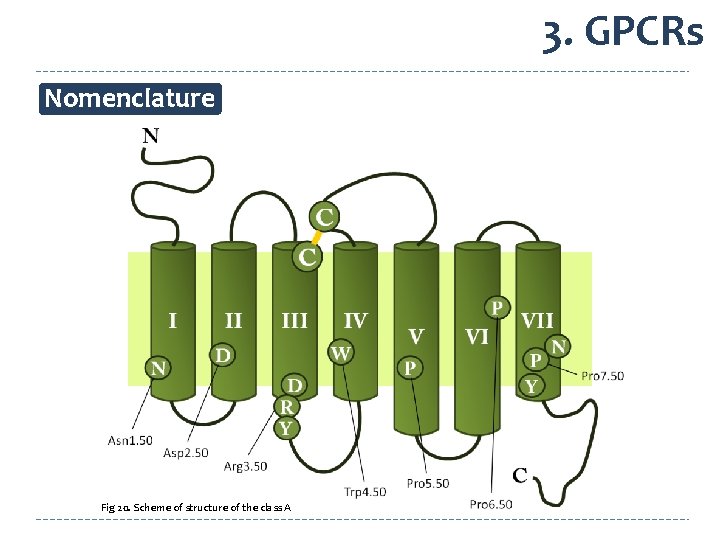
3. GPCRs Nomenclature Fig 20. Scheme of structure of the class A
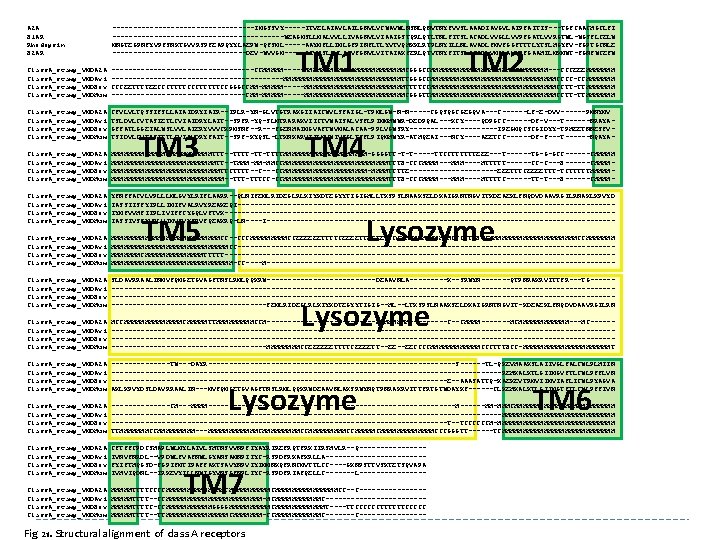
A 2 A B 1 AR Rhodopsin B 2 AR -----------------IMGSSVY-----ITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---TGFCAACHGCLFI ---------------------WEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWL-WGSFLCELW MNGTEGPNFYVPFSNKTGVVRSPFEAPQYYLAEPW-QFSML-----AAYMFLLIMLGFPINFLTLYVTVQHKKLRTPLNYILLNLAVADLFMVFGGFTTTLYTSLHGYFV-FGPTGCNLE ----------------DEV-WVVGM-----GIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWT-FGNFWCEFW Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum -----------------CCHHHHH-----HHHHHHHHHHHHCGGGCCHHHHHHHHHHHHHH---CCCEEEHHHHHHH ---------------------HHHHHHHHHHHHHHTTGGGCCHHHHHHHHHHHHHHHCCCC-CCHHHHHHH CCCEETTTTEECCCTTTTTCCGGGCCHH-HHHHH-----HHHHHHHHHHHHTTTTCCHHHHHHHHHHHHHHHCCTT-TTHHHHHHH ----------------CHH-HHHHH-----HHHHHHHHHHHHCGGGTTHHHHHHHHHHHHHHHCCTT-TTHHHHHHH Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum CFVLVLTQSSIFSLLAIAIDRYIAIR--IPLR-YN-GLVTGTRAKGIIAICWVLSFAIGL-TPMLGW-N-N-----CGQSQGCGEGQVA---C------LF-E-DVV------PMNYMV TSLDVLCVTASIETLCVIAIDRYLAIT--SPFR-YQ-SLMTRARAKVIICTVWAISALVSFLPIMMHWWR-DEDPQAL---KCY----QDPGCC------DF-V---T------NRAYAGFFATLGGEIALWSLVVLAIERYVVVCKPMSNF--R---FGENHAIMGVAFTWVMALACAA-PPLVGWSRY-----------IPEGMQCSCGIDYY-TPHEETNNESFVTSIDVLCVTASIETLCVIAVDRYFAIT--SPF-KYQSL-LTKNKARVIILMVWIVSGLTSFLPIQMHWYR-ATHQEAI---NCY----AEETCC------DF-F---T------NQAYA- Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum HHHHHHHHHHHHTTT--TTTT-TT-TTTTHHHHHHHHHH-GGGGTT-T-T-----TTCCTTTTTTEEE---T------TG-G-GCC------CHHHHHHHHHHHHHTT--THHH-HH-HHCCHHHHHHHHHHHHHTTTB-CCCHHHH----HTTTTT------CC-C---B------CHHHHHHHHHHHHHHHTTCTTTT--C---CCHHHHHHHHHH-HHHHTTTTE-----------EEETTTCEEEETTT-TCTTTTTHHHHHHHHHHHHHHHH--TTTCC-CCHHHHHHHHHHHHHTTTB-CCCHHHH----HTTTTC------TT-T---B------CHHHH- Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum YFNFFACVLVPLLLMLGVYLRIFLAARR--QLNIFEMLRIDEGLRLKIYKDTEGYYTIGIGHLLTKSPSLNAAKSELDKAIGRNTNGVITKDEAEKLFNQDVDAAVRGILRNAKLKPVYD IASSIISFYIPLLIMIFVALRVYREAKEQI---------------------------------------------IYMFVVHFIIPLIVIFFCYGQLVFTVK----------------------------------------------IASSIVSFYVPLVIMVFVYSRVFQEAKRQ-LN----I------------------------------------------ Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum HHHHHHHHHHHHHCC--CCCHHHHHCCEEEEEETTTTCEEEETTEEEECCCCHHHHHHCCCTTTBCCHHHHHHHHHHHCCHHHHHHHHHHHHHHCC---------------------------------------------HHHHHHHCHHHHHHHTTTTT----------------------------------------------HHHHHHHHHHHHHHH-CC----H------------------------------------------ Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum SLDAVRRAALINMVFQMGETGVAGFTNSLRMLQQKRW-------------DEAAVNLA-----K--SRWYN-------QTPNRAKRVITTFR---TG--------------------------------------------------------------------------------------------------------------------------------------------FEMLRIDEGLRLKIYKDTEGYYTIGIG--HL--LTKSPSLNAAKSELDKAIGRNTNGVIT-KDEAEKLFNQDVDAAVRGILRN Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum HCCHHHHHHHCHHHHHTTHHHHHCCH-------------HHHH-----C--CHHHH-------HCHHHHHH---HC--------------------------------------------------------------------------------------------------------------------------------------------HHHHCCEEEEEETTTTCEEEETT--EECCCCHHHHHHCCCTTTBCC-HHHHHHHHHHHT Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum -------TW---DAYR------------------------------S------TL-QKEVHAAKSLAIIVGLFALCWLPLHIIN -----------------------------------------------REHKALKTLGIIMGVFTLCWLPFFLVN ----------------------------------------E--AAASATTQ-KAEKEVTRMVIIMVIAFLICWLPYAGVA AKLKPVYDSLDAVRRAALIN---MVFQMGETGVAGFTNSLRMLQQKRWDEAAVNLAKSRWYNQTPNRAKRVITTFRTGTWDAYKF------CLKEHKALKTLGIIMGTFTLCWLPFFIVN Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum -------CH---HHHH------------------------------H------HH-HHHCHHHHHHHHHHHH -----------------------------------------------HHHHHHHHHHHHHH ----------------------------------------T--TCCCCCCH-HHHHHHHHHHHHHH TTHHHHHHHCCHHHHH---HHHHHHCHHHHHHHHHCCHHHHHHHHHCCCGGGTT------TCHHHHHHHHHHHHHH Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum CFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLR--Q--------IVNVFNRDL--VPDWLFVAFNWLGYANSAMNPIIYC-RSPDFRKAFKRLLA------------FYIFTHQGSD-FGPIFMTIPAFFAKTSAVYNPVIYIMMNKQFRNCMVTTLCC----GKNPSTTVSKTETSQVAPA IVHVIQDNL--IRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC-------L-------- Class. A_stamp_VMDA 2 A Class. A_stamp_VMDAvi Class. A_stamp_VMDBov Class. A_stamp_VMDHum HHHHHTTTTCCCCHHHHHHHCHHHHHHHHHHHHHHHCC--C--------HHHHHTTTT--CCHHHHHHHHHHHH-HCHHHHHHC------------HHHHHTTTTC-CCHHHHHHGGGGHHHHHCHHHHHHT----TTCCCCCCTTTTTTCCCCC HHHHHTTTT--TTHHHHHHHHCHHHHHHH-CCHHHHHHC------------ TM 1 TM 3 TM 2 TM 4 TM 5 Lysozyme TM 7 Fig 21. Structural alignment of class A receptors TM 6
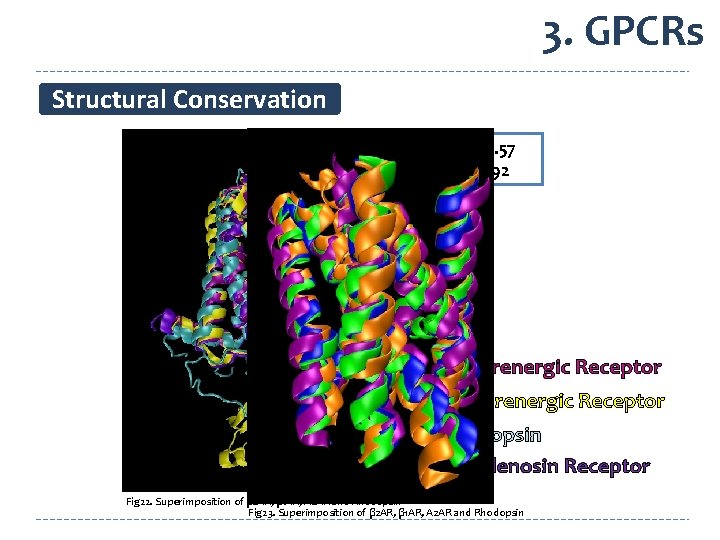
3. GPCRs Structural Conservation SCORE 5. 57 RMSD 1. 92 β 1 -Adrenergic Receptor β 2 -Adrenergic Receptor Rhodopsin A 2 -Adenosin Receptor Fig 22. Superimposition of β 2 AR, β 1 AR, A 2 AR and Rhodopsin Fig 23. Superimposition of β 2 AR, β 1 AR, A 2 AR and Rhodopsin

3. GPCRs Structural Conservation Fig 24. Superimposition of β 2 AR, β 1 AR, A 2 AR and Rhodopsin
![3. GPCRs β 2 Adrenergic Receptor >beta 2 adrenergic receptor [Homo sapiens] MGQPGNGSAFLLAPNRSHAPDHDVTQQRDEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKF ERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIET 3. GPCRs β 2 Adrenergic Receptor >beta 2 adrenergic receptor [Homo sapiens] MGQPGNGSAFLLAPNRSHAPDHDVTQQRDEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKF ERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIET](http://slidetodoc.com/presentation_image_h2/c103ba0c8751823458ac6e4a8aa96f8c/image-22.jpg)
3. GPCRs β 2 Adrenergic Receptor >beta 2 adrenergic receptor [Homo sapiens] MGQPGNGSAFLLAPNRSHAPDHDVTQQRDEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKF ERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIET LCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHQEAINCYA NETCCDFFTNQAYAIASSIVSFYVPLVIMAFVYSRVFQEAKRQLQKIDKSEGRFHVQNLSQVE QDGRTGHGLRRSSKFCLKEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNLIRKEVYILLNW IGYVNSGFNPLIYCRSPDFRIAFQELLCLRRSSLKAYGNGYSSNGNTGEQSGYHVEQEKENKL LCEDLPGTEDFVGHQGTVPSDNIDSQGRNCSTNDSLL Fig 25. β 2 Adrenergic Receptor sequence β 2 AR G proteins Adenilate cyclase AMPc PKA Smooth muscle relaxation, blood vessels dilatation, striated muscle contraction, bronchiole dilation. . . • The first non-rhodopsin GPCR cloned • One of the most extensively studied members of this large receptor family.

3. GPCRs β 2 Adrenergic Receptor MEMSAT software to predict alpha helix transmembrane regions based on Dr. Jones predictive algorithm described in 1994 Fig 26 Hydropathy plot of β 2 Adrenergic Receptor sequence Fig 27. Hydropathy plot of β 2 Adrenergic Receptor sequence
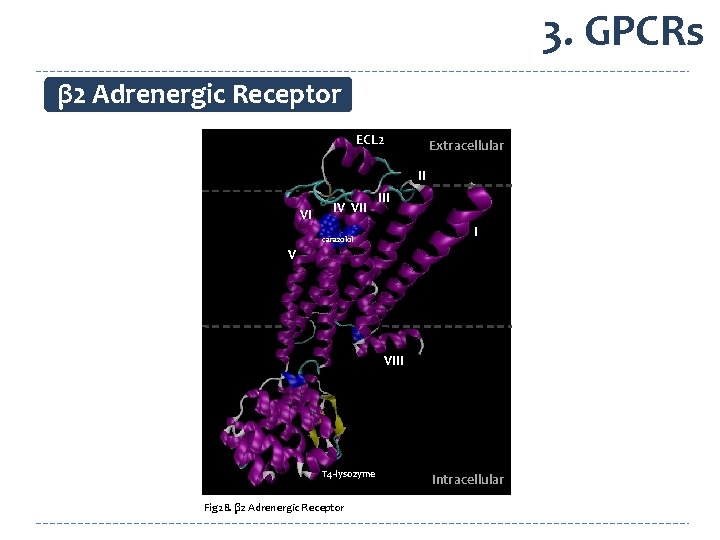
3. GPCRs β 2 Adrenergic Receptor ECL 2 Extracellular II VI IV VII I carazolol V VIII T 4 -lysozyme Fig 28. β 2 Adrenergic Receptor Intracellular
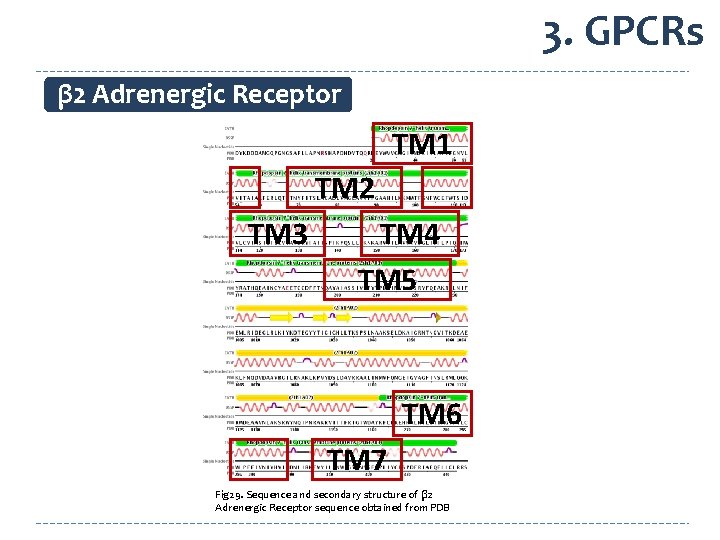
3. GPCRs β 2 Adrenergic Receptor TM 1 TM 2 TM 3 TM 4 TM 5 TM 6 TM 7 Fig 29. Sequence and secondary structure of β 2 Adrenergic Receptor sequence obtained from PDB
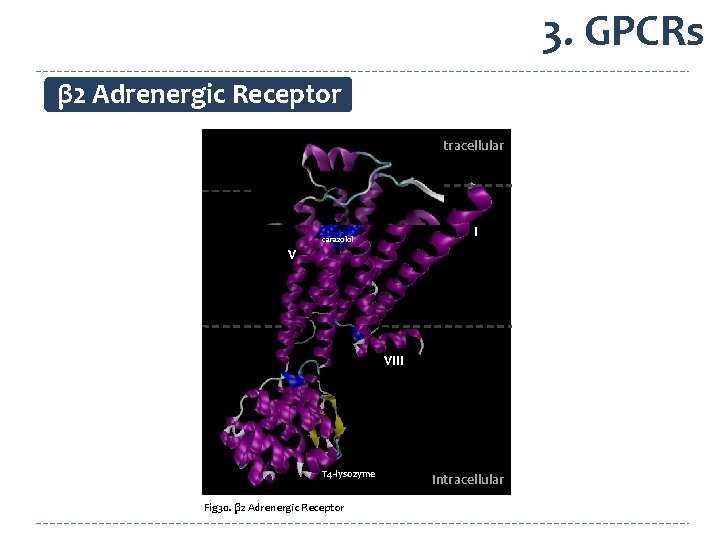
3. GPCRs β 2 Adrenergic Receptor ECL 2 Extracellular II VI IV VII I carazolol V VIII T 4 -lysozyme Fig 30. β 2 Adrenergic Receptor Intracellular
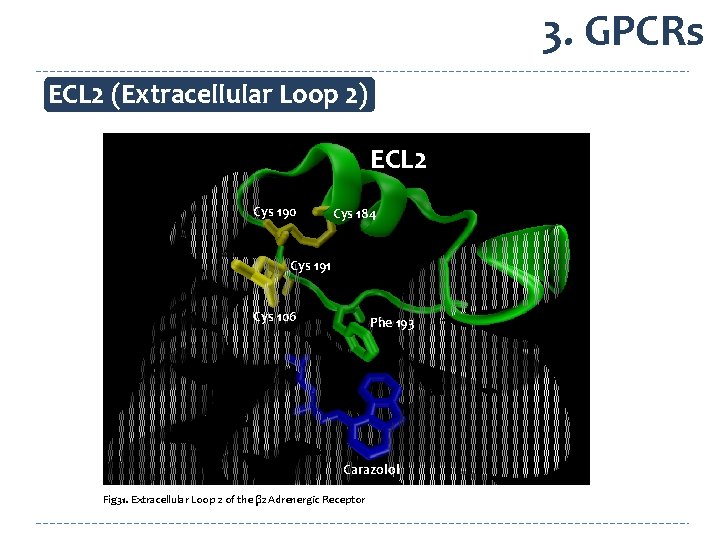
3. GPCRs ECL 2 (Extracellular Loop 2) ECL 2 Cys 190 Cys 184 Cys 191 Cys 106 Phe 193 Carazolol Fig 31. Extracellular Loop 2 of the β 2 Adrenergic Receptor
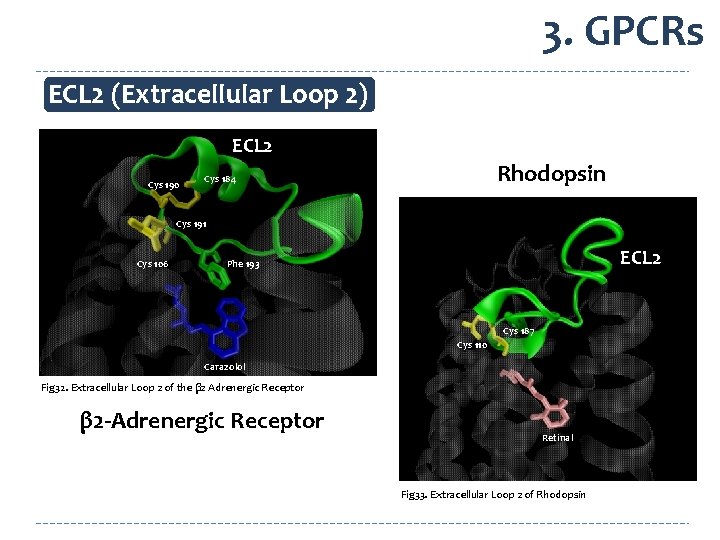
3. GPCRs ECL 2 (Extracellular Loop 2) ECL 2 Cys 190 Rhodopsin Cys 184 Cys 191 Cys 106 ECL 2 Phe 193 Cys 187 Cys 110 Carazolol Fig 32. Extracellular Loop 2 of the β 2 Adrenergic Receptor β 2 -Adrenergic Receptor Retinal Fig 33. Extracellular Loop 2 of Rhodopsin
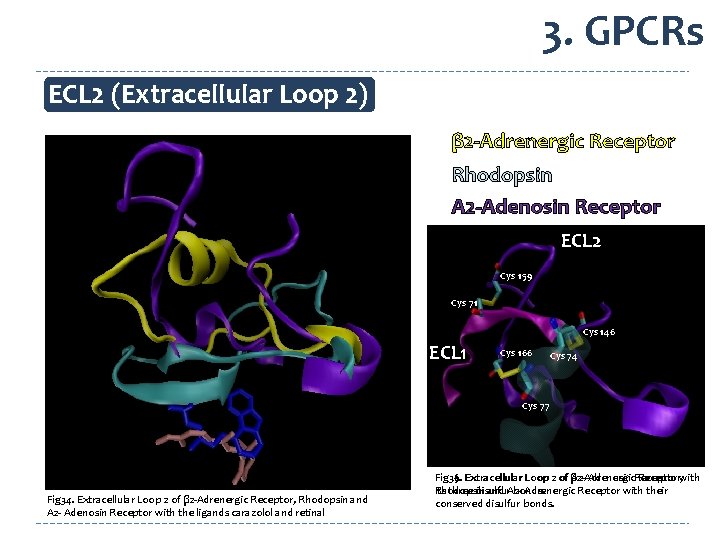
3. GPCRs ECL 2 (Extracellular Loop 2) β 2 -Adrenergic Receptor Rhodopsin A 2 -Adenosin Receptor ECL 2 Cys 159 Cys 71 Cys 146 ECL 1 Cys 166 Cys 74 Cys 77 Fig 34. Extracellular Loop 2 of β 2 -Adrenergic Receptor, Rhodopsin and A 2 - Adenosin Receptor with the ligands carazolol and retinal Fig 35. Receptor, with Fig 36. Extracellular Loop 2 of β 2 -Adrenergic A 2 - Adenosin Receptor Rhodopsin and A 2 Adrenergic Receptor with their its three disulfur bonds. conserved disulfur bonds.
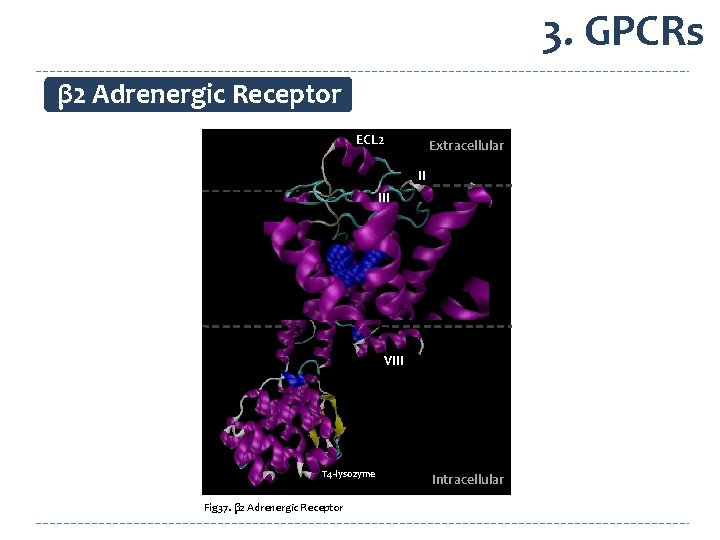
3. GPCRs β 2 Adrenergic Receptor ECL 2 Extracellular II VI IV VII I carazolol V VIII T 4 -lysozyme Fig 37. β 2 Adrenergic Receptor Intracellular
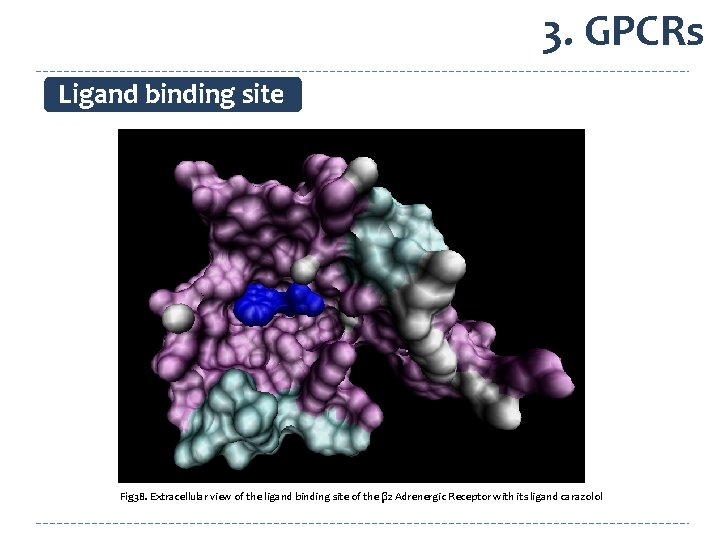
3. GPCRs Ligand binding site Fig 38. Extracellular view of the ligand binding site of the β 2 Adrenergic Receptor with its ligand carazolol
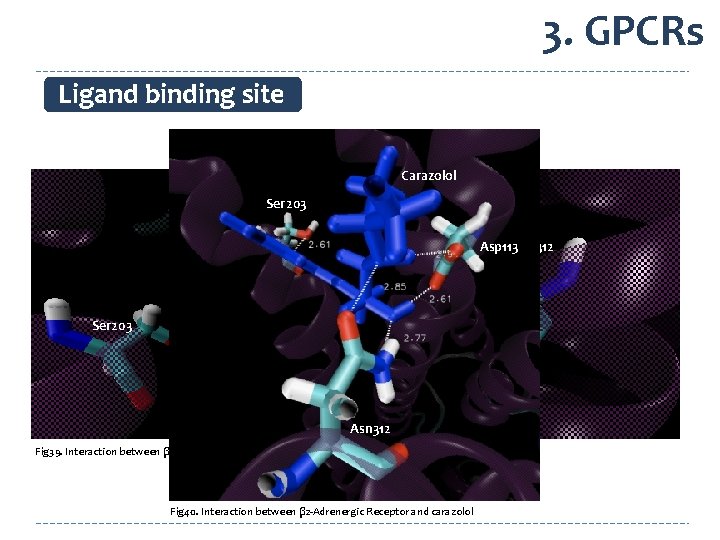
3. GPCRs Ligand binding site Carazolol Ser 203 Asp 113 Asn 312 Ser 203 Asp 113 Carazolol Asn 312 Fig 39. Interaction between β 2 -Adrenergic Receptor and carazolol Fig 40. Interaction between β 2 -Adrenergic Receptor and carazolol
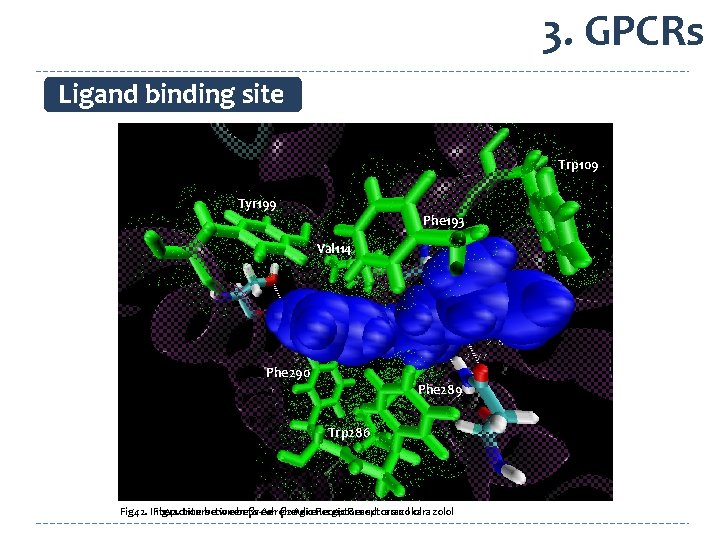
3. GPCRs Ligand binding site Trp 109 Tyr 199 Phe 193 Val 114 Trp 286 Phe 193 Val 114 Phe 289 Phe 290 Tyr 199 Phe 289 Trp 286 Phe 290 Fig 42. Interaction Fig 41. Interaction between β 2 -Adrenergic Receptor and carazolol
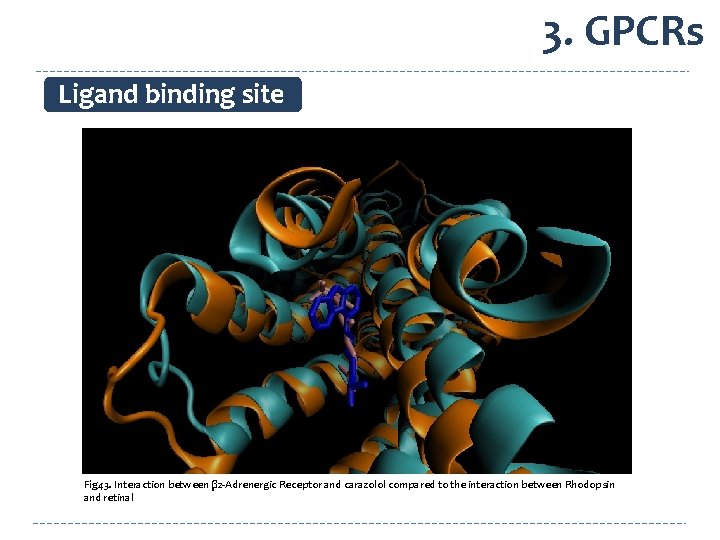
3. GPCRs Ligand binding site Fig 43. Interaction between β 2 -Adrenergic Receptor and carazolol compared to the interaction between Rhodopsin and retinal
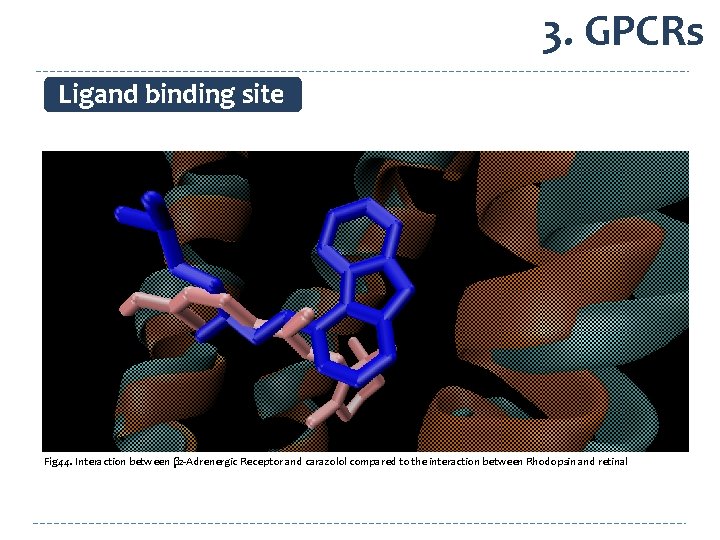
3. GPCRs Ligand binding site Fig 44. Interaction between β 2 -Adrenergic Receptor and carazolol compared to the interaction between Rhodopsin and retinal
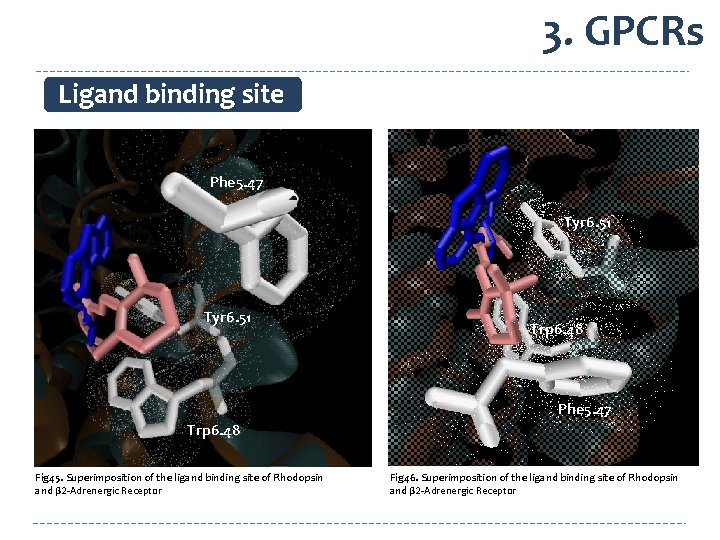
3. GPCRs Ligand binding site Phe 5. 47 Tyr 6. 51 Trp 6. 48 Phe 5. 47 Trp 6. 48 Fig 45. Superimposition of the ligand binding site of Rhodopsin and β 2 -Adrenergic Receptor Fig 46. Superimposition of the ligand binding site of Rhodopsin and β 2 -Adrenergic Receptor

3. GPCRs Ligand binding site Rhodopsin β 2 -Adrenergic Receptor Trp 6. 48 Fig 47. Interaction between Rhodopsin and retinal through Trp 6. 48 Fig 48. Interaction between β 2 -Adrenergic Receptor and carazolol and Trp 6. 48 “Toogle Switch”
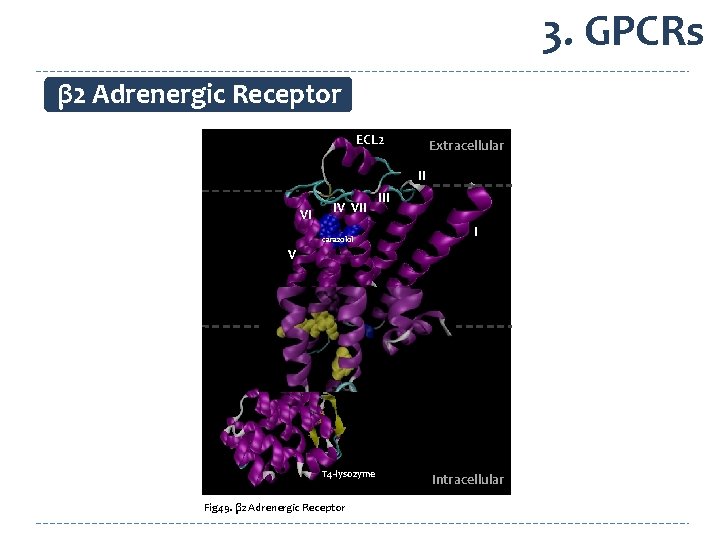
3. GPCRs β 2 Adrenergic Receptor ECL 2 Extracellular II VI IV VII I carazolol V VIII T 4 -lysozyme Fig 49. β 2 Adrenergic Receptor Intracellular
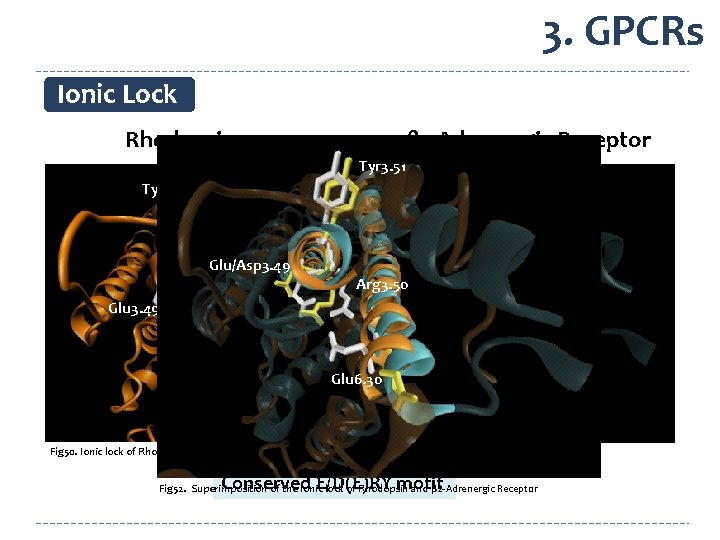
3. GPCRs Ionic Lock Rhodopsin β 2 -Adrenergic Receptor Tyr 3. 51 Asp 3. 49 Glu/Asp 3. 49 Arg 3. 50 Glu 3. 49 Glu 6. 30 Fig 50. Ionic lock of Rhodopsin Arg 3. 50 Glu 6. 30 Fig 51. Ionic lock of β 2 -Adrenergic Receptor Conserved E/D(E)RY motif Fig 52. Superimposition of the Ionic lock of Rhodopsin and β 2 -Adrenergic Receptor
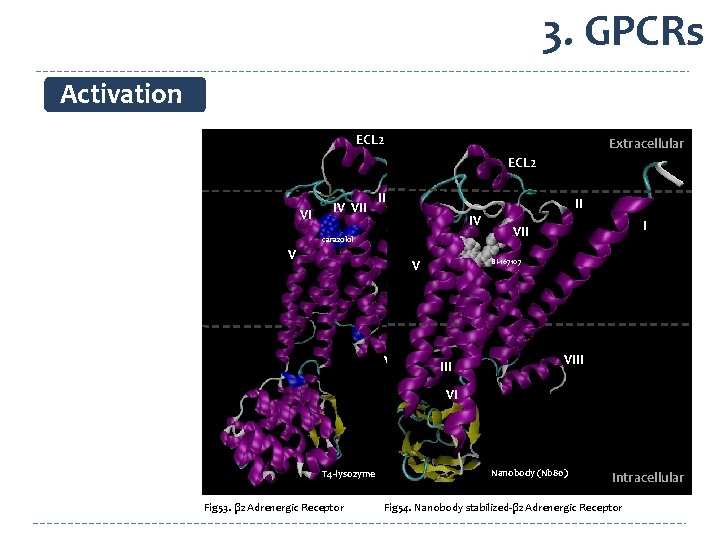
3. GPCRs Activation ECL 2 Extracellular II VI IV VII Extracellular ECL 2 III II IV I carazolol V V VIII I VII BI-167107 VIII VI T 4 -lysozyme Fig 53. β 2 Adrenergic Receptor Nanobody (Nb 80) Intracellular Fig 54. Nanobody stabilized-β 2 Adrenergic Receptor
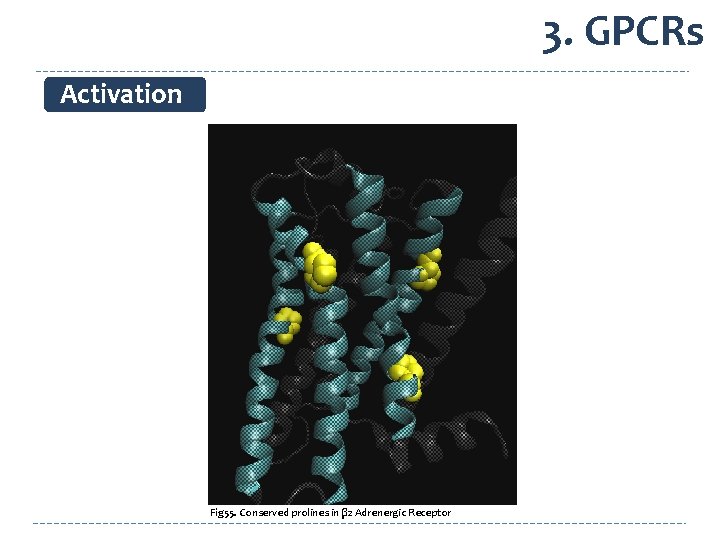
3. GPCRs Activation Fig 55. Conserved prolines in β 2 Adrenergic Receptor
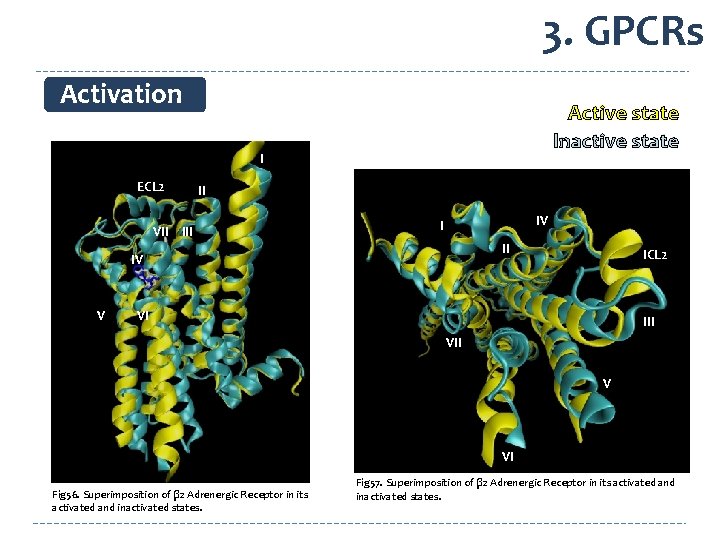
3. GPCRs Activation Active state Inactive state I ECL 2 II VII IV I II IV V ICL 2 VI III V VI Fig 56. Superimposition of β 2 Adrenergic Receptor in its activated and inactivated states. Fig 57. Superimposition of β 2 Adrenergic Receptor in its activated and inactivated states.
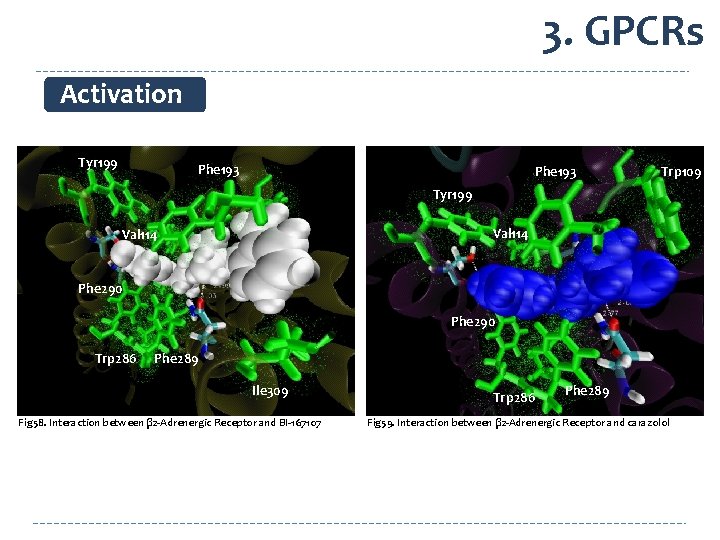
3. GPCRs Activation Tyr 199 Phe 193 Trp 109 Tyr 199 Val 114 Phe 290 Trp 286 Phe 289 Ile 309 Fig 58. Interaction between β 2 -Adrenergic Receptor and BI-167107 Trp 286 Phe 289 Fig 59. Interaction between β 2 -Adrenergic Receptor and carazolol
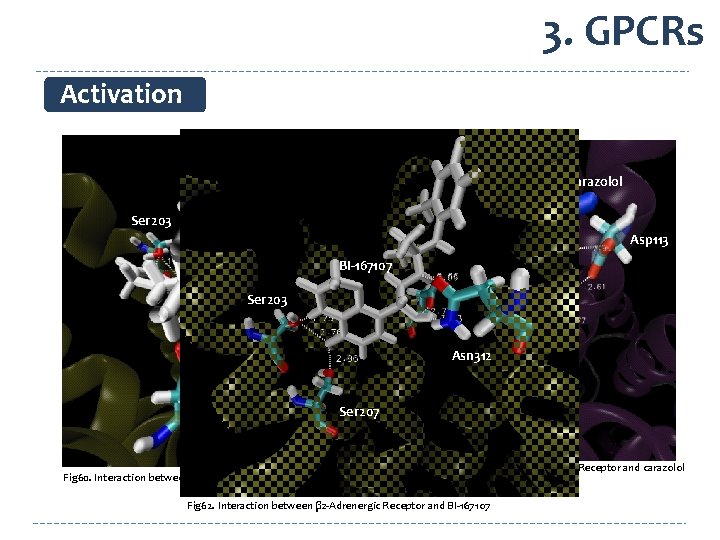
3. GPCRs Activation Carazolol BI-167107 Ser 203 Asp 113 BI-167107 Ser 203 Asn 312 Ser 207 Fig 60. Interaction between β 2 -Adrenergic Receptor and BI-167107 Fig 61. Interaction between β 2 -Adrenergic Receptor and carazolol Fig 62. Interaction between β 2 -Adrenergic Receptor and BI-167107
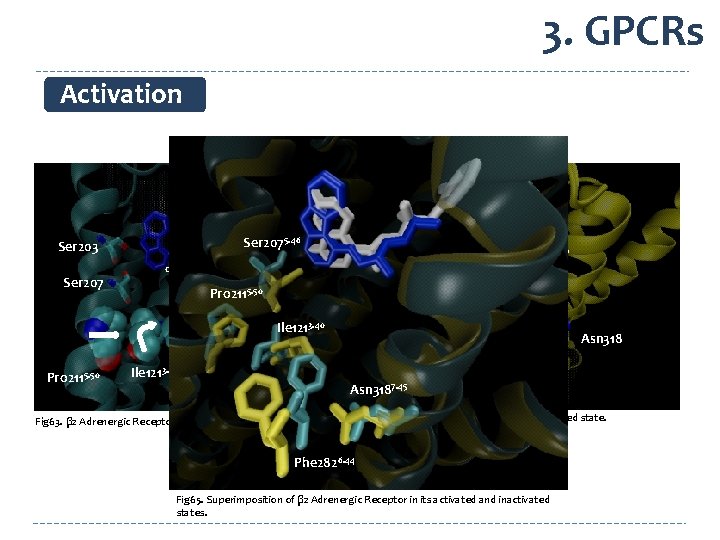
3. GPCRs Activation Ser 2075. 46 Ser 203 Ser 207 BI-167107 carazolol Ser 203 Pro 2115. 50 Ser 207 Ile 1213. 40 Ile 121 Asn 3187. 45 Pro 2115. 50 Ile 1213. 40 Phe 2826. 44 Pro 211 Asn 3187. 45 Phe 282 Fig 64. Adrenergic Receptor in its activated state. Fig 63. β 2 Adrenergic Receptor in its inactivated state. Phe 2826. 44 Fig 65. Superimposition of β 2 Adrenergic Receptor in its activated and inactivated states.
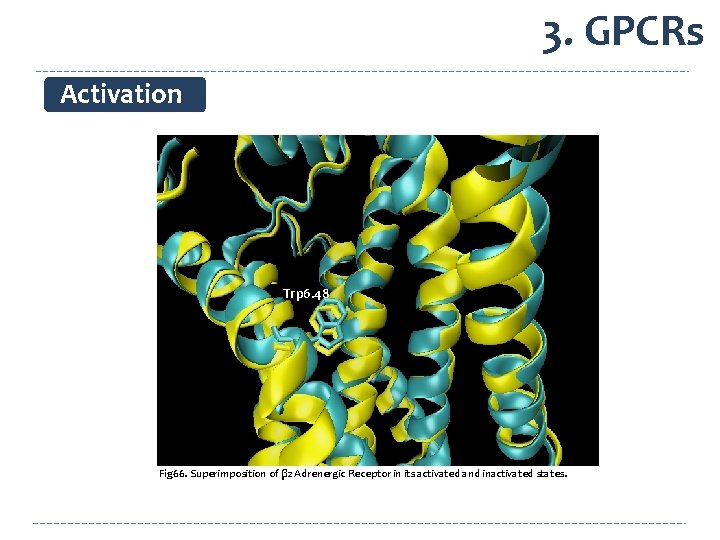
3. GPCRs Activation Trp 6. 48 Fig 66. Superimposition of β 2 Adrenergic Receptor in its activated and inactivated states.
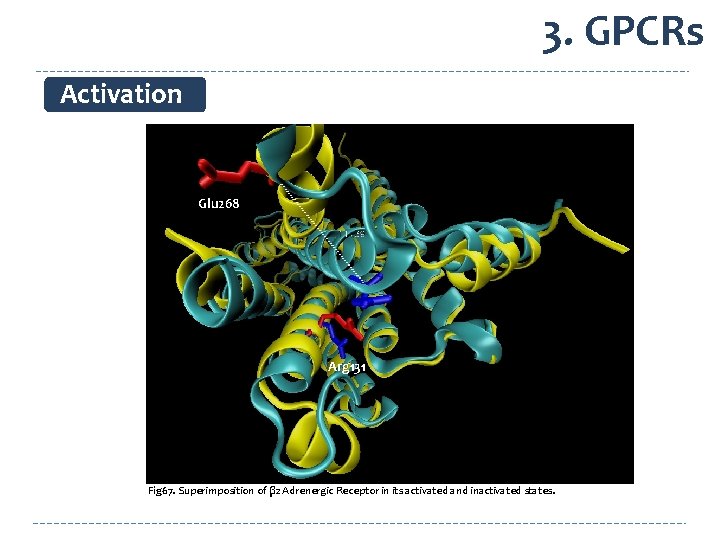
3. GPCRs Activation Glu 268 Arg 131 Fig 67. Superimposition of β 2 Adrenergic Receptor in its activated and inactivated states.
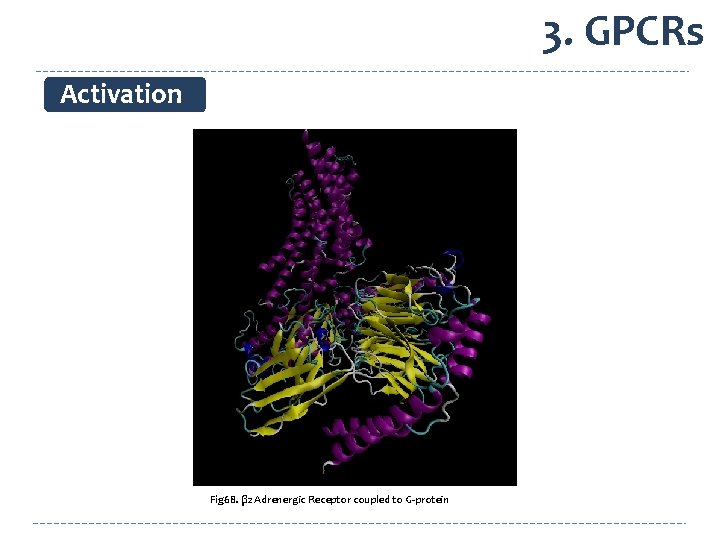
3. GPCRs Activation Glu 268 Arg 131 Fig 68. β 2 Adrenergic Receptor coupled to G-protein

4. Conclusions G-protein-coupled receptors represent the largest class of drug targets, and its structure determination is very important to design new specific drugs. Although the crystallization methods have improved incredibly, we need to crystallize new states of structures that have already been crystallized to understand better GPCR dynamics such as the activation process. In addition, the crystallization of new structures is needed to amplify the knowledge of these receptors and to design new drugs.

5. Bibliography Costanzi S. On the Applicability of GPCR Homology Models to Computer-Aided Drug Discovery: A Comparison between In Silico and Crystal Structures of the β 2 - Adrenergic receptor. J. Med. Chem. 2008; 51: 2907 -2914. Cherezov V, Rosenbaum D. M, Hanson M. A, Rasmussen S. G. F, Thian F. S, Kobilka T. S. , Choi H, Kuhn P, Weis W. I, Kobilka B. K, Stevents R. C. High – Resolution Crystal Structure of an Engineered Human β 2 Adrenergic G Protein – Coupled Receptor. Deupi X, Kobilka B. K. Energy landscapes as a tool to integrate GPCR structure, dynamics, and function. Physiology (Bethesda). 2010: 25; 293– 303 Deupi X, Standfuss J. Structural insights into agonist-induced activation of G-protein-coupled receptors. Current Opinion in Structural Biology. 2011; 21: 541 – 551. Dror R. O, Arlow D. H, Maragakis P, Mildorf T. J, Pan A. C, Xu H, Borhani D. W, Shaw D. E. Activation mechanism of the β 2 - Adrenergic receptor. PNAS. 2011; 108: 18684 – 18689. Fairman J. W, Noinaj N, Buchanan S. K. The structural biology of β- barrel membrane proteins: a summary of recent report. Current Opinion Structural Biology. 2011; 21: 523 – 531. Goetz A, Lanig, Gmeiner P, Clark T. Molecular Dynamics Simulations of the Effect of the G-Protein and Diffusible Ligands β 2 -Adrenergic Receptors. J. Mol. Biol. 2011; 414: 611 – 623. González A, Perez-Acle T, Pardo L, Deupi X. Molecular Basis of Ligand Dissociation in β-Adrenergic Receptor. Plos ONE. 2011; 6.

5. Bibliography Jones D. T. , Taylor W. R. , Thornton J. M. A model recognition Approach to the prediction of All – Helical Membrane Protein Structure and Toxicology. Biochemistry. 1994; 33 (10): 3038 -3049. Kahsai A. W, Xiao K, Rajagopal S, Ahn S, Shukla A, Sun J, Oas T. G, Lefkowitz R. J. Multiple ligand – specific conformations of the β 2 - adrenergic receptor. Nature Chemical Biology. 2011; 7: 692 – 700. Katritch V, Abagyan R. GPCR agonist binding revealed by modeling and crystallography. Trends in Pharmacological Science. 2011; 31 (11): 637 – 643. Kobilka B. K. Structural insights into andrenergic receptor function and pharmacology. Trends in Pharmacological Science. 2011; 21 (4): 213 -218. Kolb P, et al. Structure-based discovery of β 2 - Adrenergic receptor ligands. Proc. Natl. Acad. Sci. U. S. A. 2009; 106: 6843– 6848 López Muñoz L. Homology modeling and structural analysis of the antipsychotic drugs receptorome. [Doctoral Thesis]. Universitat Pompeu Fabra; 2010. Massotte D, Kieffler B. L. Structure – Function Relationships in G Proteins – Coupled Receptors Handbook. In: Devi L. A, editor. The G Protein – Coupled Receptors. New Yersey: Humana Press; 2005. 3 -39. Palczewski K, et al. Crystal structure of rhodopsin: a G protein-Coupled receptor. Science. 2000; 289: 739– 745.

5. Bibliography Rasmussen S. G, et al. Crystal structure of the human β 2 -Adrenergic G-protein-coupled receptor. Nature. 2007; 450: 383– 387. Rasmussen S. G, De. Vree B. T, Zou Y. , Kruse A. C, Chung K. Y, Kobilka T. S. Crystal structure of the β 2 Adrenergic Receptor-Gs protein complex. Nature. 2011; 477: 549 -555. Rasmussen S. G, et al. Structure of a nanobody-stabilized active state of the β 2 - adrenoceptor. Nature. 2011; 469: 175– 180. Rosenbaum D. M, et al. GPCR engineering yields high-resolution structural insights into β 2 -adrenergic receptor function. Science. 2007; 318: 1266– 1273 Rosenbaum D. M, Rasmussen S. G, Kobilka B. K. The β 2 - Adrenergic Receptor as a Model for G-Protein. Coupled Receptor Structure and Activation by Diffusible Hormones. Handbook of Cell Signaling. 2010; Chapter 25: 163 -168. Rosenbaum D. M, et al. Structure and function of an irreversible agonist-β 2 adrenoceptor complex. Nature. 2011; 469: 236– 240. Sabio M, et al. Use of the X-ray structure of the β 2 - Adrenergic receptor for drug discovery. Part 2: identification of active compounds. Bioorg. Med. Chem. Lett. 2008; 18: 5391– 5395. Sansuk K, et al. A structural insight into the reorientation of transmembrane domains 3 and 5 during family A GPCR activation. Mol. Pharmacol. 2011; 79: 262– 269

5. Bibliography Schulz G. β- barrel membrane proteins. Current Opinion in Structural Biology. 2000; 10: 443 – 447. Topiol S, Sabio M. X-ray structure breakthroghs in the GPCR transmembrane region. Biochemical Pharmacology. 2009; 79: 11 – 20. Warne T, et al. Structure of a β 1 - Adrenergic G-protein-coupled receptor. Nature. 2008; 454: 486– 491. Weis W. I, Kobilka B. K. Structural insights into G-protein-coupled receptor activation. Current Opinion Structural Biology. 2008; 18: 734 – 740. Wimley W. C. The versatile β- barrel membrane protein. Current Opinion in Structural Biology. 2003; 13: 404 – 411. Worth C. L. , Kleinau G, Krause G. Comparative Sequence and Structural Analysen of G-protein-coupled Receptor Crystal Structures and Implications for Molecular Models. Plos One. 2009; 9.

THANKS FOR YOUR ATTENTION!!!! QUESTIONS?
- Slides: 54