MedCORDEX database netcdf files File System XFS file

Med-CORDEX database = = = netcdf files File System XFS file server + + their info relational database mysql db LAMP server Linux, Apache, Mysql and PHP www. medcordex. eu 1

file server NETAPP FAS 3240 HA Storage System § dual controller § RAID DP technology (two simultaneus disk failures allowed) environment: § dual power supply (one coming from UPS) § air-conditioned room www. medcordex. eu 2

LAMP server HP DL 575 G 7 Linux Server § § SLES 11 SP 2 Operating System no users: the machine is devoted to act as a webserver (not only for Med-CORDEX database) Apache 2. 4. 6 JVM 1. 7. 0_55 PHP 5. 5. 10 mysql 5. 0. 96 Tomcat 7. 0. 52 pure-ftpd 1. 0. 36 Environment: § dual power supply (one coming from UPS) § air-conditioned room www. medcordex. eu 3

paths & filenames ATMOSPHERIC DATA According to “CORDEX Archive Design” O. B. Christensen, W. J Gutowski, G. Nikulin, and S. Legutke http : //cordex. dmi. dk • PATH /MEDCORDEX/<Domain>/<Institution>/<GCMModel. Name>/ <CMIP 5 Experiment. Name>/<CMIP 5 Ensemble. Member>/<RCMMode l. Name>/<RCMVersion. ID>/<Frequency>/<Variable. Name> Our PATH shortcut: /MEDCORDEX/ALL (files are not listable) • FILENAME Variable. Name_Domain_GCMModel. Name_CMIP 5 Experiment. Name _CMIP 5 Ensemble. Member_RCMModel. Name_RCMVersion. ID_Frequ ency[_Start. Time-End. Time]. nc www. medcordex. eu 4

paths & filenames OCEAN DATA Not yet defined a standard (AFAIK) shall we use http: //cmip-pcmdi. llnl. gov/cmip 5/output_req. html#req_list ? www. medcordex. eu 5

paths & filenames All tokens which form the PATH are derived from FILENAME but the Institution which is the name of the directory where files have been placed by each data providers e. g. /incoming_MEDCORDEX/ENEA In the db we use all tokens and one more info: realm which is atmosphere or ocean. Realm is deduced from the Variable. Name THUS WE HAVE A CONSTRAINT ! variables must ALL be unique regardless to the realm they belong to! www. medcordex. eu 6

uploading files Data providers having data to upload can use ANY ftp client to do: ftp cd mput ftp: //user: passw@www. medcordex. eu /incoming_MEDCORDEX/$INST *. nc (all files into the same flat dir) PLEASEGO. txt (any size, also empty) where $INST is the code of their institution (eg: ENEA) Then they wait for the automatic daily procedure to start (at 20: 00) www. medcordex. eu 7

ingesting files Every day at 20: 00 is automatically run the “ingesting procedure” § For each dir /incoming_MEDCORDEX/$INST with PLEASEGO. txt: § for each other file in the dir, the procedure: 1. verifies it’s a netcdf file ncdump -h works properly 2. splits filenames in tokens and checks their compliance to CORDEX standard 3. checks validity of variable name 4. creates the right $PATH in /MEDCORDEX 5. moves the file into its $PATH 6. inserts/updates the file’s record in the db it is already known also ncdump –h continue www. medcordex. eu 8

ingesting files § When data provider’s files are all processed a mail is sent to him/her with the log of what happened ingesting his/her data After ingesting all files of all data providers, the procedure: 1. computes some statistics and publishes them on www. medcordex. eu/stats taking figures from db & ftp logs 2. makes all links in /MEDCORDEX/ALL 3. copies the whole /MEDCORDEX directory to another host www. medcordex. eu 9
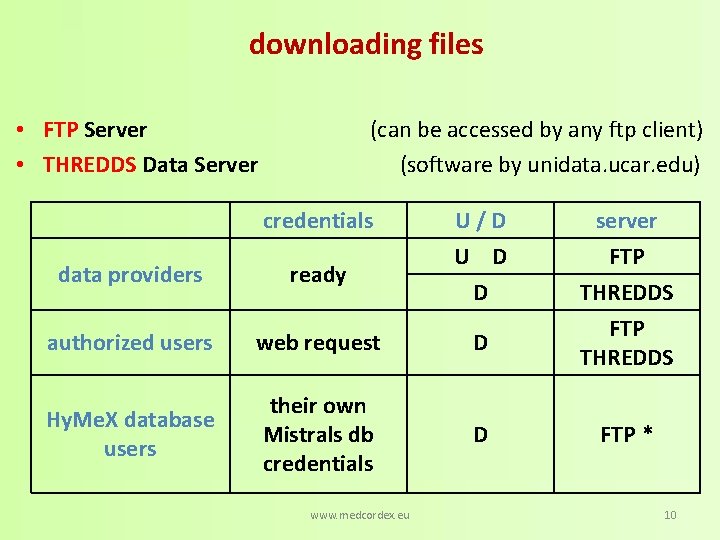
downloading files • FTP Server • THREDDS Data Server (can be accessed by any ftp client) (software by unidata. ucar. edu) credentials U/D U D D data providers ready authorized users web request D Hy. Me. X database users their own Mistrals db credentials D www. medcordex. eu server FTP THREDDS FTP * 10

downloading data (using any FTP client) cmd line: § ftp $f/$p/ ; dir ; get filen. nc “dir” not in /ALL § ncftp –u $hymex www. medcordex. eu ; cd $p ; get filen. nc § wget $f/$p/file. nc § wget -r $f/$p recursive get, not in /ALL browser: § $f/$p/filen. nc where: $f = ftp: //user: passw@www. medcordex. eu $p = MEDCORDEX/MED-xx/…/…/…. $p = MEDCORDEX/ALL www. medcordex. eu 11

downloading data (using THREDDS) services: § § § § (password required only to get netcdf files) Op. ENDAP HTTP server netcdf subset WCS WMS NCML ISO UDDC use files remotely , download them download files select & download sections of each file Web Coverage Service serves data to WCS clients Web Map Service serves data to WMS clients Net. CDF Markup Language to define a CDM ds description of the file in ISO 19115(-2) metadata. Unidata Attribute Convention for Data Discovery provides recommendations for net. CDF attributes that can be added to net. CDF files www. medcordex. eu 12

downloading data (using THREDDS) cmd line: § ncdump –h § cdo showdate § cdo copy § ferret: use $t/dods. C/$p/file. nc local. nc $t/dods. C/$p/file. nc tested with: netcdf 4. 3. 1. 1, cdo 1. 6. 4 rc 6, ferret 6. 9 browser: www. medcordex. eu/tds MEDCORDEX/ALL is invisible where: $p=MEDCORDEX/MED-xx/…/…/…. $p=MEDCORDEX/ALL $t=https: //user: passw@www. medcordex. eu: 8290/medcordex www. medcordex. eu 13

db fields for each ingested netcdf file are recorded: code path fname size ncdump realm Institution Variable. Name Domain GCMModel. Name CMIP 5 Experiment. Name CMIP 5 Ensemble. Member RCMModel. Name RCMVersion. ID RCMmodel Frequency Start. Time End. Time www. medcordex. eu 14
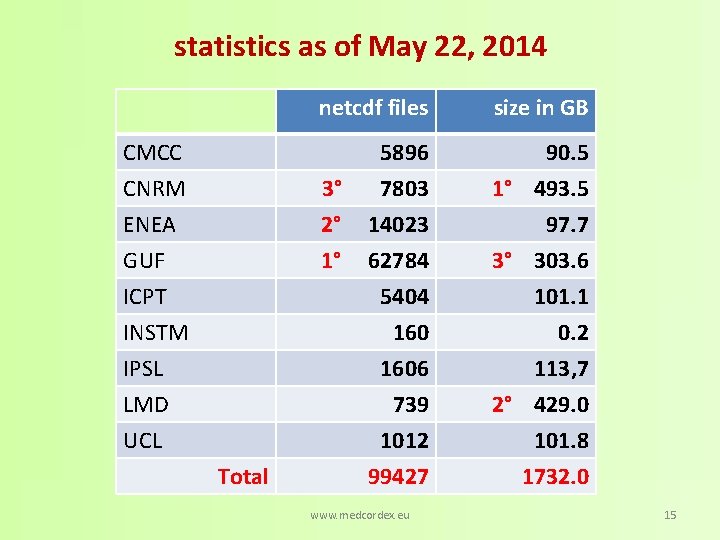
statistics as of May 22, 2014 CMCC CNRM ENEA netcdf files size in GB 5896 7803 14023 90. 5 1° 493. 5 97. 7 62784 5404 1606 739 1012 99427 3° 303. 6 101. 1 0. 2 113, 7 2° 429. 0 101. 8 1732. 0 3° 2° GUF ICPT INSTM IPSL LMD UCL 1° Total www. medcordex. eu 15
- Slides: 15