Mechanism of enzymes catalysis enzymesubstrate complex formation Coenzymes
Mechanism of enzymes catalysis, enzymesubstrate complex formation. Co-enzymes and its properties Dr. C. Mercy, Assistant professor, PG Department of zoology, Sarah Tucker College, Tirunelveli.

The mechanism of enzyme catalysis • In order for a reaction to occur, reactant molecules must contain sufficient energy to cross a potential energy barrier, the activation energy. • All molecules possess varying amounts of energy depending, for example, on their recent collision history but, generally, only a few have sufficient energy for reaction. • The lower the potential energy barrier to reaction, the more reactants have sufficient energy and, hence, the faster the reaction will occur. • All catalysts, including enzymes, function by forming a transition state, with the reactants, of lower free energy than would be found in the uncatalysed reaction. • Even quite modest reductions in this potential energy barrier may produce large increases in the rate of reaction.

• There a number of mechanisms by which this activation energy decrease may be achieved. • The most important of these involves the enzyme initially binding the substrate(s), in the correct orientation to react, close to the catalytic groups on the active enzyme complex and any other substrates. • In this way the binding energy is used partially in order to reduce the contribution of the considerable activation entropy, due to the loss of the reactants' (and catalytic groups') translational and rotational entropy, towards the total activation energy.

Enzyme catalysis: • Enzyme catalysis is the increase in the rate of a chemical reaction by the active site of a protein. • The protein catalyst(enzyme) may be part of a multi-subunit complex, and/or may transiently or permanently associate with a Cofactor (e. g. adenosine triphosphate). • Catalysis of biochemical reactions in the cell is vital due to the very low reaction rates of the uncatalysed reactions at room temperature and pressure. • A key driver of protein evolution is the optimization of such catalytic activities via protein dynamics • The mechanism of enzyme catalysis is similar in principle to other types of chemical catalysis. • By providing an alternative reaction route the enzyme reduces the energy required to reach the highest energy transition state of the reaction.
• The reduction of activation energy increases the amount of reactant molecules that achieve a sufficient level of energy, such that they reach the activation energy and form the product. • As with other catalysts, the enzyme is not consumed during the reaction (as a substrate is) but is recycled such that a single enzyme performs many rounds of catalysis.

• Enzyme- substrate complex formation: • Enzyme–substrate complex : The intermediate formed when a substrate molecule interacts with the active site of an enzyme. • Following the formation of an enzyme–substrate complex, the substrate molecule undergoes a chemical reaction and is converted into a new product. • Various mechanisms for the formation of enzyme–substrate complexes have been suggested, including the induced-fit modeland the lock-and-key mechanism • Enzyme Kinetics: The Enzyme Substrate Complex • A theory to explain the catalytic action of enzymes was proposed by the Swedish chemist Savante Arrhenius in 1888. He proposed that the substrate and enzyme formed some intermediate substance which is known as the enzyme substrate complex. The reaction can be represented as:
![• If this reaction is combined with the original reaction equation [1], the • If this reaction is combined with the original reaction equation [1], the](http://slidetodoc.com/presentation_image_h2/db5a04ddc753e91e67157e46d8db354d/image-7.jpg)
• If this reaction is combined with the original reaction equation [1], the following results: • The existence of an intermediate enzyme-substrate complex has been demonstrated in the laboratory, for example, using catalase and a hydrogen peroxide derivative. • At Yale University, Kurt G. Stern observed spectral shifts in catalase as the reaction it catalyzed proceeded. • This experimental evidence indicates that the enzyme first unites in some way with the substrate and then returns to its original form after the reaction is concluded.
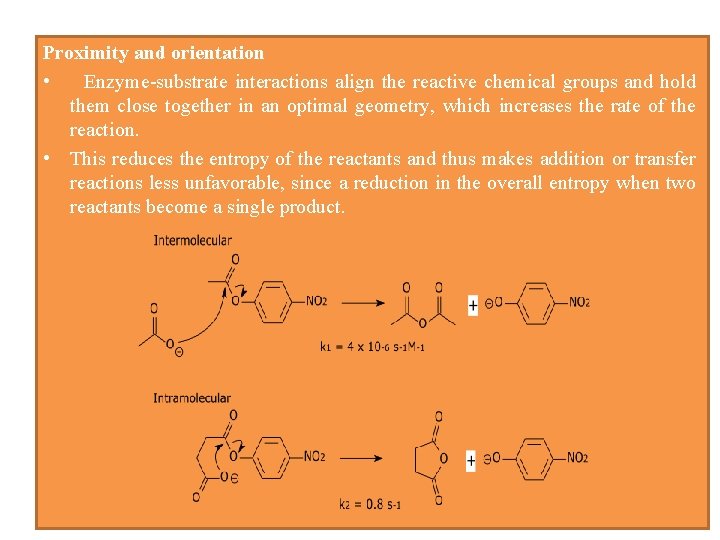
Proximity and orientation • Enzyme-substrate interactions align the reactive chemical groups and hold them close together in an optimal geometry, which increases the rate of the reaction. • This reduces the entropy of the reactants and thus makes addition or transfer reactions less unfavorable, since a reduction in the overall entropy when two reactants become a single product.
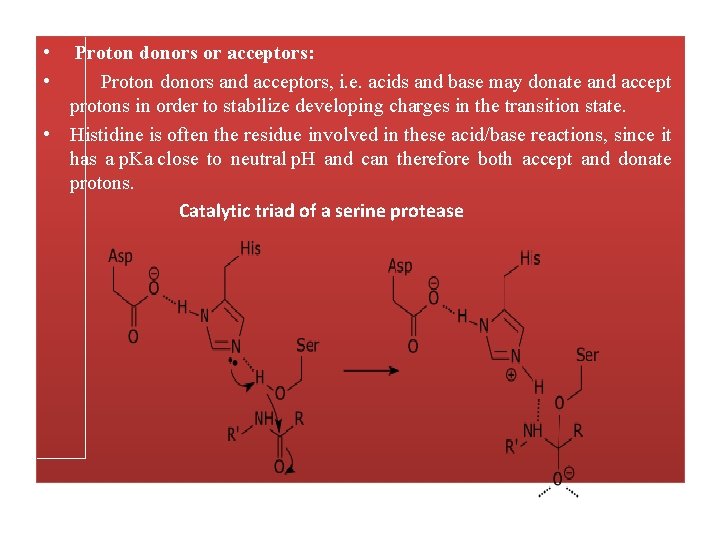
• • Proton donors or acceptors: Proton donors and acceptors, i. e. acids and base may donate and accept protons in order to stabilize developing charges in the transition state. • Histidine is often the residue involved in these acid/base reactions, since it has a p. Ka close to neutral p. H and can therefore both accept and donate protons. Catalytic triad of a serine protease

• Electrostatic catalysis: • Stabilization of charged transition states can also be by residues in the active site forming ionic bonds (or partial ionic charge interactions) with the intermediate. • These bonds can either come from acidic or basic side chains found on amino acids such as lysine, arginine, aspartic acid or glutamic acid or come from metal cofactors such as zinc. • Metal ions are particularly effective and can reduce the p. Ka of water enough to make it an effective nucleophile. • This is very different from transition state stabilization in water, where the water molecules must pay with "reorganization energy". • In order to stabilize ionic and charged states. Thus, the catalysis is associated with the fact that the enzyme polar groups are preorganized. • The magnitude of the electrostatic field exerted by an enzyme's active site has been shown to be highly correlated with the enzyme's catalytic rate enhancement
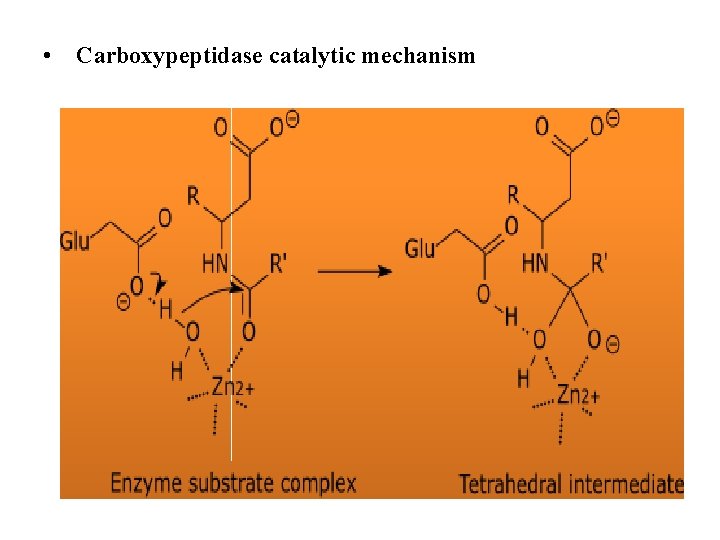
• Carboxypeptidase catalytic mechanism

Covalent catalysis: • Covalent catalysis involves the substrate forming a transient covalent bond with residues in the enzyme active site or with a cofactor. • This adds an additional covalent intermediate to the reaction, and helps to reduce the energy of later transition states of the reaction. • The covalent bond must, at a later stage in the reaction, be broken to regenerate the enzyme. • This mechanism is utilised by the catalytic triad of enzymes such as proteases like chymotrypsin and trypsin, where an acyl-enzyme intermediate is formed. • An alternative mechanism is schiff base formation using the free amine from a lysine residue, as seen in the enzyme aldolase during glycolysis. • Some enzymes utilize non-amino acid cofactors such as pyridoxal phosphate (PLP) or thiamine pyrophosphate (TPP) to form covalent intermediates with reactant molecules.

Metal ion catalysis: • A metal ion in the active site participates in catalysis by coordinating charge stabilization and shielding. • Because of a metal's positive charge, only negative charges can be stabilized through metal ions. • However, metal ions are advantageous in biological catalysis because they are not affected by changes in p. H. • Metal ions can also act to ionize water by acting as a Lewis acid. Metal ions may also be agents of oxidation and reduction. Substrate, bound substrate, and transition state conformations of lysozyme • • The substrate, on binding, is distorted from the half chair conformation of the hexose ring (because of the steric hindrance with amino acids of the protein forcing the equatorial c 6 to be in the axial position) into the chair conformation

Quantum tunneling: • These traditional "over the barrier" mechanisms have been challenged in some cases by models and observations of "through the barrier" mechanisms (quantum tunneling). • Some enzymes operate with kinetics which is faster than what would be predicted by the classical ΔG. • In "through the barrier" models, a proton or an electron can tunnel through activation barriers. Quantum tunneling for protons has been observed in tryptamine oxidation by aromatic amine dehydrogenase. Active enzyme: • The binding energy of the enzyme-substrate complex cannot be considered as an external energy which is necessary for the substrate activation. • Examples of catalytic mechanisms: • In reality, most enzyme mechanisms involve a combination of several different types of catalysis.

Triose phosphate isomerase: • Triose phosphate isomerase (EC 5. 3. 1. 1) catalyses the reversible interconvertion of the two triose phosphates isomers dihydroxyacetone phosphate and D-glyceraldehyde 3 -phosphate. Trypsin: Trypsin (EC 3. 4. 21. 4) is a serine protease that cleaves protein substrates after lysine or arginine residues using a catalytic triad to perform covalent catalysis, and an oxyanion holeto stabilise charge-buildup on the transition states. Aldolase: • Aldolase (EC 4. 1. 2. 13) catalyses the breakdown of fructose 1, 6 bisphosphate (F-1, 6 -BP) into glyceraldehyde 3 phosphate and dihydroxyacetone phosphate (DHAP).

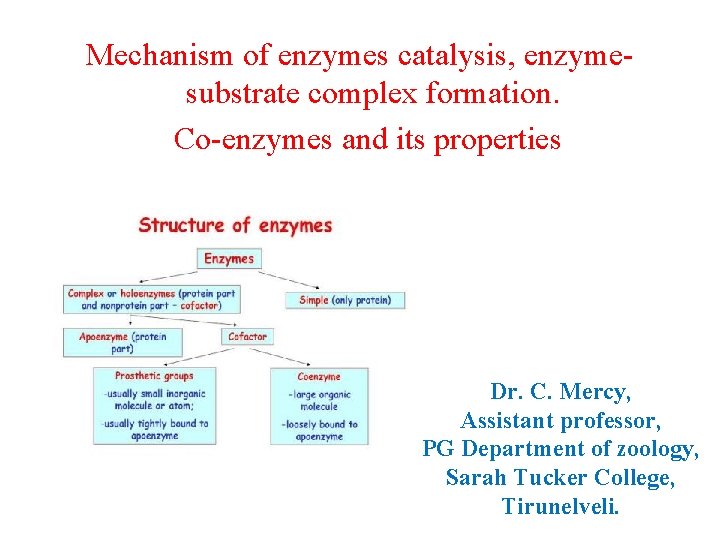
• Co-enzymes and its properties: Definition • A coenzyme is an organic non-protein compound that binds with an enzyme to catalyze a reaction. • Coenzymes are often broadly called cofactors, but they are chemically different. • A coenzyme cannot function alone, but can be reused several times when paired with an enzyme. Functions of Coenzymes: • An enzyme without a coenzyme is called an apoenzyme. • Without coenzymes or cofactors, enzymes cannot catalyze reactions effectively. • In fact, the enzyme may not function at all. If reactions cannot occur at the normal catalyzed rate, then an organism will have difficulty sustaining life. • When an enzyme gains a coenzyme, it then becomes a holoenzyme, or active enzyme. • Active enzymes change substrates into the products an organism needs to carry out essential functions, whether chemical or physiological.

• Coenzymes, like enzymes, can be reused and recycled without changing reaction rate or effectiveness. They attach to a portion of the active site on an enzyme, which enables the catalyzed reaction to occur. • When an enzyme is denatured by extreme temperature or p. H, the coenzyme can no longer attach to the active site. Types of Enzymes: • Cofactors are molecules that attach to an enzyme during chemical reactions. In general, all compounds that help enzymes are called cofactors. • However, cofactors can be broken down into three subgroups based on chemical makeup and function: Coenzymes: • These are reusable non-protein molecules that contain carbon (organic). • They bind loosely to an enzyme at the active site to help catalyze reactions. • Most are vitamins, vitamin derivatives, or form from nucleotides.

Cofactors: • Unlike coenzymes, true cofactors are reusable non-protein molecules that do not contain carbon (inorganic). • Usually, cofactors are metal ions such as iron, zinc, cobalt, and copper that loosely binds to an enzyme’s active site. • They must also be supplemented in the diet as most organisms do not naturally synthesize metal ions. • Prosthetic groups: • These can be organic vitamins, sugars, lipids, or inorganic metal ions. • However, unlike coenzymes or cofactors, these groups bind very tightly or covalently to an enzyme to aid in catalyzing reactions. • These groups are often used in cellular respiration and photosynthesis.

THANK YOU
- Slides: 19