Measuring the Randomness of Data via Multiscale Bootstrap

Measuring the Randomness of Data via Multiscale Bootstrap with Applications to Evolutionary Tree Inference Hidetoshi Shimodaira Tokyo Inst. Tech. talk outline Bootstrap resampling method Phylogenetic inference Testing procedures Multiscale bootstrap (approximately unbiasd test) • Asymptotic theory of “nearly flat surface” • •
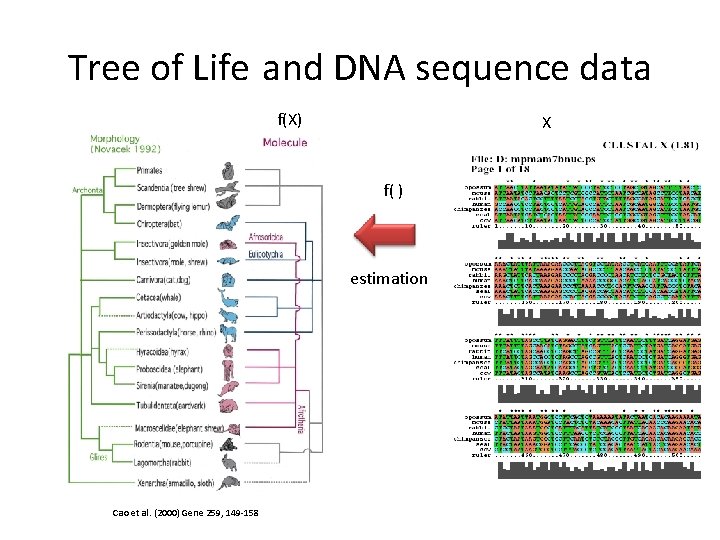
Tree of Life and DNA sequence data f(X) X f( ) estimation Cao et al. (2000) Gene 259, 149 -158

“The problem of regions” Efron and Tibshirani (1998) (example) Model Selection of Polynomial Regression 0: constant, 1: linear, 2: quadratic 4
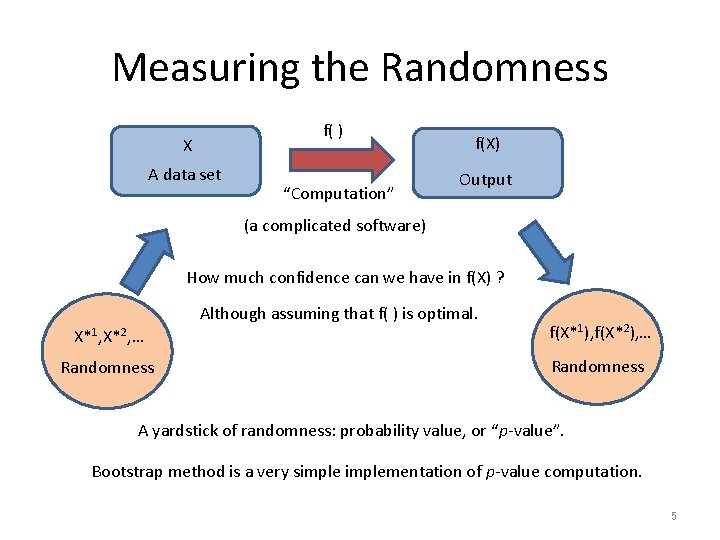
Measuring the Randomness f( ) X A data set “Computation” f(X) Output (a complicated software) How much confidence can we have in f(X) ? Although assuming that f( ) is optimal. X*1, X*2, … f(X*1), f(X*2), … Randomness A yardstick of randomness: probability value, or “p-value”. Bootstrap method is a very simplementation of p-value computation. 5
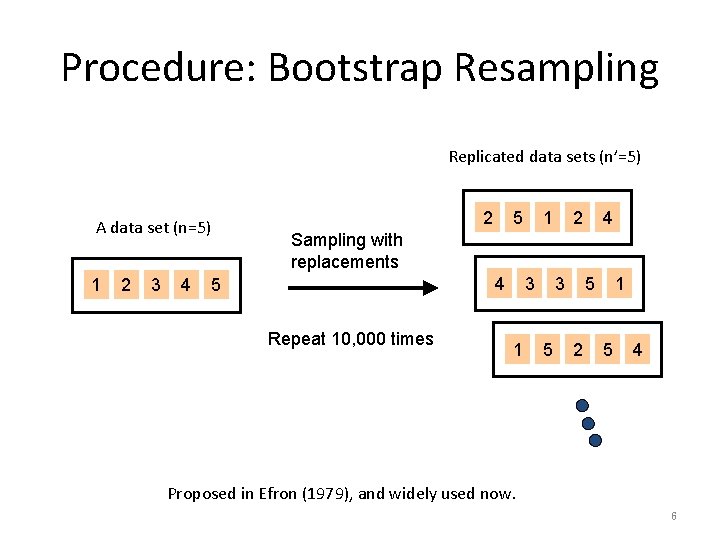
Procedure: Bootstrap Resampling Replicated data sets (n’=5) A data set (n=5) 1 2 3 4 2 5 1 2 4 Sampling with replacements 4 5 Repeat 10, 000 times 3 1 3 5 5 2 1 5 4 Proposed in Efron (1979), and widely used now. 6
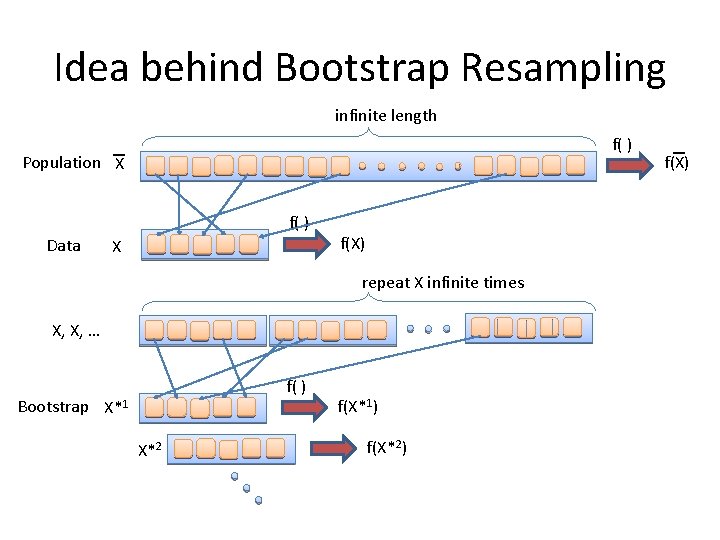
Idea behind Bootstrap Resampling infinite length f( ) Population X f( ) Data X f(X) repeat X infinite times X, X, … f( ) Bootstrap X*1 X*2 f(X*1) f(X*2) f(X)
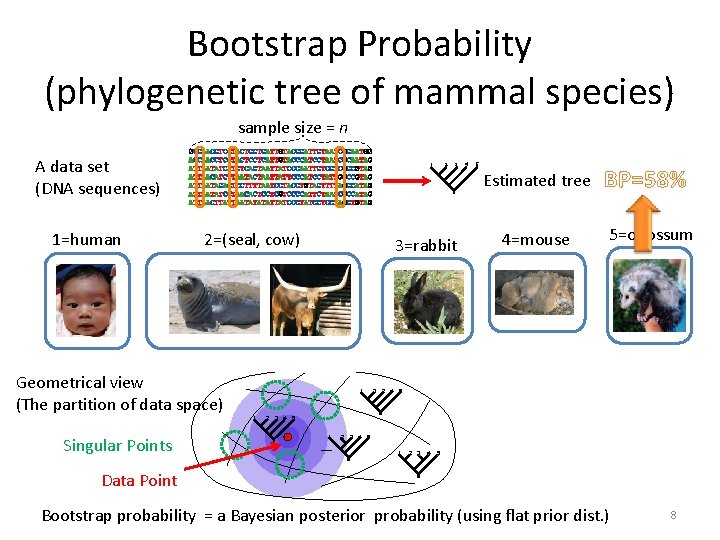
Bootstrap Probability (phylogenetic tree of mammal species) sample size = n A data set (DNA sequences) 1=human Data sets replicated 10, 000 times 1 3 4 5 Estimated tree n’ 2=(seal, cow) 1 2 3 4 5 3=rabbit 1 1 2 3 4 2 1 1 3 4 4=mouse 5 5 1 Geometrical view (The partition of data space) Singular Points 2 3 2 4 5 BP=58% 5=opossum Frequencies of Trees 3 2 4 5 2 3 4 5 1 2 3 4 5 Data Point Bootstrap probability = a Bayesian posterior probability (using flat prior dist. ) 8
Shimodaira
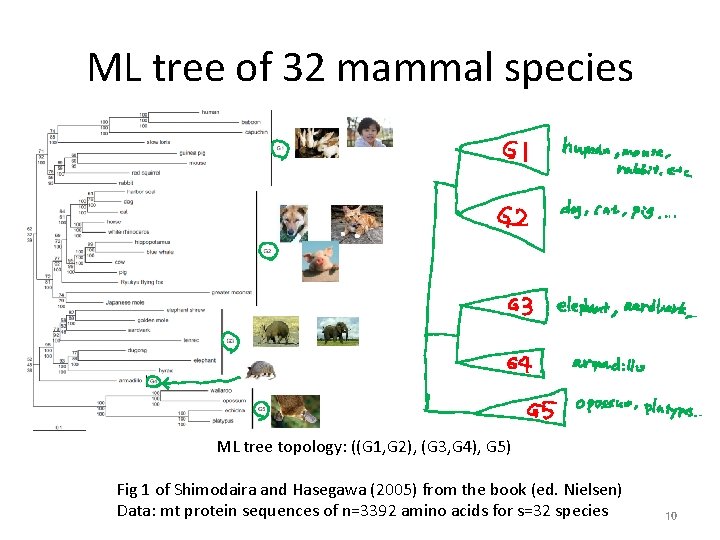
ML tree of 32 mammal species ML tree topology: ((G 1, G 2), (G 3, G 4), G 5) Fig 1 of Shimodaira and Hasegawa (2005) from the book (ed. Nielsen) Data: mt protein sequences of n=3392 amino acids for s=32 species 10

Data: mt protein sequences 11
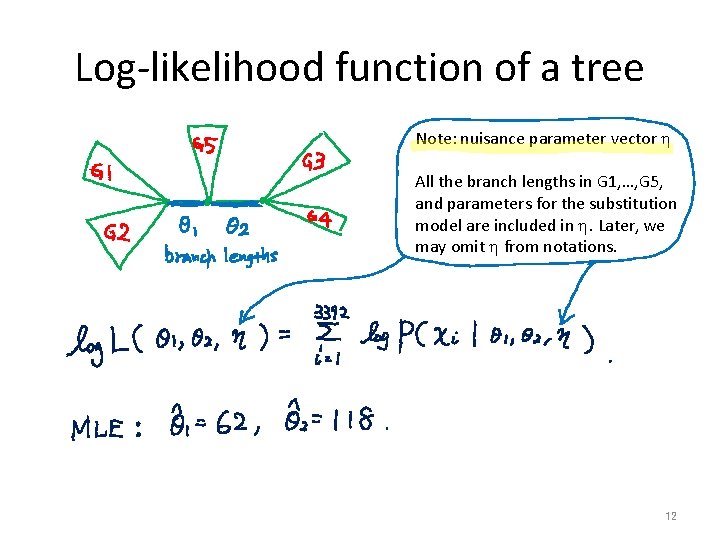
Log-likelihood function of a tree Note: nuisance parameter vector h All the branch lengths in G 1, …, G 5, and parameters for the substitution model are included in h. Later, we may omit h from notations. 12
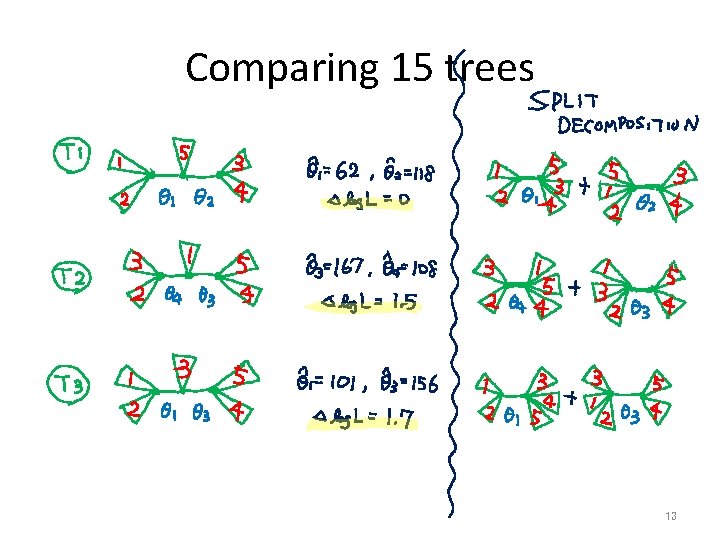
Comparing 15 trees 13

Comparing 15 trees (cont. ) 14

Likelihood Ratio Test (each tree v. s. full model) Rejects all the possible tree topologies! => Model is misspecified 15
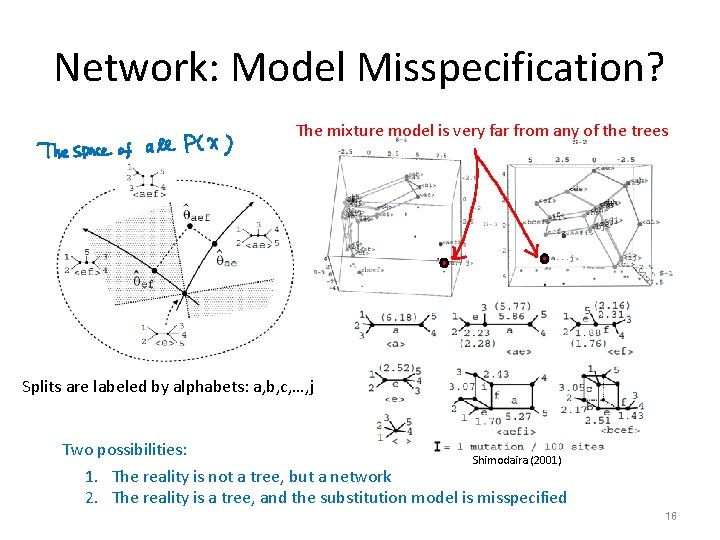
Network: Model Misspecification? The mixture model is very far from any of the trees Splits are labeled by alphabets: a, b, c, …, j Two possibilities: Shimodaira (2001) 1. The reality is not a tree, but a network 2. The reality is a tree, and the substitution model is misspecified 16

Phylogenetic inference: Which methodology should we use? Measure of confidence Hypotheses Correct (Traditional framework) Frequentist p-value (“modern” theory) Bayesian posterior probability (250 years old theory) Posterior probability of tree (often implemented via MCMC) Likelihood ratio test Cox test Kishino-Hasegawa test Best (Akaike’s framework) Shimodaira-Hasegawa test Bootstrap probability Approximately unbiased test (via multiscale bootstrap) bias adjustment 17
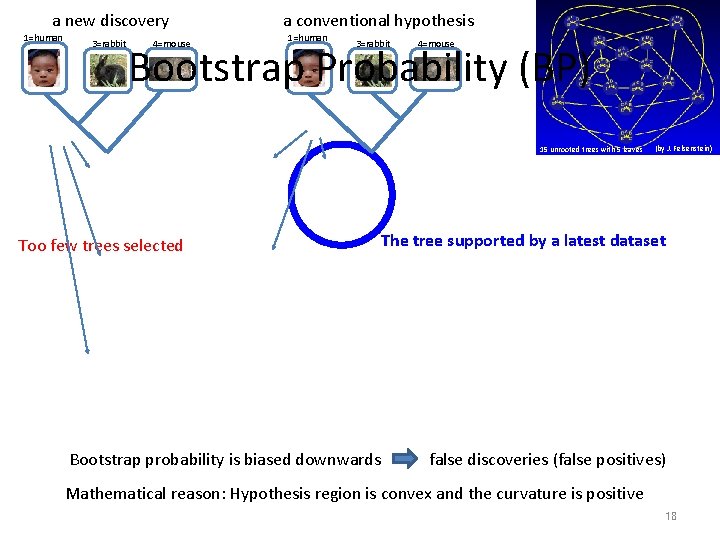
a new discovery 1=human 3=rabbit 4=mouse a conventional hypothesis 1=human 3=rabbit 4=mouse Bootstrap Probability (BP) 15 unrooted trees with 5 leaves Too few trees selected (by J. Felsenstein) The tree supported by a latest dataset Bootstrap probability is biased downwards false discoveries (false positives) Mathematical reason: Hypothesis region is convex and the curvature is positive 18

Shimodaira-Hasegawa test (a multiple comparisons procedure) Too many trees selected The tree supported by a latest dataset SH-test is biased downwards false undiscovery (false negatives) Mathematical reason: Multiple comparisons method evaluates the worst-case scenario and is conservative 19

Multiscale Bootstrap (AU test) Moderate number of trees selected The tree supported by a latest dataset Approximately Unbiased (third-order asymptotic accuracy for smooth surfaces) 20
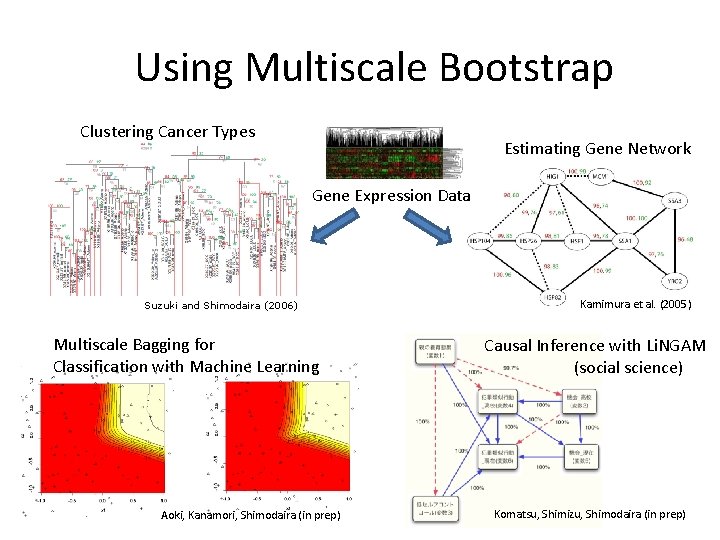
Using Multiscale Bootstrap Clustering Cancer Types Estimating Gene Network Gene Expression Data Suzuki and Shimodaira (2006) Multiscale Bagging for Classification with Machine Learning Aoki, Kanamori, Shimodaira (in prep) Kamimura et al. (2005) Causal Inference with Li. NGAM (social science) Komatsu, Shimizu, Shimodaira (in prep)
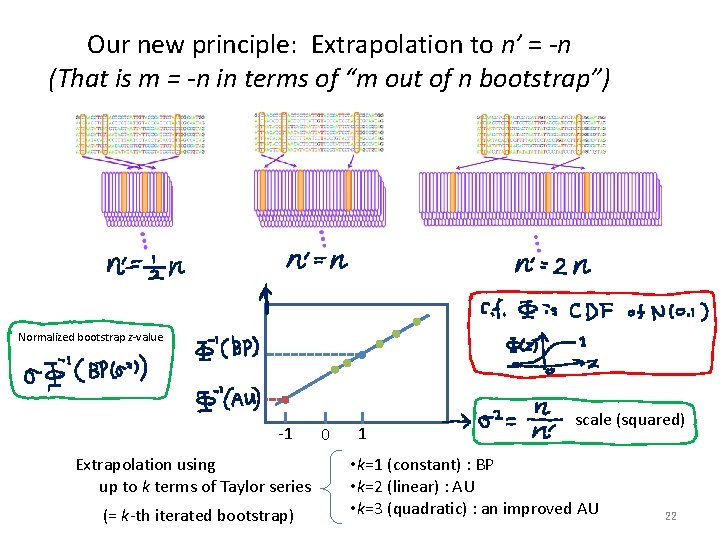
Our new principle: Extrapolation to n’ = -n (That is m = -n in terms of “m out of n bootstrap”) Normalized bootstrap z-value -1 Extrapolation using up to k terms of Taylor series (= k-th iterated bootstrap) 0 1 scale (squared) • k=1 (constant) : BP • k=2 (linear) : AU • k=3 (quadratic) : an improved AU 22
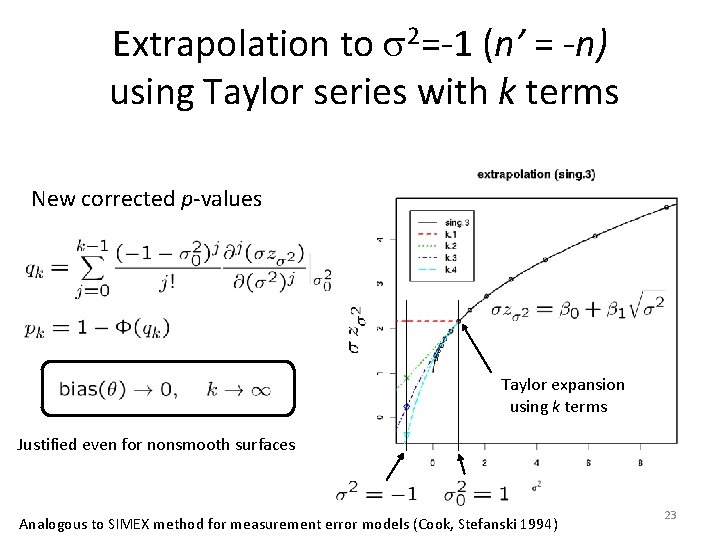
Extrapolation to s 2=-1 (n’ = -n) using Taylor series with k terms New corrected p-values Taylor expansion using k terms Justified even for nonsmooth surfaces Analogous to SIMEX method for measurement error models (Cook, Stefanski 1994) 23

Mammal phylogeny p-values are computed by “scaleboot” t 1: ML tree t 4: “True tree” d: the distance to the boundary surface 24

What does bootstrap do? (multivariate normal model)
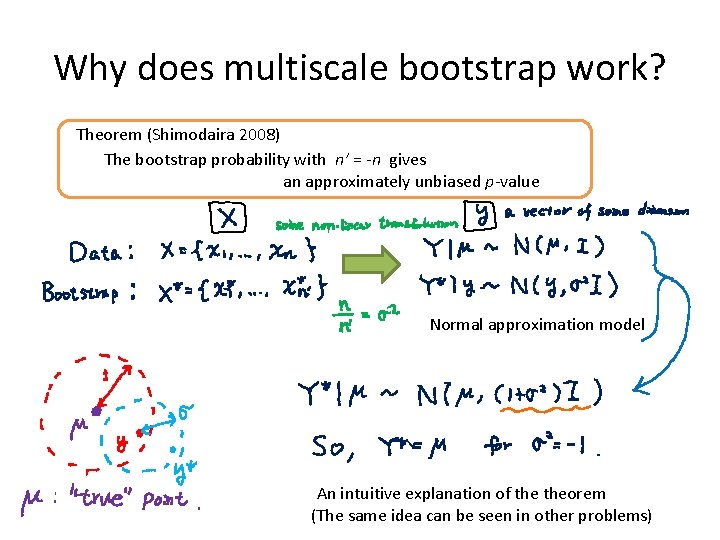
Why does multiscale bootstrap work? Theorem (Shimodaira 2008) The bootstrap probability with n’ = -n gives an approximately unbiased p-value Normal approximation model An intuitive explanation of theorem (The same idea can be seen in other problems)

What is an “unbiased test” ? The rejection probability is equal to the significance level (e. g. , a=0. 05) on the boundary of the null hypothesis. This is a frequentist idea

Flat Boundary Surface (or one dim. ) Bayesian (s=1) Frequentist
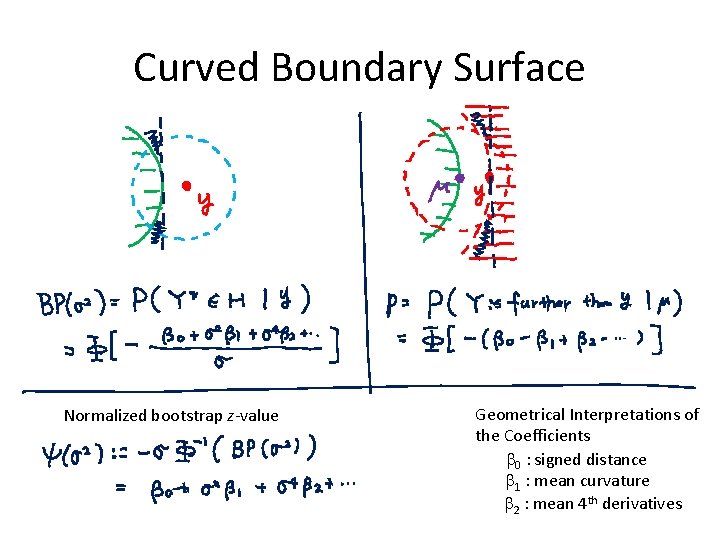
Curved Boundary Surface Normalized bootstrap z-value Geometrical Interpretations of the Coefficients b 0 : signed distance b 1 : mean curvature b 2 : mean 4 th derivatives
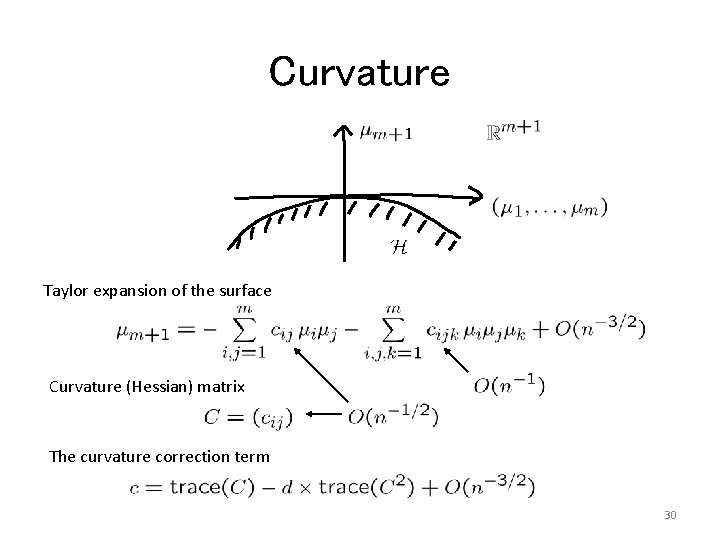
Curvature Taylor expansion of the surface Curvature (Hessian) matrix The curvature correction term 30
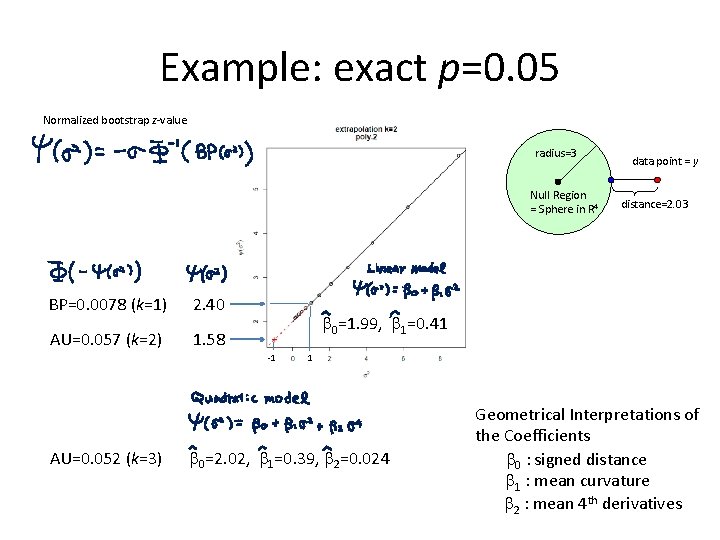
Example: exact p=0. 05 Normalized bootstrap z-value radius=3 Null Region = Sphere in R 4 BP=0. 0078 (k=1) 2. 40 AU=0. 057 (k=2) 1. 58 distance=2. 03 b 0=1. 99, b 1=0. 41 -1 AU=0. 052 (k=3) data point = y 1 b 0=2. 02, b 1=0. 39, b 2=0. 024 Geometrical Interpretations of the Coefficients b 0 : signed distance b 1 : mean curvature b 2 : mean 4 th derivatives
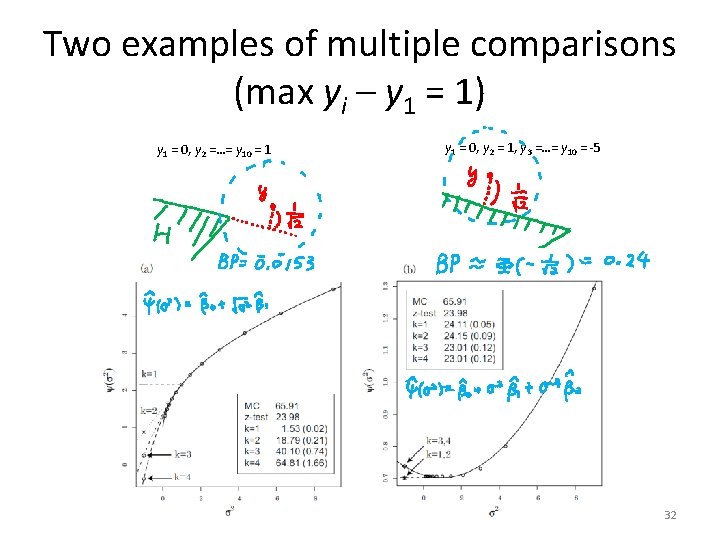
Two examples of multiple comparisons (max yi – y 1 = 1) y 1 = 0, y 2 =…= y 10 = 1 y 1 = 0, y 2 = 1, y 3 =…= y 10 = -5 32
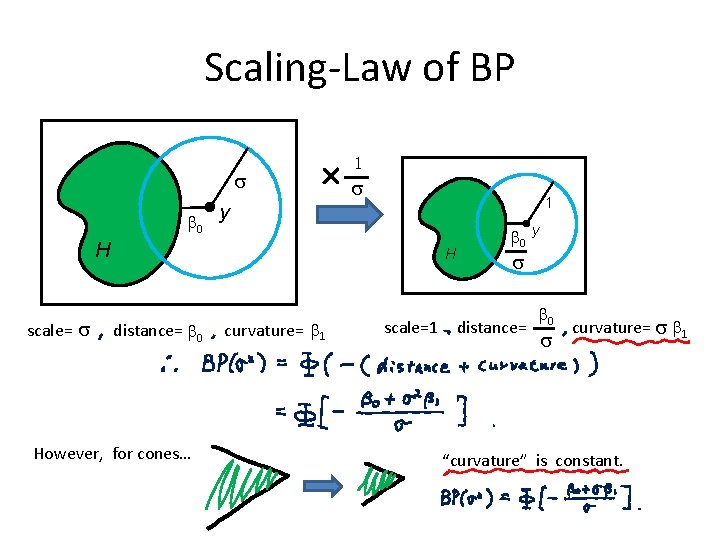
Scaling-Law of BP s b 0 y 1 s 1 H scale= s H distance= b 0 However, for cones… curvature= b 1 scale=1 b 0 y s distance= b 0 s curvature= s b 1 “curvature” is constant.
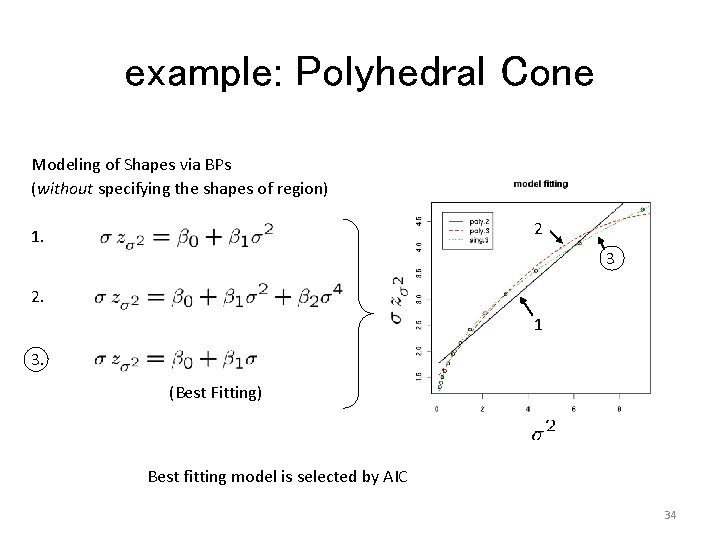
example: Polyhedral Cone Modeling of Shapes via BPs (without specifying the shapes of region) 2 1. 3 2. 1 3. (Best Fitting) Best fitting model is selected by AIC 34
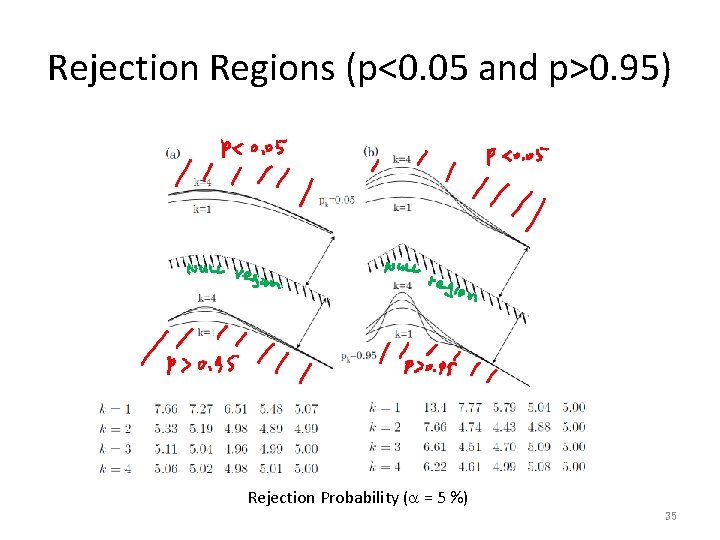
Rejection Regions (p<0. 05 and p>0. 95) Rejection Probability (a = 5 %) 35
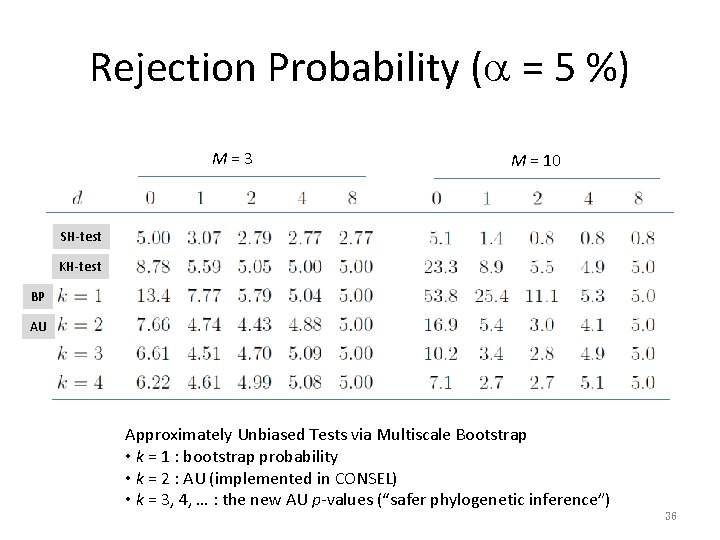
Rejection Probability (a = 5 %) M=3 M = 10 SH-test KH-test BP AU Approximately Unbiased Tests via Multiscale Bootstrap • k = 1 : bootstrap probability • k = 2 : AU (implemented in CONSEL) • k = 3, 4, … : the new AU p-values (“safer phylogenetic inference”) 36
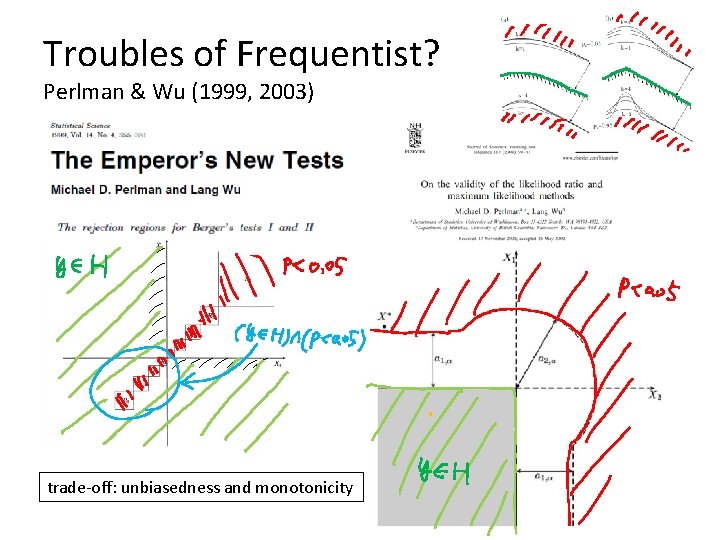
Troubles of Frequentist? Perlman & Wu (1999, 2003) trade-off: unbiasedness and monotonicity
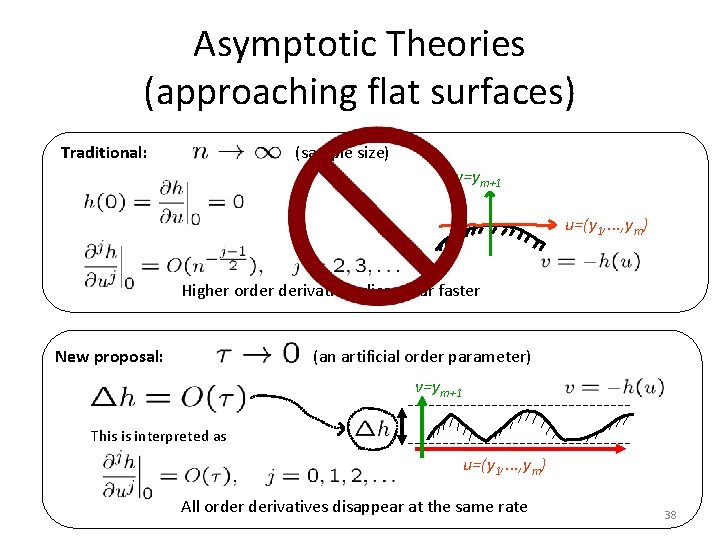
Asymptotic Theories (approaching flat surfaces) Traditional: (sample size) v=ym+1 u=(y 1, . . . , ym) Higher order derivatives disappear faster New proposal: (an artificial order parameter) v=ym+1 This is interpreted as u=(y 1, . . . , ym) All order derivatives disappear at the same rate 38

Nearly Flat Surfaces Three conditions 1. (i. e. , approaches a flat surface) 2. Fourier transform: for 3. 39
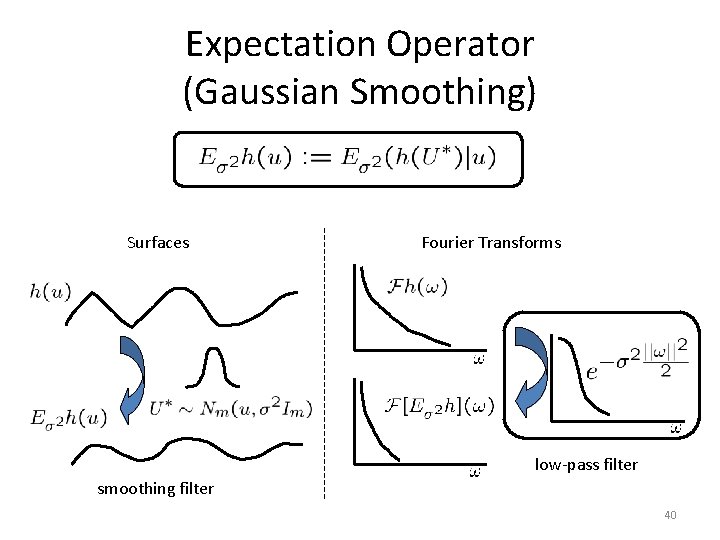
Expectation Operator (Gaussian Smoothing) Surfaces Fourier Transforms low-pass filter smoothing filter 40
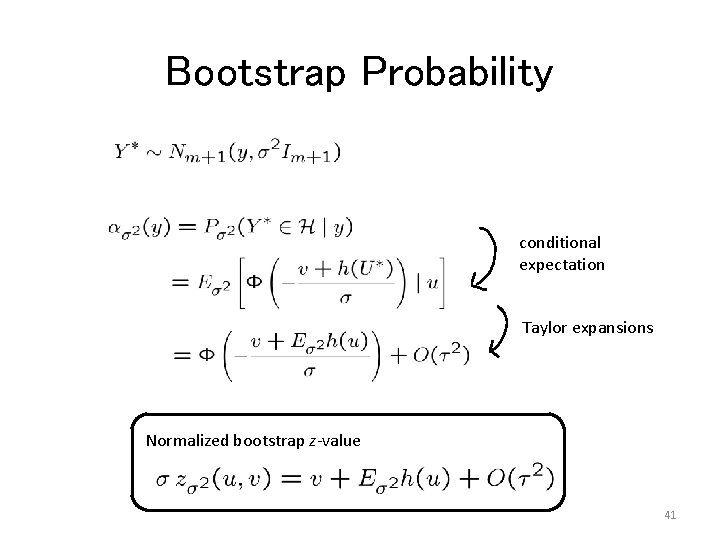
Bootstrap Probability conditional expectation Taylor expansions Normalized bootstrap z-value 41

Models for Curve Fitting 1. Smooth Surfaces: parameters: 2. Cone Surfaces: parameters: 42
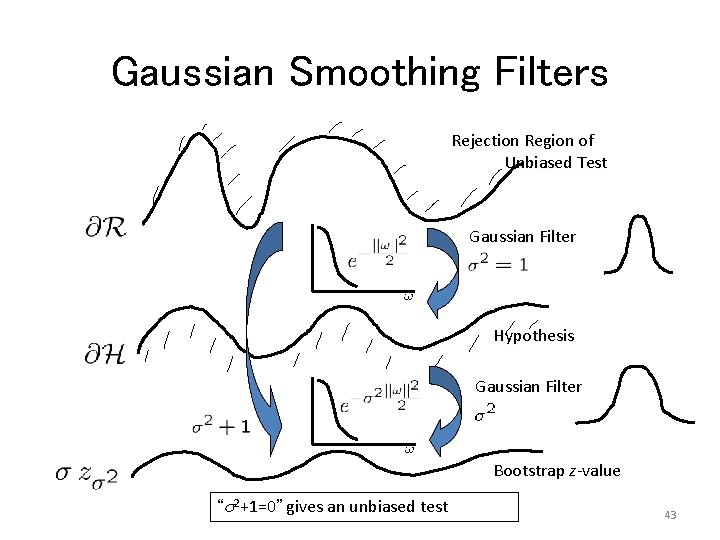
Gaussian Smoothing Filters Rejection Region of Unbiased Test Gaussian Filter Hypothesis Gaussian Filter Bootstrap z-value “s 2+1=0” gives an unbiased test 43

Unbiased Tests But, the extrapolation does not exist for nonsmooth surfaces. 44

Our Method and Generalization Our corrected p-values are represented as: …(*) Generalization: (*) defines a new p-value from a given Jk(w) For our method, Jk(w) is defined by 45

A Class of Approximately Unbiased Tests Our pk satisfies the following conditions (i)-(iv). the condition for unbiasedness pk is unbiased when h is polynomial of degree 2 k-1 46
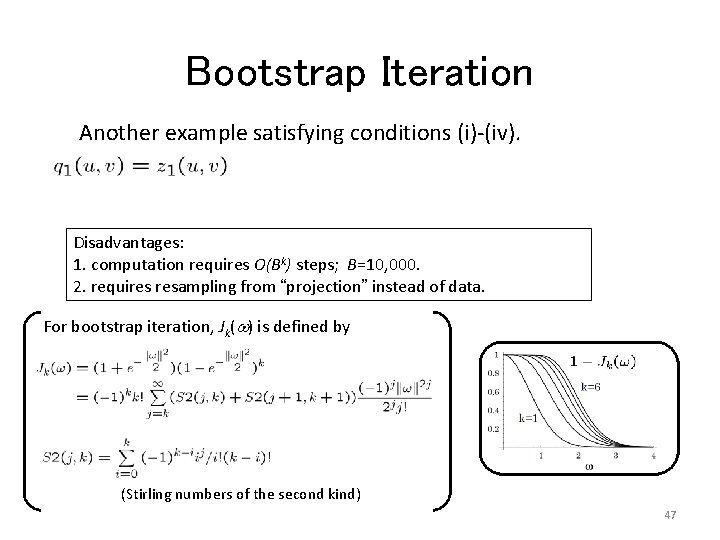
Bootstrap Iteration Another example satisfying conditions (i)-(iv). Disadvantages: 1. computation requires O(Bk) steps; B=10, 000. 2. requires resampling from “projection” instead of data. For bootstrap iteration, Jk(w) is defined by (Stirling numbers of the second kind) 47

Frequentist and Bayesian-like p-values Bootstrap Probability Frequentist unbiased p-value Bayesian-like p-value 48
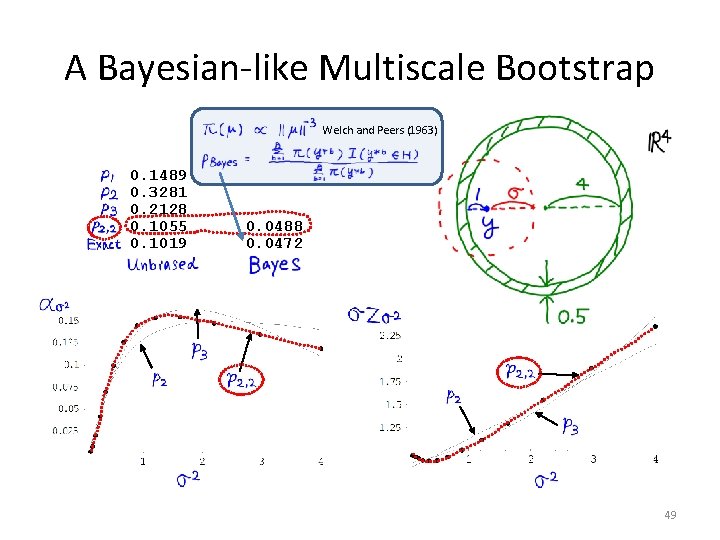
A Bayesian-like Multiscale Bootstrap Welch and Peers (1963) 0. 1489 0. 3281 0. 2128 0. 1055 0. 1019 0. 0488 0. 0472 49
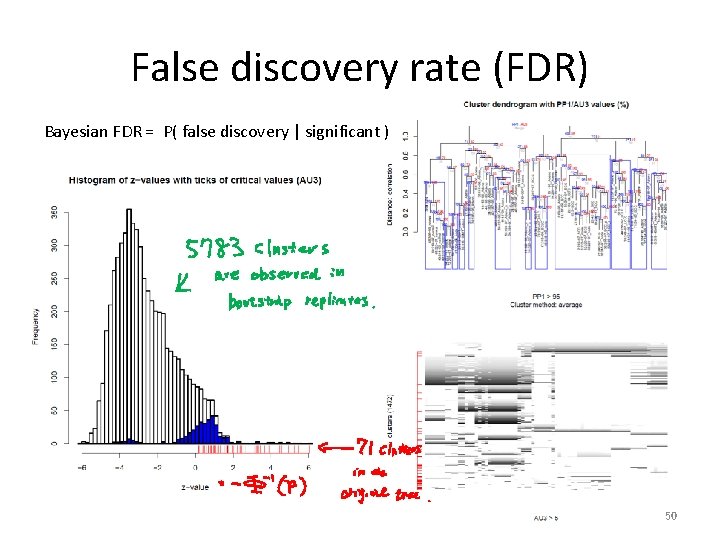
False discovery rate (FDR) Bayesian FDR = P( false discovery | significant ) 50
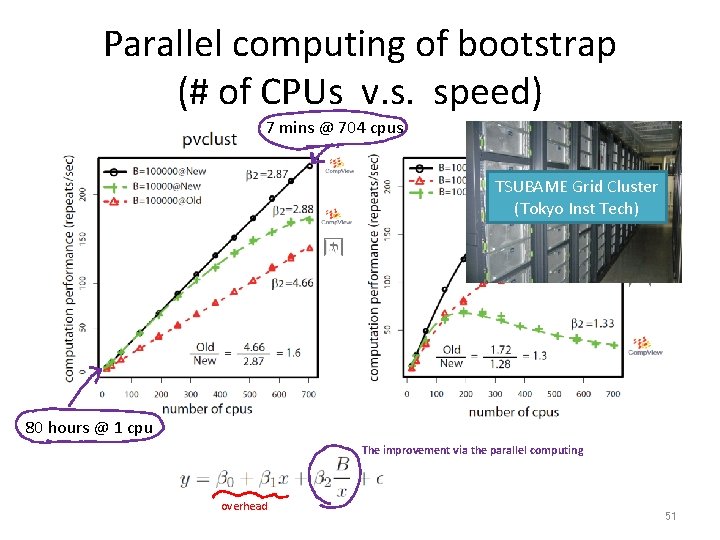
Parallel computing of bootstrap (# of CPUs v. s. speed) 7 mins @ 704 cpus TSUBAME Grid Cluster (Tokyo Inst Tech) 80 hours @ 1 cpu The improvement via the parallel computing overhead 51

Summary • The new principle (extrapolation to n’=-n) is proposed • AU is safer than BP, but AU is liberal than MC (BP < AU < MC) • The software “scaleboot” is available from CRAN • Exactly unbiased tests do not exist anyway for singular surfaces • We do not know yet the right method of inductive inference even after 250 years of investigations since Thomas Bayes • Work in progress of myself: a Bayesian-like multiscale bootstrap and FDR issues 52
- Slides: 51