MCB 317 Genetics and Genomics MCB 317 Topic

MCB 317 Genetics and Genomics MCB 317 Topic 10, part 5 A Story of Transcription
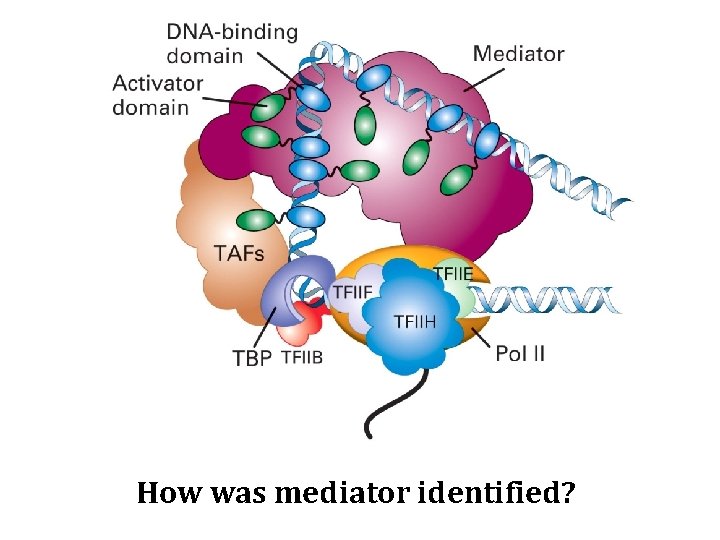
How was mediator identified?
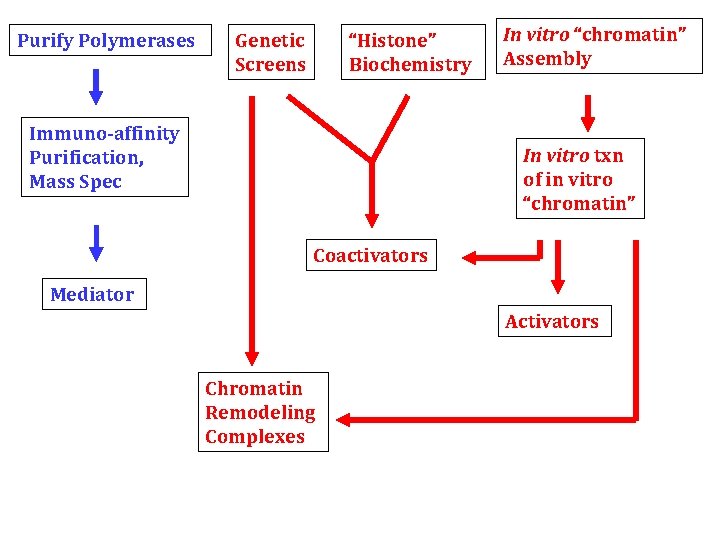
Purify Polymerases Genetic Screens “Histone” Biochemistry Immuno-affinity Purification, Mass Spec In vitro “chromatin” Assembly In vitro txn of in vitro “chromatin” Coactivators Mediator Activators Chromatin Remodeling Complexes

RNAPs Purified Based on in vivo txn of naked genomic DNA- nonpecific synthesis of RNA, but… … is the “structure” (subunit composition) of RNAPII the same in vivo as defined in vitro?

Hypothesis: Steps involved in purification of RNAPs may have dissociated some subunits. Test: “Purify” RNAPII by the most gentle method possible Method: Immunoaffinity purification and Immunoprecipitation from crude extracts
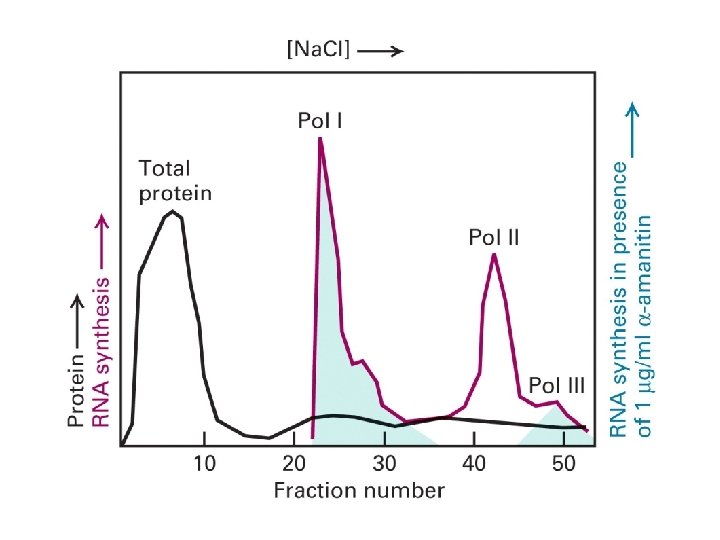
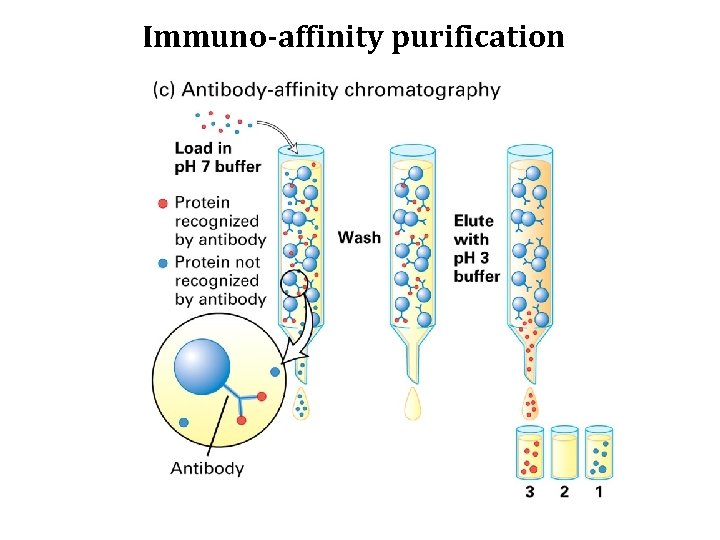
Immuno-affinity purification
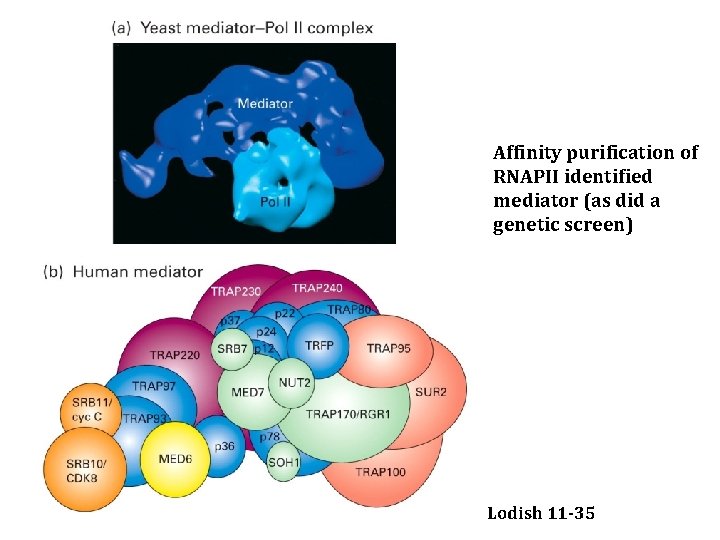
Affinity purification of RNAPII identified mediator (as did a genetic screen) Lodish 11 -35
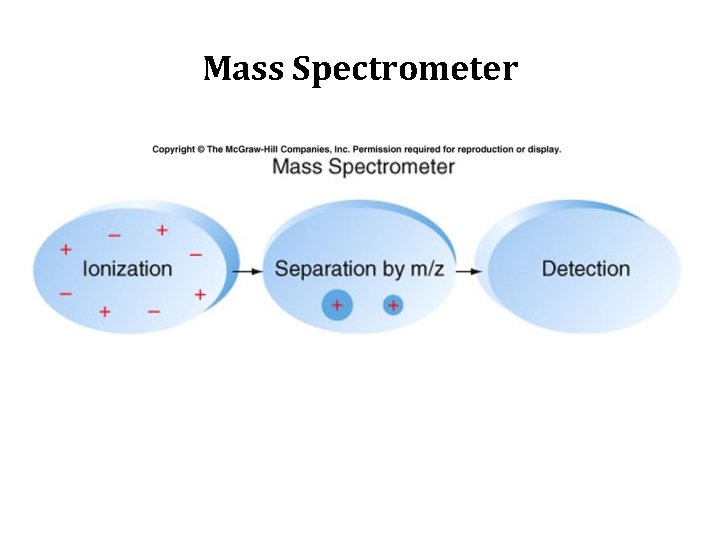
Mass Spectrometer

Mass Spectrometer Two spectrometers working in tandem 1. Separate “large” fragments of proteins 2. Those fragments analyzed by a second spectrometer -> masses of peptides 3. Masses of peptides = sequence of peptide fragments 4. Computer compares sequence of peptide fragments with predicted products of genes in genome to identify the gene that encodes the protein

One Subunit -> Complex Protein -> Immuno-affinity purification -> mass spec -> genome database -> genes that encode subunits of a complex
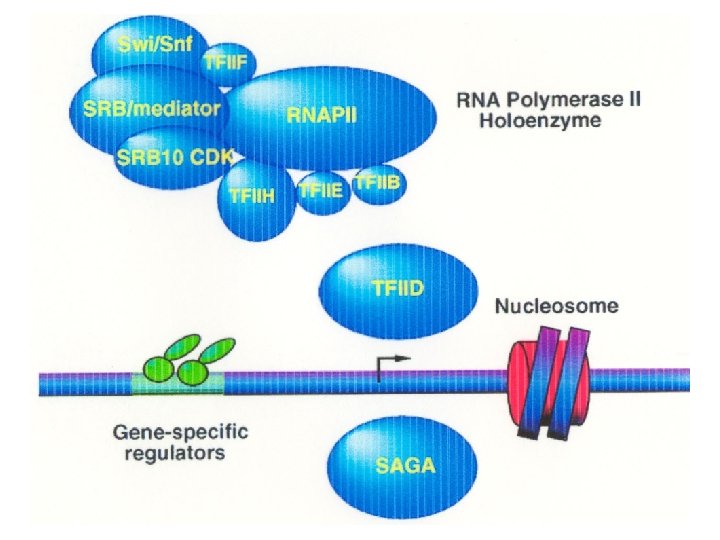
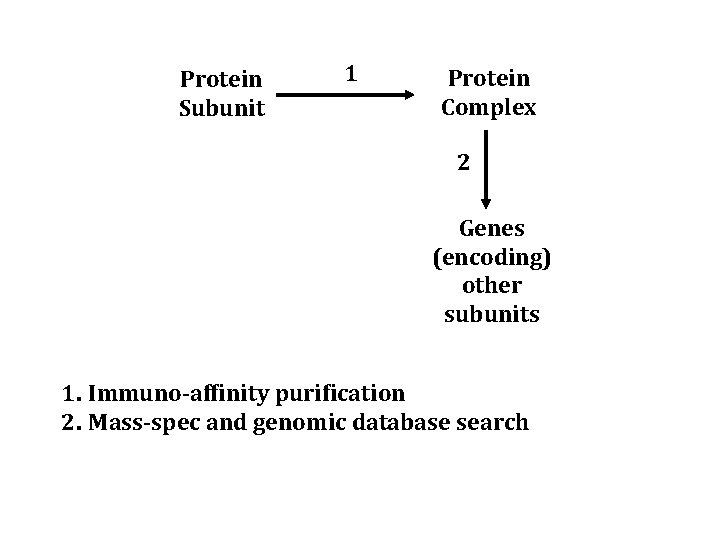
Protein Subunit 1 Protein Complex 2 Genes (encoding) other subunits 1. Immuno-affinity purification 2. Mass-spec and genomic database search
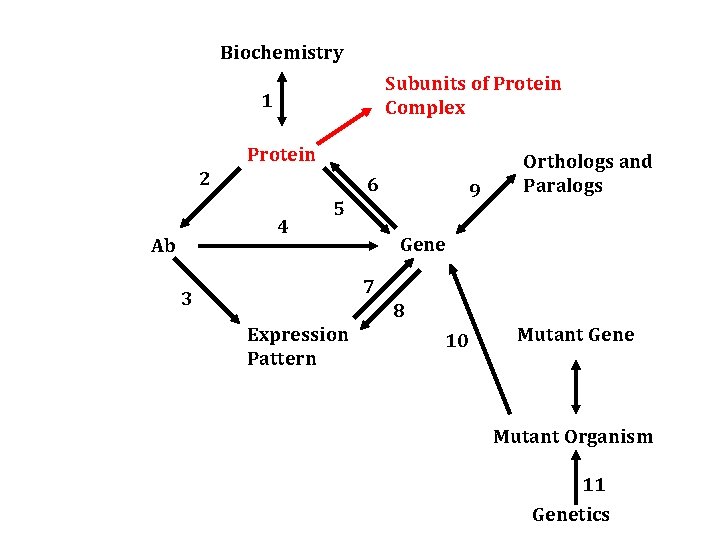
Biochemistry Subunits of Protein Complex 1 Protein 2 6 4 Ab 9 5 Orthologs and Paralogs Gene 7 3 8 Expression Pattern 10 Mutant Gene Mutant Organism 11 Genetics
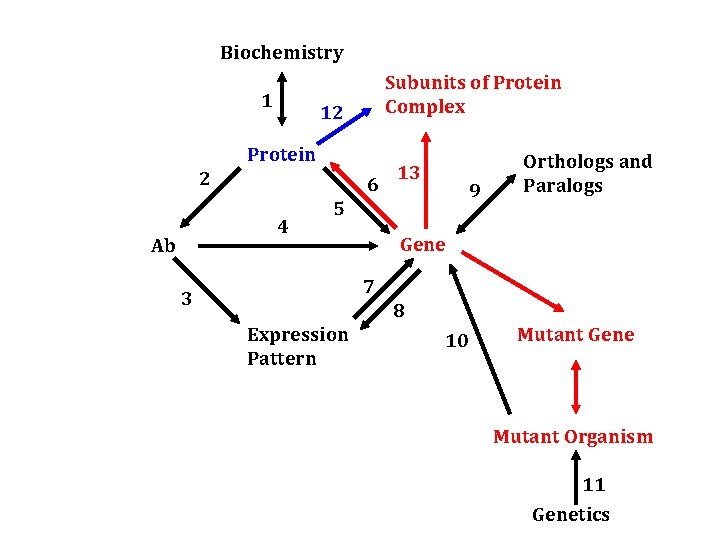
Biochemistry 1 Subunits of Protein Complex 12 Protein 2 6 4 Ab 13 9 5 Orthologs and Paralogs Gene 7 3 8 Expression Pattern 10 Mutant Gene Mutant Organism 11 Genetics

Yeast Genome Manipulation via Homologous Recombination • Gene disruption – Determine null phenotype of a gene • Gene replacement – Create mutant alleles of a gene [pt mut, deletion series, etc] • Epitope TAG • GFP fusions and protein localization
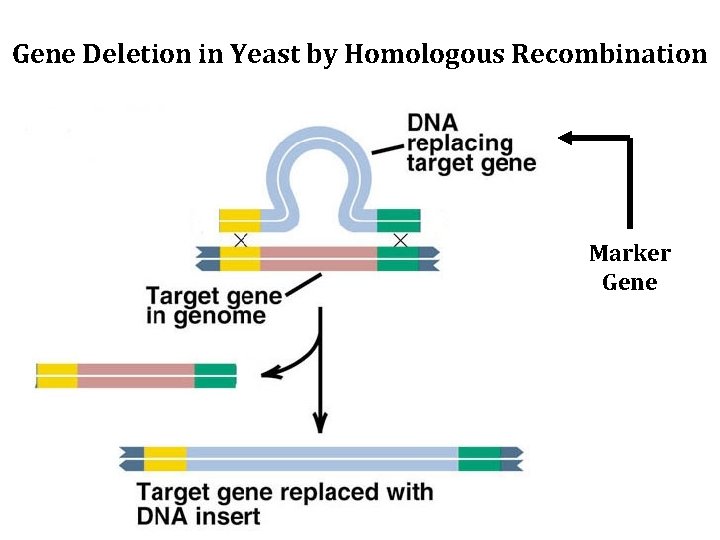
Gene Deletion in Yeast by Homologous Recombination Marker Gene

Gene Disruption in Yeast
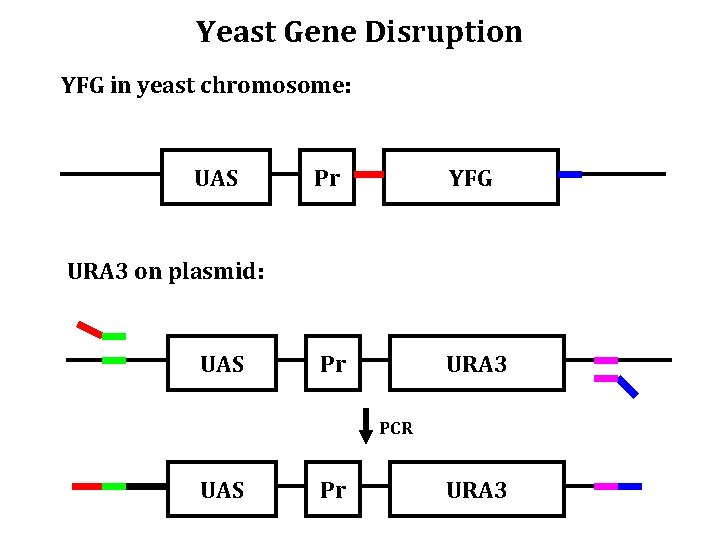
Yeast Gene Disruption YFG in yeast chromosome: UAS Pr YFG Pr URA 3 on plasmid: UAS PCR UAS Pr URA 3
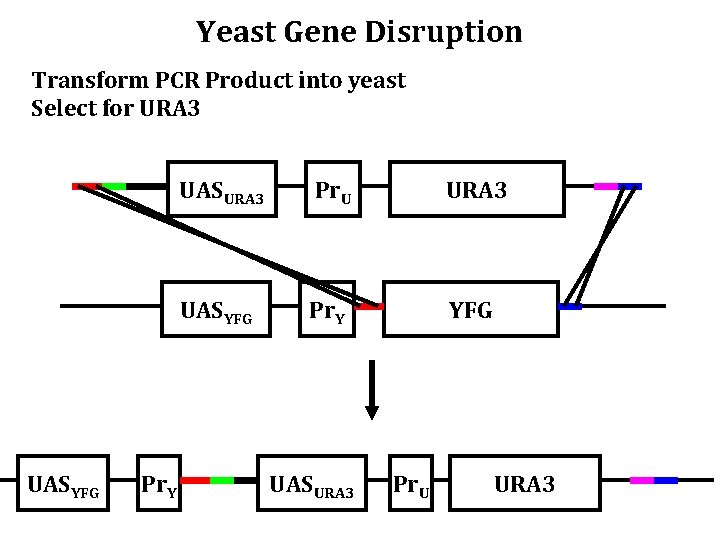
Yeast Gene Disruption Transform PCR Product into yeast Select for URA 3 UASYFG Pr. Y UASURA 3 Pr. U URA 3 UASYFG Pr. Y YFG UASURA 3 Pr. U URA 3
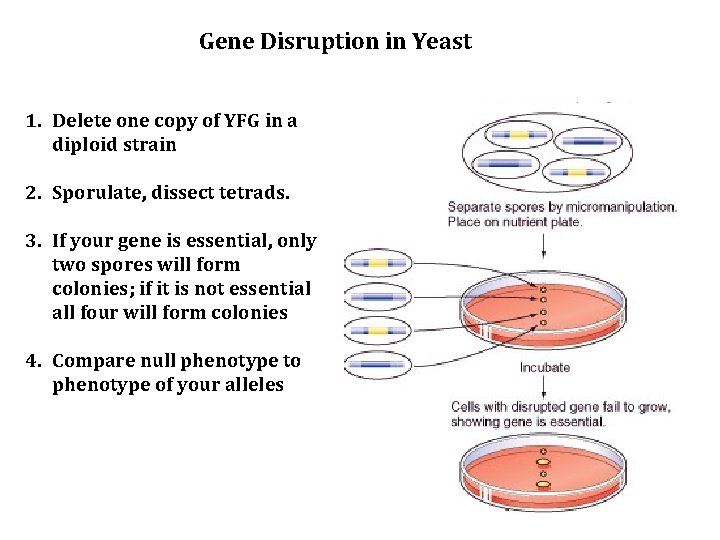
Gene Disruption in Yeast 1. Delete one copy of YFG in a diploid strain 2. Sporulate, dissect tetrads. 3. If your gene is essential, only two spores will form colonies; if it is not essential all four will form colonies 4. Compare null phenotype to phenotype of your alleles
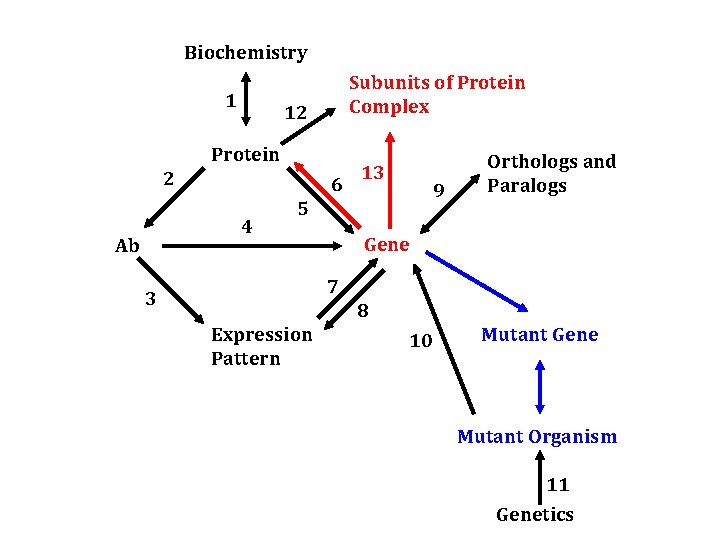
Biochemistry 1 Subunits of Protein Complex 12 Protein 2 6 4 Ab 13 9 5 Orthologs and Paralogs Gene 7 3 8 Expression Pattern 10 Mutant Gene Mutant Organism 11 Genetics
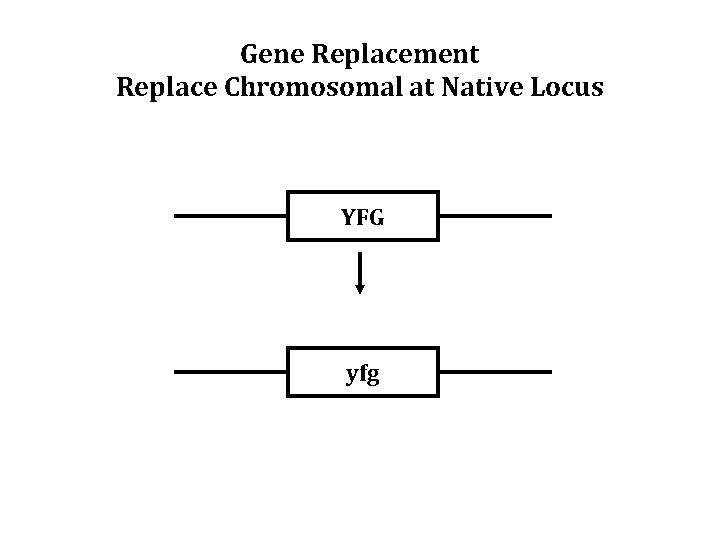
Gene Replacement Replace Chromosomal at Native Locus YFG yfg
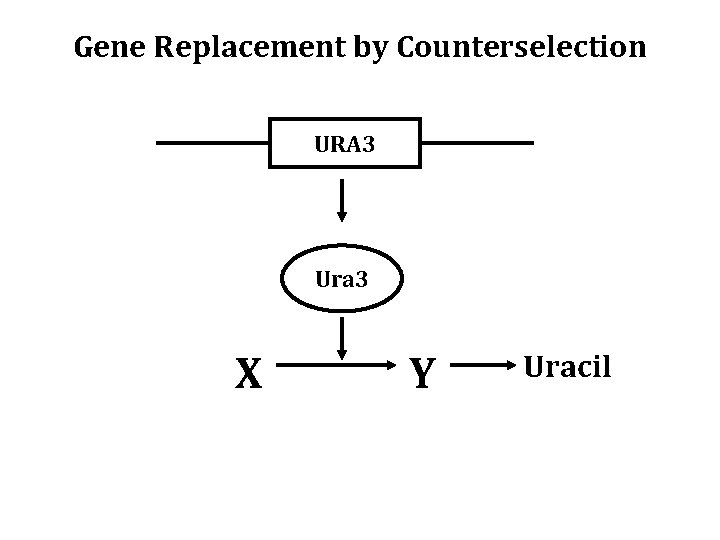
Gene Replacement by Counterselection URA 3 Ura 3 X Y Uracil
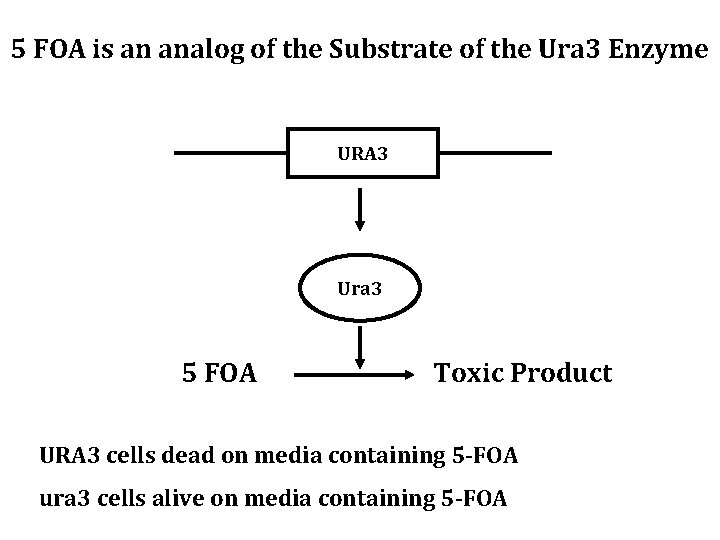
5 FOA is an analog of the Substrate of the Ura 3 Enzyme URA 3 Ura 3 5 FOA Toxic Product URA 3 cells dead on media containing 5 -FOA ura 3 cells alive on media containing 5 -FOA
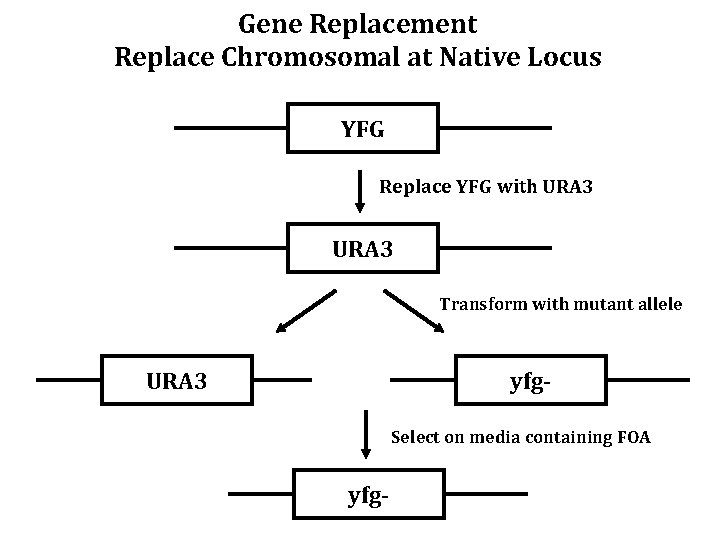
Gene Replacement Replace Chromosomal at Native Locus YFG Replace YFG with URA 3 Transform with mutant allele URA 3 yfg. Select on media containing FOA yfg-
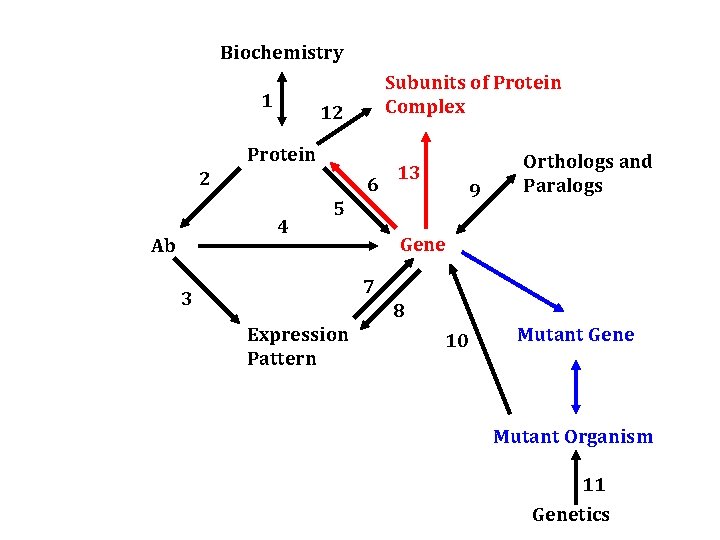
Biochemistry 1 Subunits of Protein Complex 12 Protein 2 6 4 Ab 13 9 5 Orthologs and Paralogs Gene 7 3 8 Expression Pattern 10 Mutant Gene Mutant Organism 11 Genetics
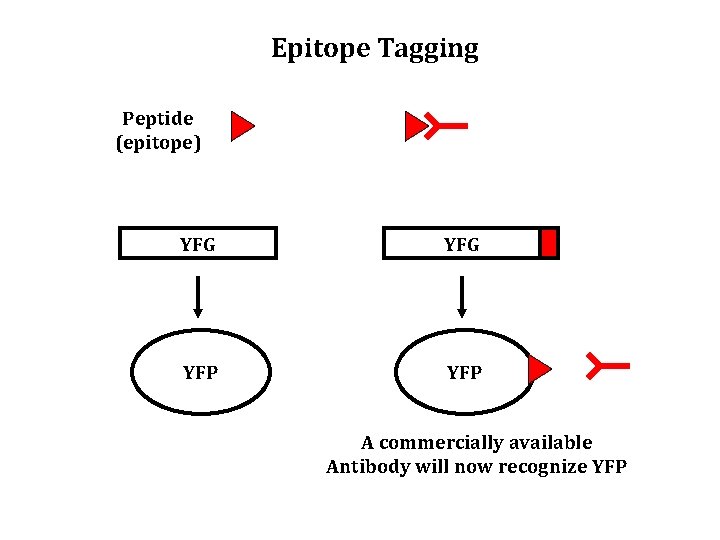
Epitope Tagging Peptide (epitope) YFG YFP A commercially available Antibody will now recognize YFP
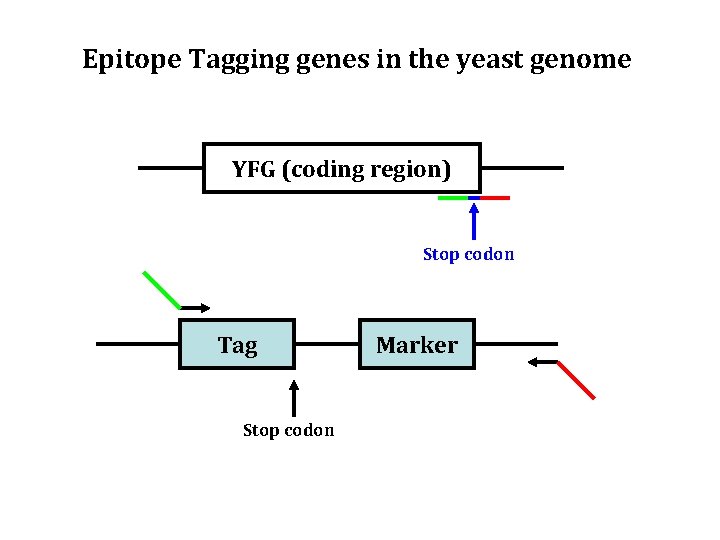
Epitope Tagging genes in the yeast genome YFG (coding region) Stop codon Tag Stop codon Marker
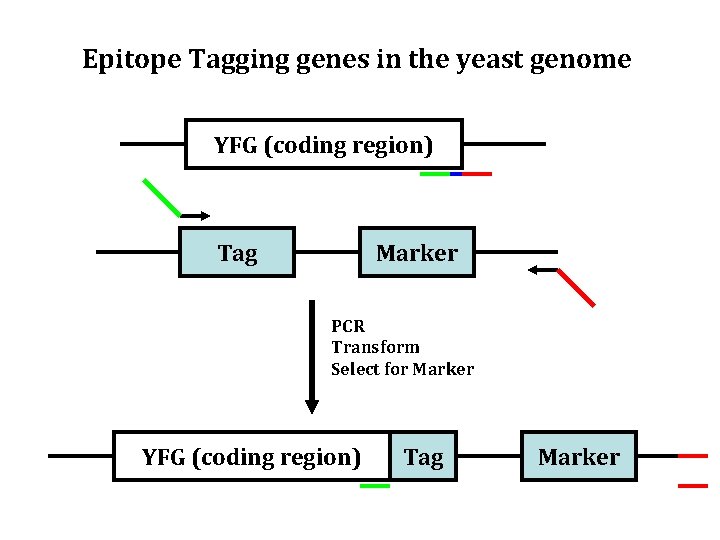
Epitope Tagging genes in the yeast genome YFG (coding region) Tag Marker PCR Transform Select for Marker YFG (coding region) Tag Marker

Is YFP part of a complex? If so, what other proteins are in the complex? 1. Identify YFG (genetic screen for instance) 2. Epitope tag 3. Immuno-affinity purification 4. Mass spec
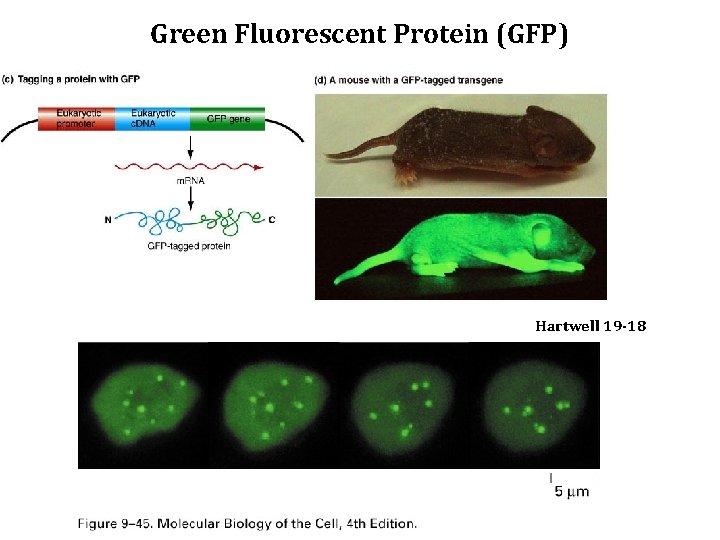
Green Fluorescent Protein (GFP) Hartwell 19 -18
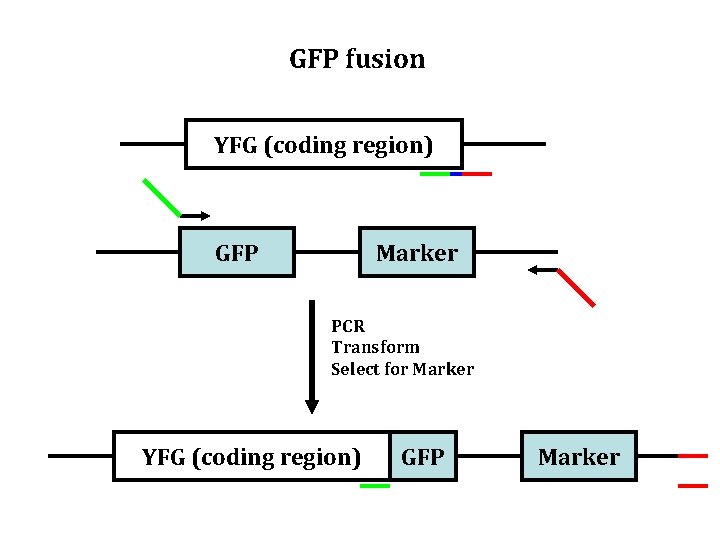
GFP fusion YFG (coding region) GFP Marker PCR Transform Select for Marker YFG (coding region) GFP Marker

Yeast Genome Manipulation via Homologous Recombination • Gene disruption – Determine null phenotype of a gene • Gene replacement – Create mutant alleles of a gene [pt mut, deletion series, etc] • Epitope TAG • GFP fusions and protein localization
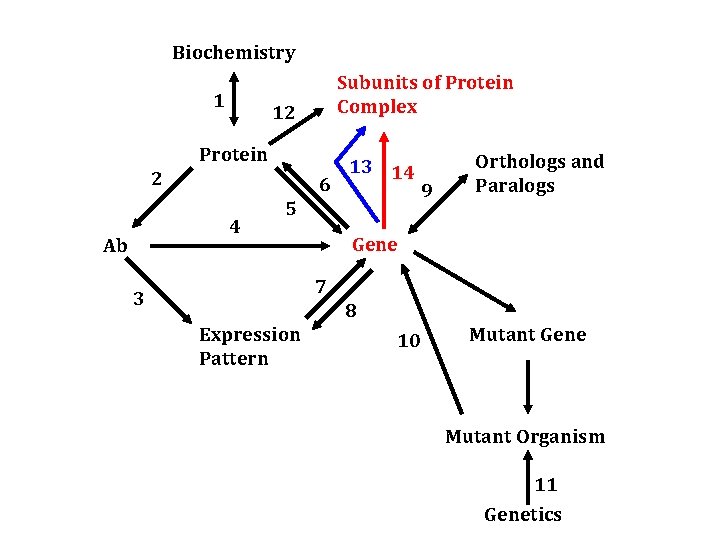
Biochemistry 1 Subunits of Protein Complex 12 Protein 2 6 4 Ab 13 14 5 9 Orthologs and Paralogs Gene 7 3 8 Expression Pattern 10 Mutant Gene Mutant Organism 11 Genetics
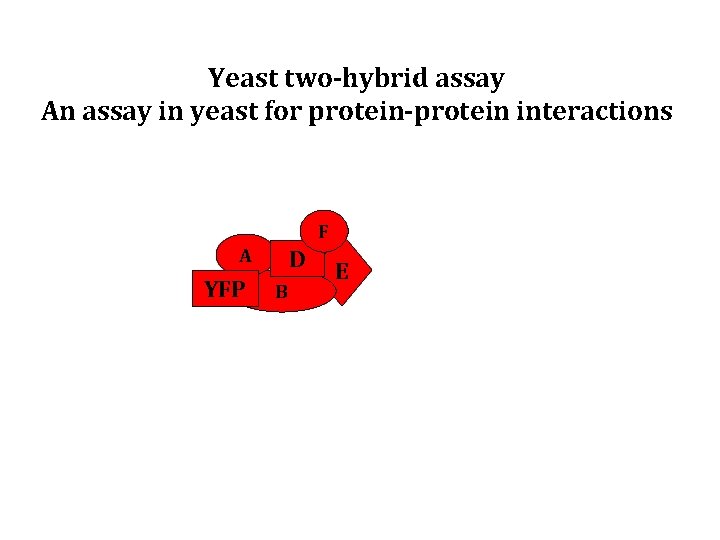
Yeast two-hybrid assay An assay in yeast for protein-protein interactions F A YFP D B E
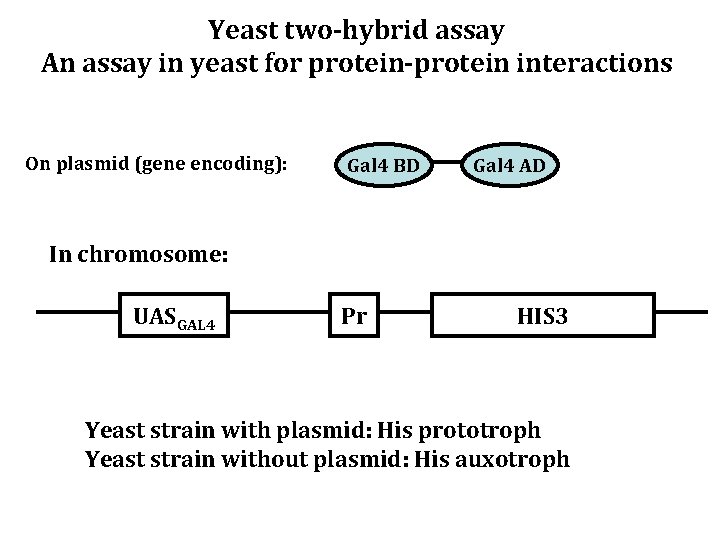
Yeast two-hybrid assay An assay in yeast for protein-protein interactions On plasmid (gene encoding): Gal 4 BD Gal 4 AD In chromosome: UASGAL 4 Pr HIS 3 Yeast strain with plasmid: His prototroph Yeast strain without plasmid: His auxotroph
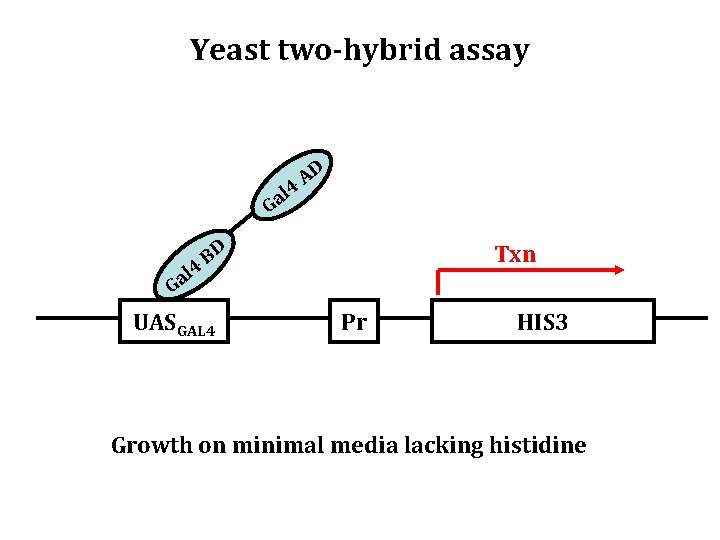
Yeast two-hybrid assay l 4 a G l 4 Ga AD BD UASGAL 4 Txn Pr HIS 3 Growth on minimal media lacking histidine
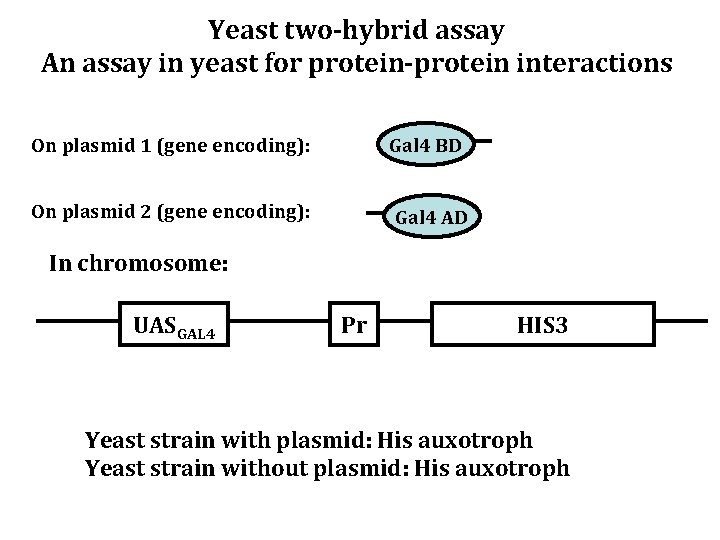
Yeast two-hybrid assay An assay in yeast for protein-protein interactions On plasmid 1 (gene encoding): Gal 4 BD On plasmid 2 (gene encoding): Gal 4 AD In chromosome: UASGAL 4 Pr HIS 3 Yeast strain with plasmid: His auxotroph Yeast strain without plasmid: His auxotroph
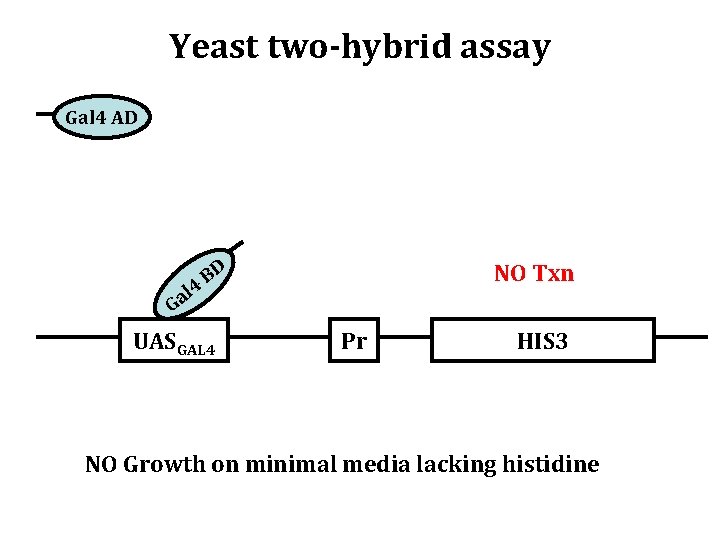
Yeast two-hybrid assay Gal 4 AD l 4 Ga BD UASGAL 4 NO Txn Pr HIS 3 NO Growth on minimal media lacking histidine
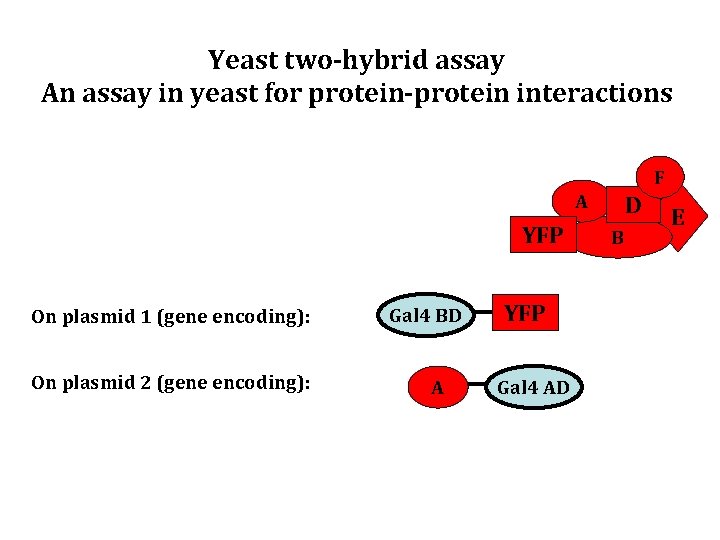
Yeast two-hybrid assay An assay in yeast for protein-protein interactions F A YFP On plasmid 1 (gene encoding): On plasmid 2 (gene encoding): Gal 4 BD A YFP Gal 4 AD D B E
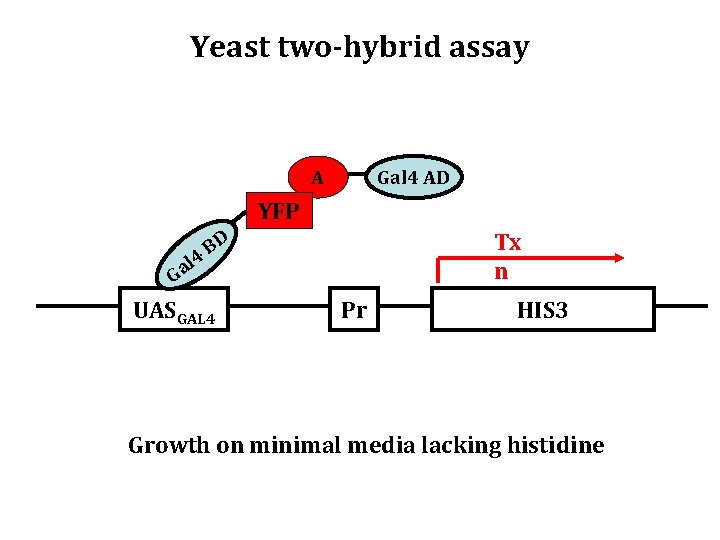
Yeast two-hybrid assay A Gal 4 AD YFP 4 l Ga Tx n BD UASGAL 4 Pr HIS 3 Growth on minimal media lacking histidine
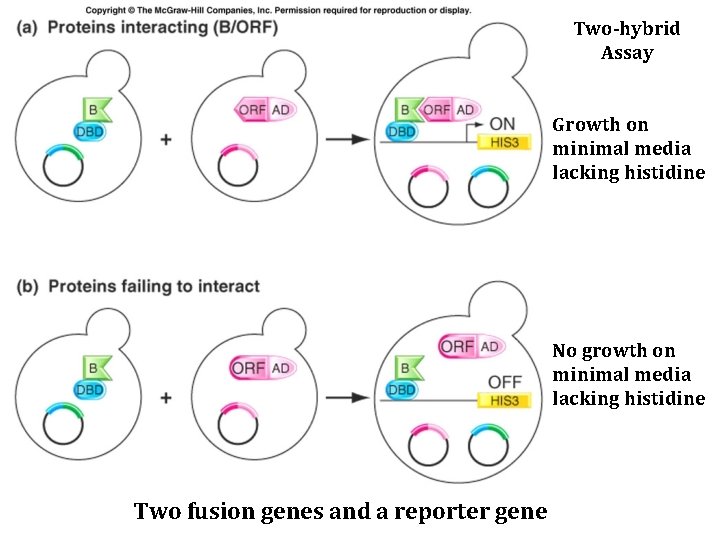
Two-hybrid Assay Growth on minimal media lacking histidine No growth on minimal media lacking histidine Two fusion genes and a reporter gene
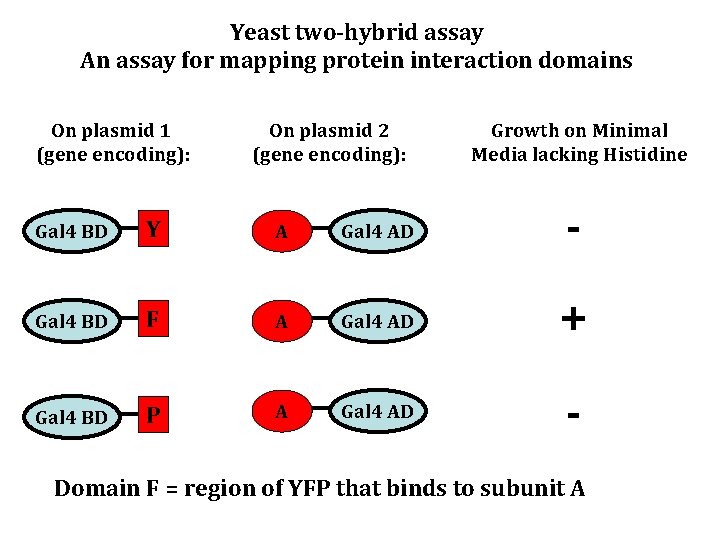
Yeast two-hybrid assay An assay for mapping protein interaction domains On plasmid 1 (gene encoding): On plasmid 2 (gene encoding): Growth on Minimal Media lacking Histidine Gal 4 BD Y A Gal 4 AD - Gal 4 BD F A Gal 4 AD + Gal 4 BD P A Gal 4 AD - Domain F = region of YFP that binds to subunit A
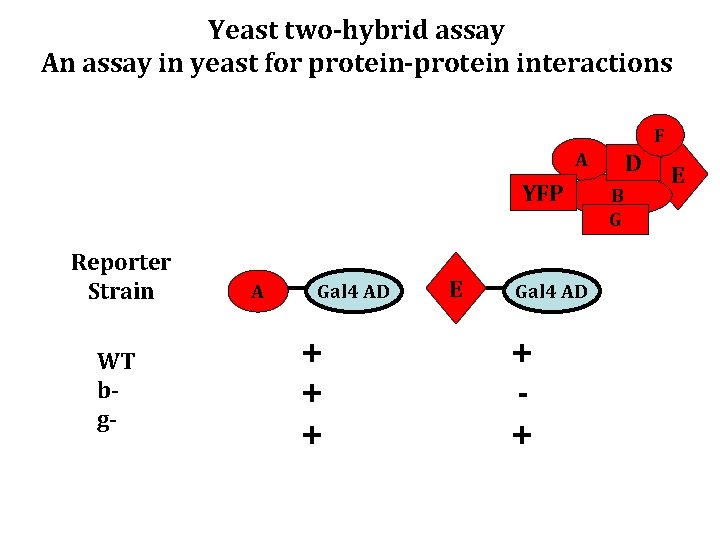
Yeast two-hybrid assay An assay in yeast for protein-protein interactions F A YFP Reporter Strain WT bg- A Gal 4 AD + + + E Gal 4 AD + + D B G E
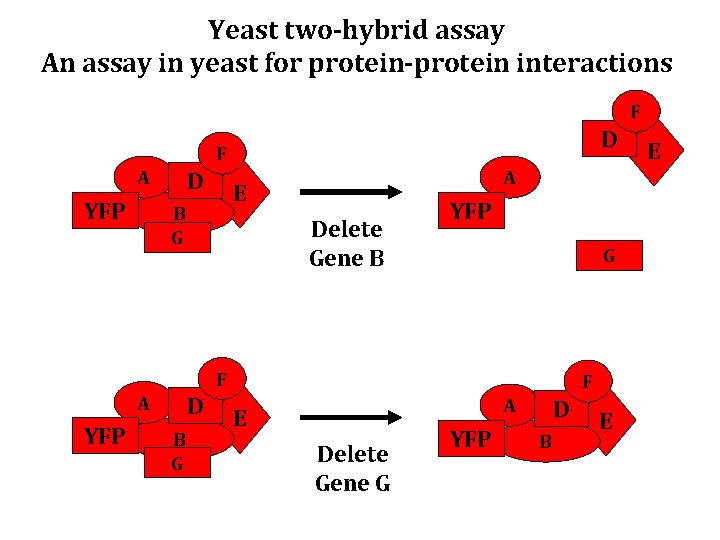
Yeast two-hybrid assay An assay in yeast for protein-protein interactions F D F A YFP D A E B G Delete Gene B YFP G F A YFP D B G F A E Delete Gene G YFP D B E E
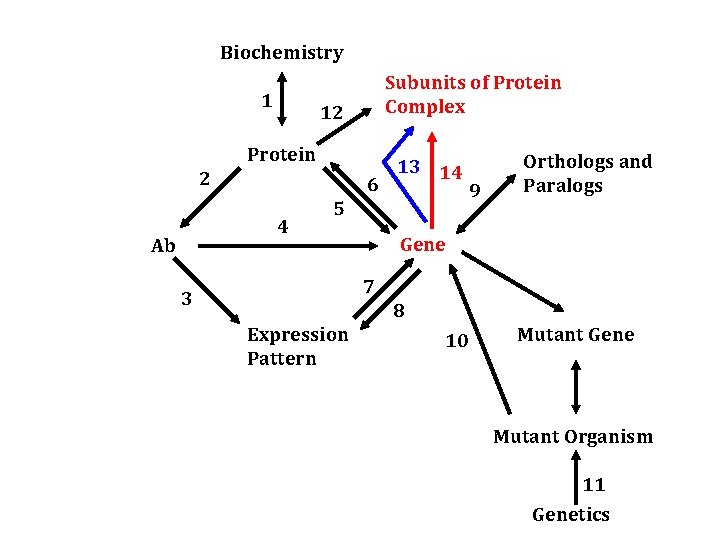
Biochemistry 1 Subunits of Protein Complex 12 Protein 2 6 4 Ab 13 14 5 9 Orthologs and Paralogs Gene 7 3 8 Expression Pattern 10 Mutant Gene Mutant Organism 11 Genetics
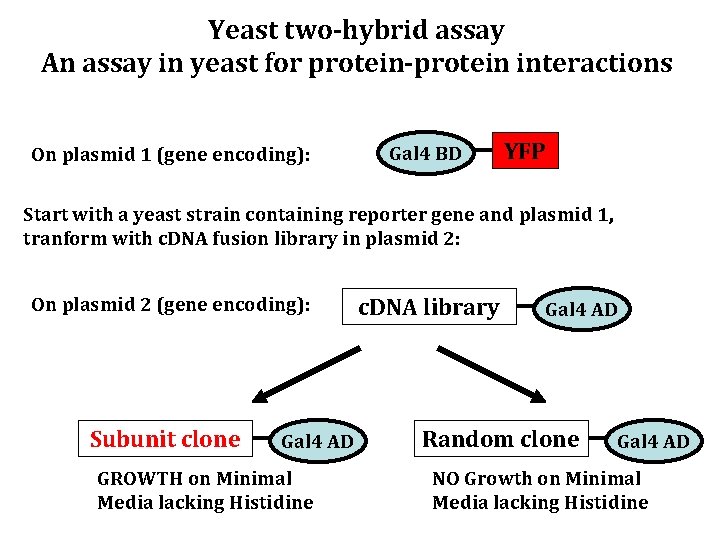
Yeast two-hybrid assay An assay in yeast for protein-protein interactions On plasmid 1 (gene encoding): Gal 4 BD YFP Start with a yeast strain containing reporter gene and plasmid 1, tranform with c. DNA fusion library in plasmid 2: On plasmid 2 (gene encoding): Subunit clone Gal 4 AD GROWTH on Minimal Media lacking Histidine c. DNA library Gal 4 AD Random clone Gal 4 AD NO Growth on Minimal Media lacking Histidine
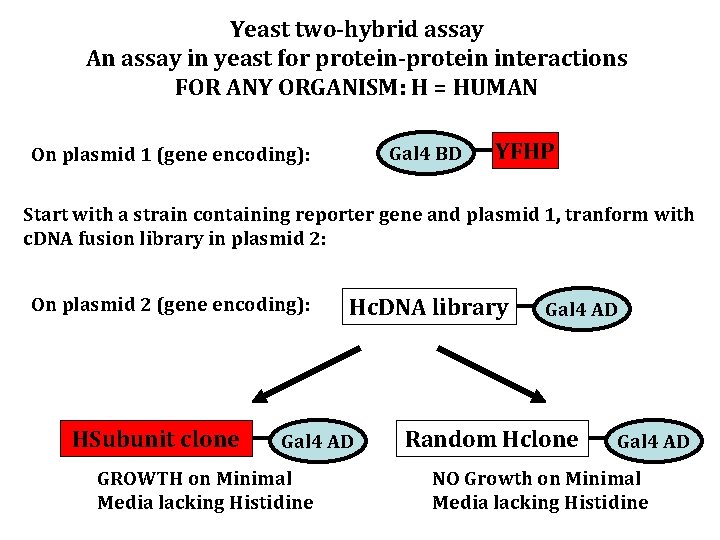
Yeast two-hybrid assay An assay in yeast for protein-protein interactions FOR ANY ORGANISM: H = HUMAN Gal 4 BD On plasmid 1 (gene encoding): YFHP Start with a strain containing reporter gene and plasmid 1, tranform with c. DNA fusion library in plasmid 2: On plasmid 2 (gene encoding): HSubunit clone Hc. DNA library Gal 4 AD GROWTH on Minimal Media lacking Histidine Gal 4 AD Random Hclone Gal 4 AD NO Growth on Minimal Media lacking Histidine
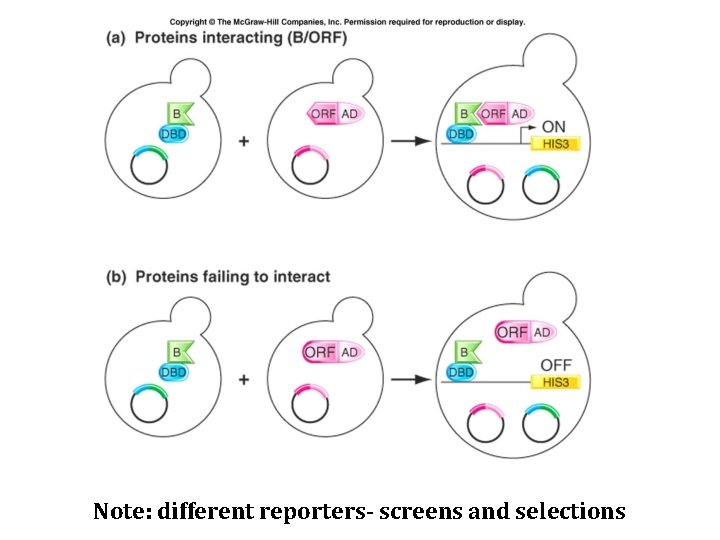
Note: different reporters- screens and selections

Some uses of two-hybrid assay • Screen genomic library for additional subunits of a protein complex NOTE: Genes/Libraries can be from any organism • Infer some aspects of the architecture of a complex (combine with mutant) • Mapping interaction regions/domains • Test candidate interactions (genes identified in a screen, for instance)
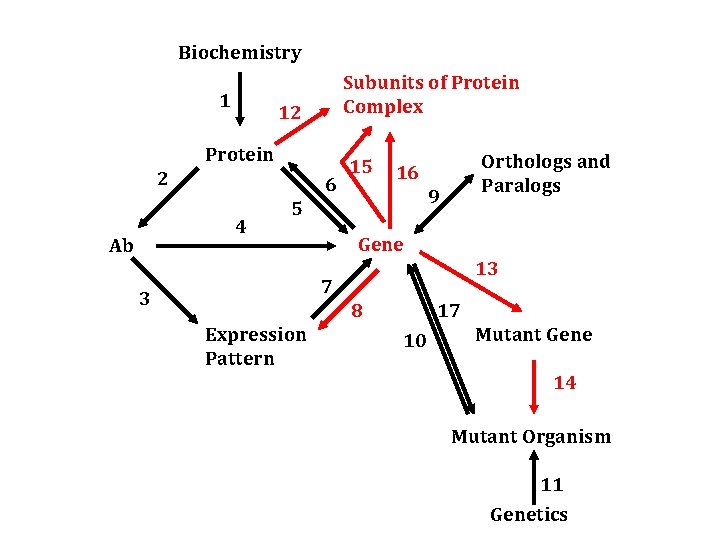
Biochemistry 1 Subunits of Protein Complex 12 Protein 2 6 4 Ab 15 Orthologs and Paralogs 16 9 5 Gene 13 7 3 8 Expression Pattern 17 10 Mutant Gene 14 Mutant Organism 11 Genetics

Molecular Genetics Summary 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. 16. 17. Column Chromatograpy (ion exch, gel filtr) + in vitro assay A. Make Polyclonal Ab; B. Make Monoclonal Ab Western blot, in situ immuno-fluorescence (subcellular, tissue) Screen expression library (with an Ab) Screen library with degenerate probe Protein expression (E. coli) A. Differential hybridization A. Northern blot (RNA), in situ hybridization (RNA or Protein), GFP tag (Protein pattern and sub-cellular localization) A. low stringency hybridization; B. computer search/clone by phone; C. computer search PCR Clone by complementation (yeast, E. coli) A. Genetic screen; B. genetic selection Immuno-affinity purification + Mass spec + Computer search In vitro mutagenesis (site directed, deletion, etc) Gene replacement (Yeast, Homologous Recombination) Epitope tag + immuno-affinity chromatography Yeast two-hybrid analysis and screens RNAi
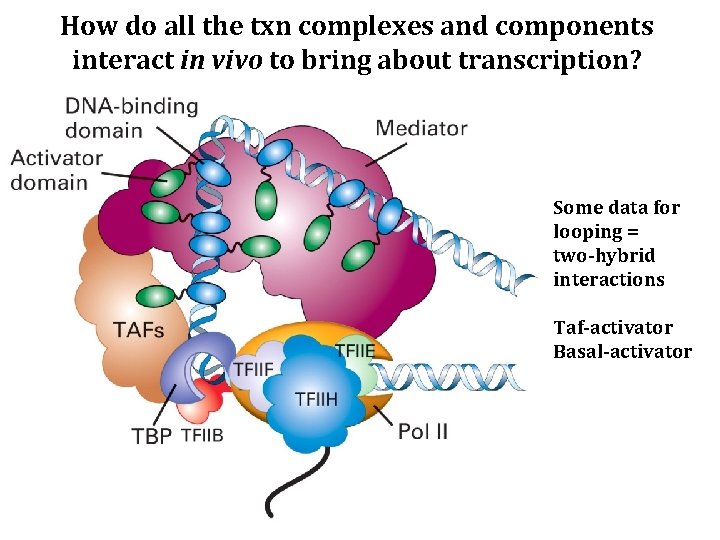
How do all the txn complexes and components interact in vivo to bring about transcription? Some data for looping = two-hybrid interactions Taf-activator Basal-activator
- Slides: 54