MCB 130 L Lecture 1 1 How to

MCB 130 L Lecture 1 1. How to get the most from your time in lab 2. Recombinant DNA 3. Tips on giving a Powerpoint talk

1. How to get the most from your time in lab 1. Be well prepared: know what you are doing and why 2. Be organized 3. Be systematic in your work 4. Take careful and thoughtful notes 5. Clean up after yourself when done!!

2. Recombinant DNA technology Recombinant DNA: Creation of a novel combination (e. g. , human and bacteria DNA) Applications: 1. Cloning = obtaining multiple copies 2. Sequencing 3. Modification: Mutagenesis …………

Importance • Biotechnology (e. g. , insulin, growth hormone) • Basic research (gene structure, function, conservation) • Gene therapy

Which genomes have been sequenced? • • • Viruses Phage Organelle genomes Plants Model Organisms: yeasts, flies, worms Vertebrates (including humans)
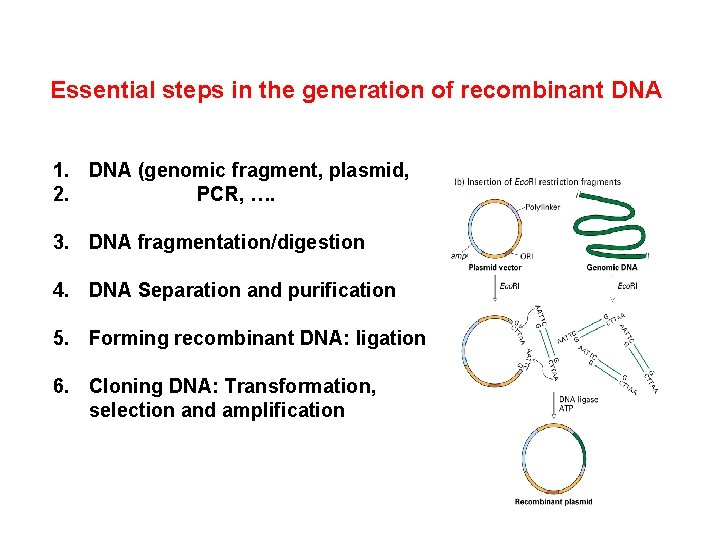
Essential steps in the generation of recombinant DNA 1. DNA (genomic fragment, plasmid, 2. PCR, …. 3. DNA fragmentation/digestion 4. DNA Separation and purification 5. Forming recombinant DNA: ligation 6. Cloning DNA: Transformation, selection and amplification
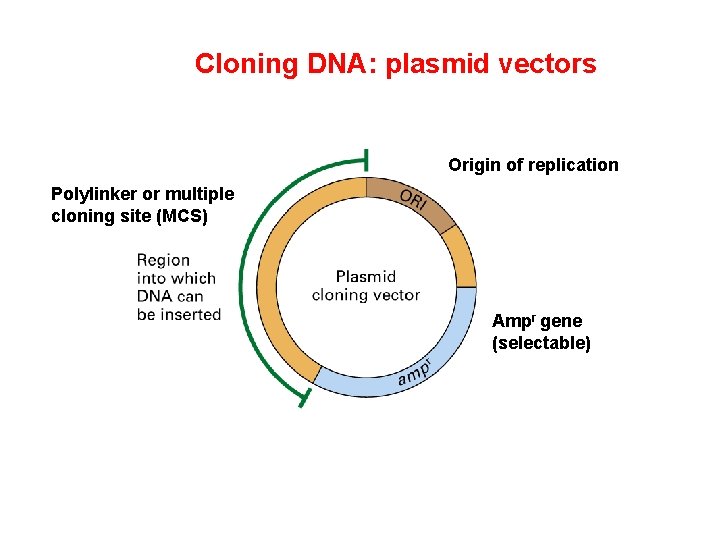
Cloning DNA: plasmid vectors Origin of replication Polylinker or multiple cloning site (MCS) Ampr gene (selectable)
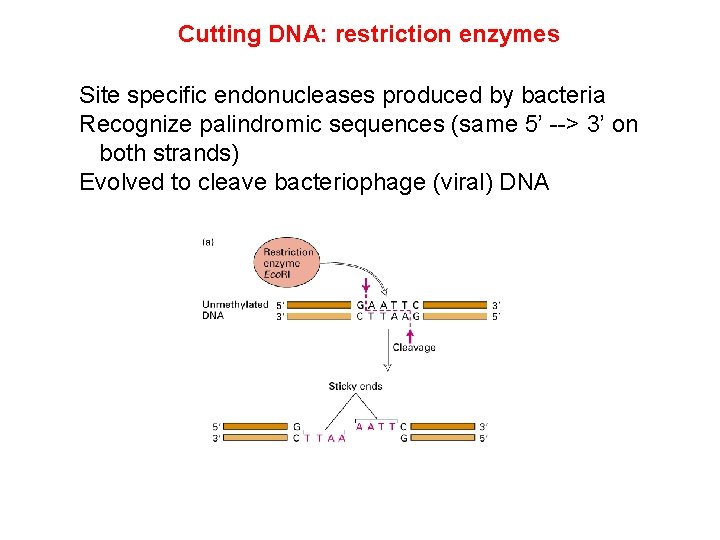
Cutting DNA: restriction enzymes Site specific endonucleases produced by bacteria Recognize palindromic sequences (same 5’ --> 3’ on both strands) Evolved to cleave bacteriophage (viral) DNA

Cutting DNA: restriction enzymes blunt ends Sticky ends: 5’ overhang 3’ overhang

Cutting DNA: restriction enzymes 1. Numerous restriction enzymes 2. Most cleave at a unique sequence 3. Named for bacterial species
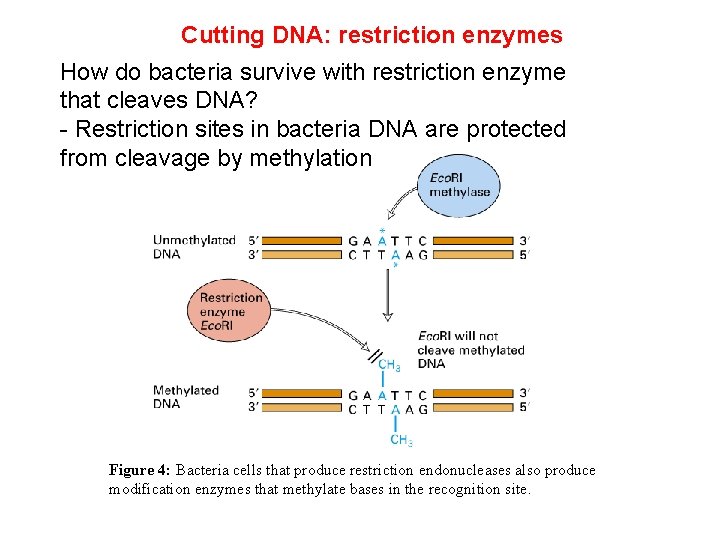
Cutting DNA: restriction enzymes How do bacteria survive with restriction enzyme that cleaves DNA? - Restriction sites in bacteria DNA are protected from cleavage by methylation Figure 4: Bacteria cells that produce restriction endonucleases also produce modification enzymes that methylate bases in the recognition site.
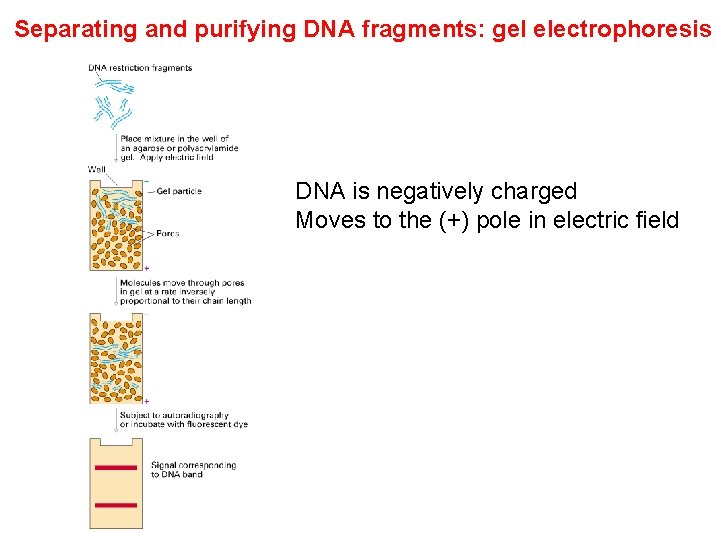
Separating and purifying DNA fragments: gel electrophoresis DNA is negatively charged Moves to the (+) pole in electric field
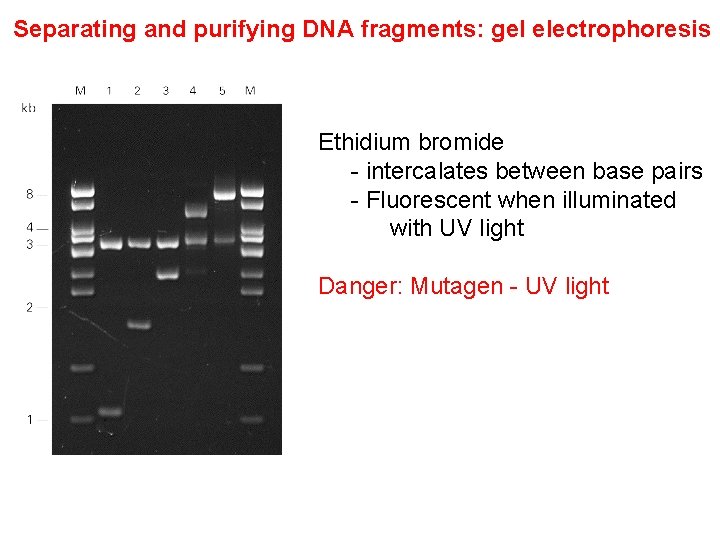
Separating and purifying DNA fragments: gel electrophoresis Ethidium bromide - intercalates between base pairs - Fluorescent when illuminated with UV light Danger: Mutagen - UV light
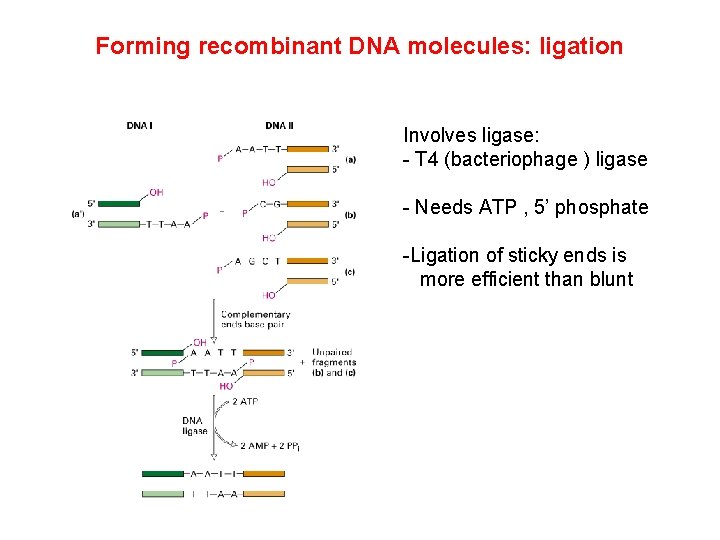
Forming recombinant DNA molecules: ligation Involves ligase: - T 4 (bacteriophage ) ligase - Needs ATP , 5’ phosphate -Ligation of sticky ends is more efficient than blunt
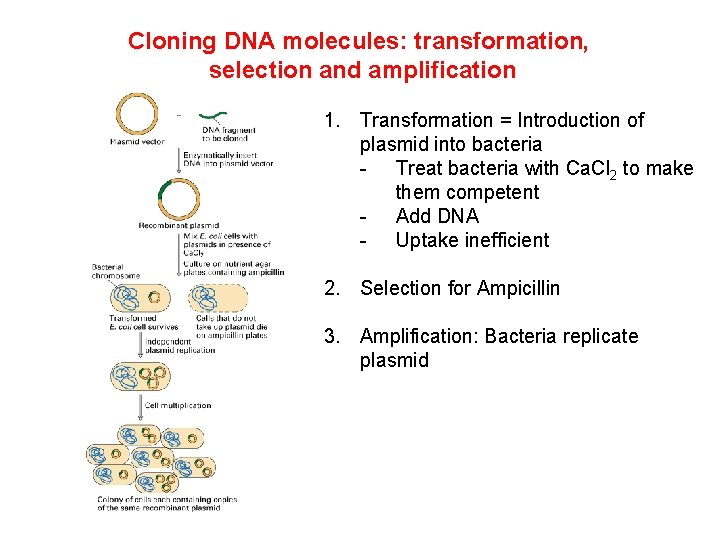
Cloning DNA molecules: transformation, selection and amplification 1. Transformation = Introduction of plasmid into bacteria - Treat bacteria with Ca. Cl 2 to make them competent - Add DNA - Uptake inefficient 2. Selection for Ampicillin 3. Amplification: Bacteria replicate plasmid

Amplification of specific DNA sequences: Polymerase Chain Reaction (PCR) Applications: 1. 2. 3. 4. general amplification diagnostics isolating DNA from ancient organisms forensics…. Invented by Kerry Mullis, UCB Ph. D, while at Cetus 1993 Nobel Prize in Chemistry

Amplification of specific DNA sequences: Polymerase Chain Reaction (PCR) 1. Logarithmic amplification: # of copies = 2 n, n= # of cycles 2. Sensitive: a single molecule can be amplified 3. Contamination a problem!

Amplification of specific DNA sequences: Polymerase Chain Reaction (PCR) Technique uses: 1. DNA polymerase from thermophilic bacteria ex: Taq from thermus aquaticus (no proofreading, error rate 1/105) 2. d. NTPs (d. ATP, d. CTP, d. TTP, d. GTP) 3. Template = DNA to be amplified 4. primers: 18 -20 nucleotides complementary to template 5. Temperature cycling: 20 -30 cycles 95ºC: denaturation 55ºC to 60ºC annealing 72ºC Extension
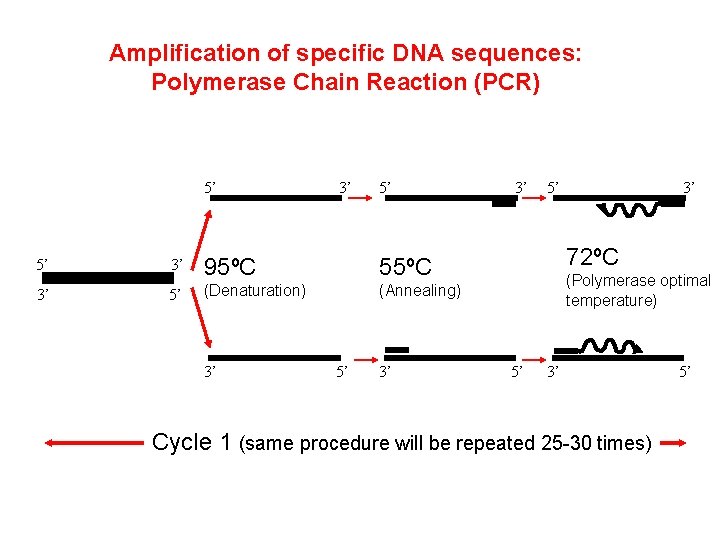
Amplification of specific DNA sequences: Polymerase Chain Reaction (PCR) 5’ 3’ 5’ 5’ 3’ 95ºC 55ºC 3’ 5’ (Denaturation) (Annealing) 3’ 5’ 3’ 72ºC (Polymerase optimal temperature) 5’ 3’ Cycle 1 (same procedure will be repeated 25 -30 times) 5’
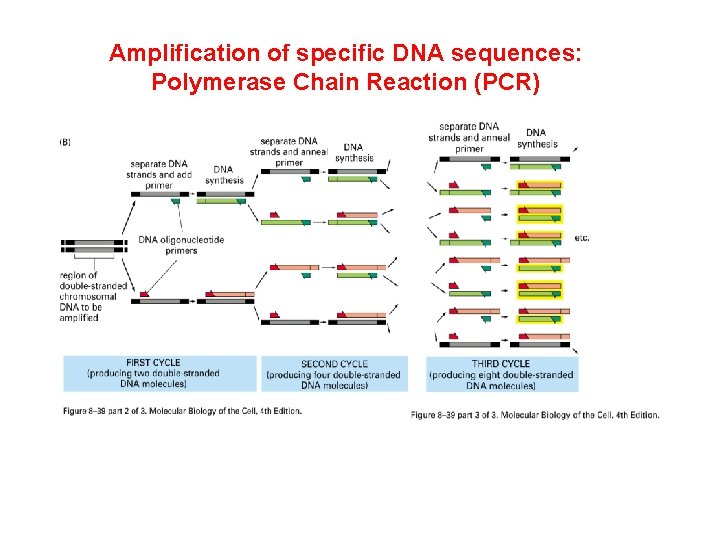
Amplification of specific DNA sequences: Polymerase Chain Reaction (PCR)
- Slides: 20