MBSW 2009 May 18 20 Muncie IN Response

MBSW’ 2009, May 18 -20, Muncie, IN Response Optimization in Oncology In Vivo Studies: a Multiobjective Modeling Approach Maksim Pashkevich, Ph. D (Early Phase Oncology Statistics) Joint work with Philip Iversen, Ph. D (Pre-Clinical Oncology Statistics) Harold Brooks, Ph. D (Growth and Translational Genetics) Eli Lilly and Company

Outline • Problem overview – In vivo studies in oncology drug development – Efficacy and toxicity measures for in vivo studies – Optimal regimen as balance between efficacy / toxicity • Models for efficacy and toxicity – Modified Simeoni model of tumor growth inhibition – Animal body weight loss model to describe toxicity – Statistical estimation of model parameters in Matlab • Optimal regimen simulation – Multiobjective representation of simulation results – Pareto-optimal set of optimal dosing regimens 2

Motivation In vivo studies in oncology • • • Typical way to assess cancer compound activity Cancer tumors are implanted in mice or rats Tumor size and animal weight are measured over time Efficacy and toxicity measures • • Tumor growth delay is a standard efficacy measure Body weight loss is a typical surrogate for toxicity Optimal dosing regimen is unknown • • • Goal is to achieve balance between efficacy and toxicity Number of possible dosing regimens is very significant Modeling should help to select promising regimens 3
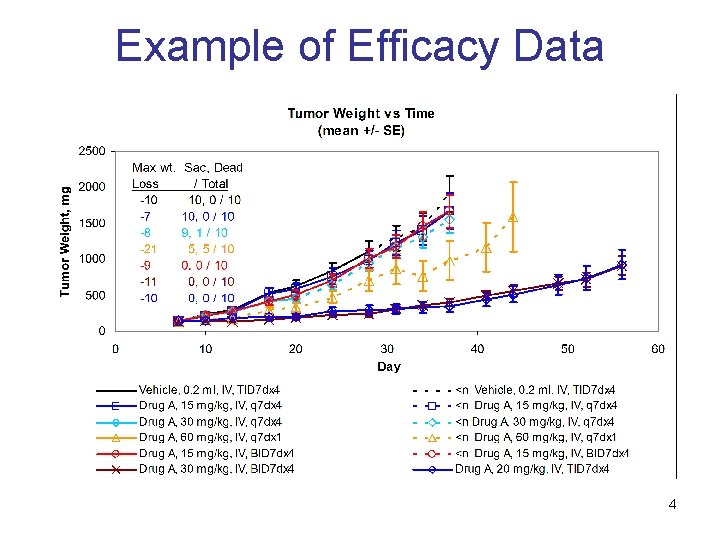
Example of Efficacy Data 4
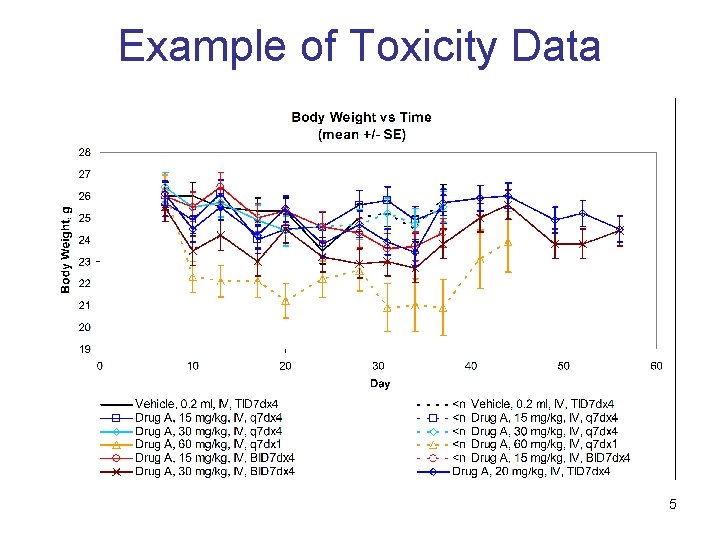
Example of Toxicity Data 5
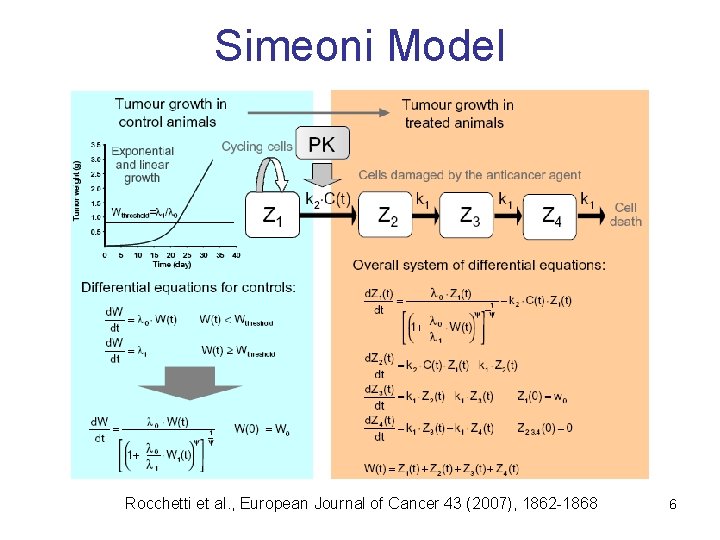
Simeoni Model Rocchetti et al. , European Journal of Cancer 43 (2007), 1862 -1868 6
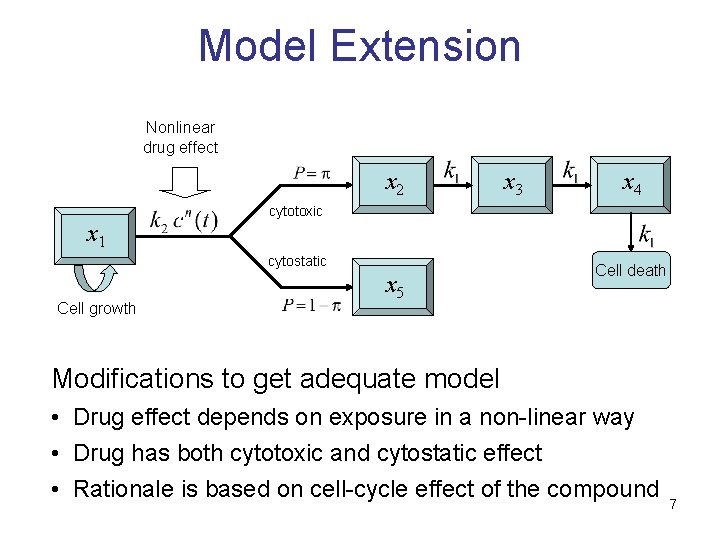
Model Extension Nonlinear drug effect x 2 x 1 x 4 cytotoxic cytostatic Cell growth x 3 x 5 Cell death Modifications to get adequate model • Drug effect depends on exposure in a non-linear way • Drug has both cytotoxic and cytostatic effect • Rationale is based on cell-cycle effect of the compound 7

Developed Efficacy Model Dynamic model: system of ordinary differential equations Initial conditions: with random effect for initial tumor weight 8
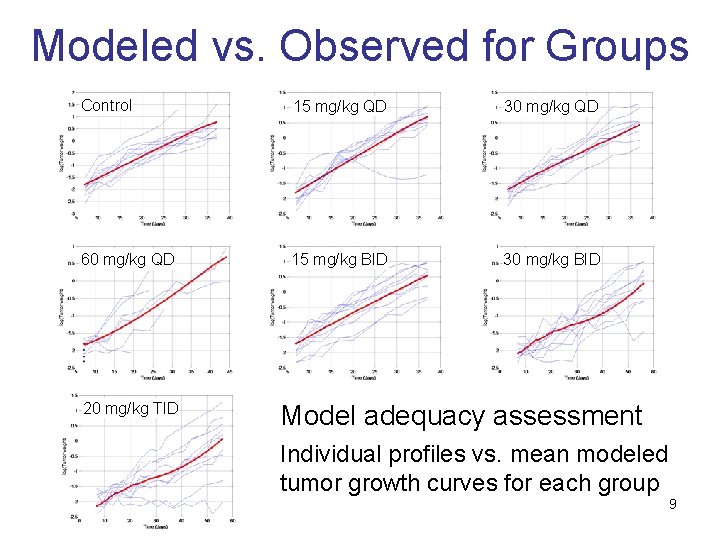
Modeled vs. Observed for Groups Control 15 mg/kg QD 30 mg/kg QD 60 mg/kg QD 15 mg/kg BID 30 mg/kg BID 20 mg/kg TID Model adequacy assessment Individual profiles vs. mean modeled tumor growth curves for each group 9
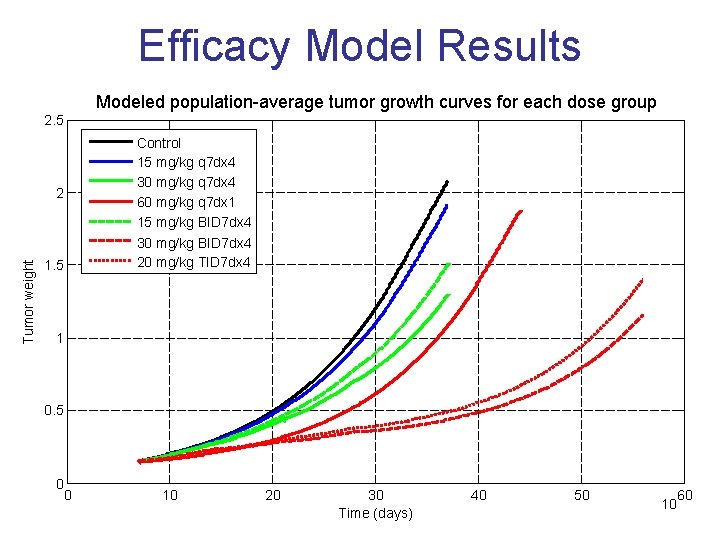
Efficacy Model Results Modeled population-average tumor growth curves for each dose group 2. 5 Control 15 mg/kg q 7 dx 4 30 mg/kg q 7 dx 4 60 mg/kg q 7 dx 1 15 mg/kg BID 7 dx 4 30 mg/kg BID 7 dx 4 20 mg/kg TID 7 dx 4 Tumor weight 2 1. 5 1 0. 5 0 0 10 20 30 Time (days) 40 50 10 60
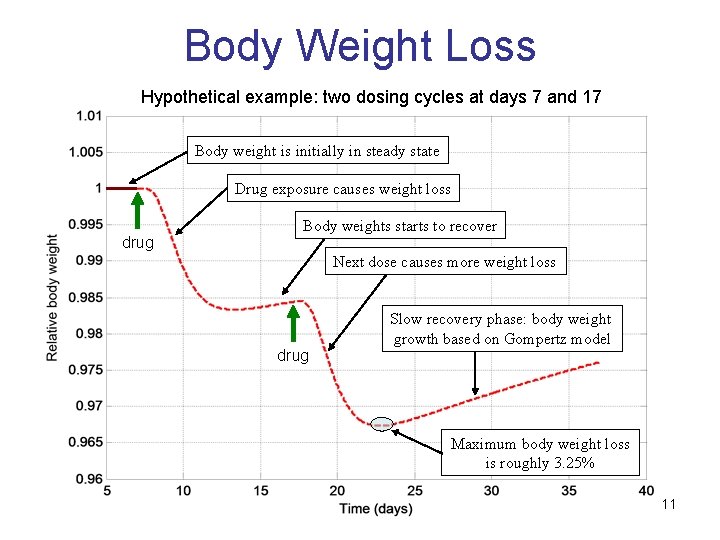
Body Weight Loss Hypothetical example: two dosing cycles at days 7 and 17 Body weight is initially in steady state Drug exposure causes weight loss drug Body weights starts to recover Next dose causes more weight loss drug Slow recovery phase: body weight growth based on Gompertz model Maximum body weight loss is roughly 3. 25% 11

Developed Toxicity Model Dynamic model: system of ordinary differential equations Initial conditions: with random effect for initial body weight 12
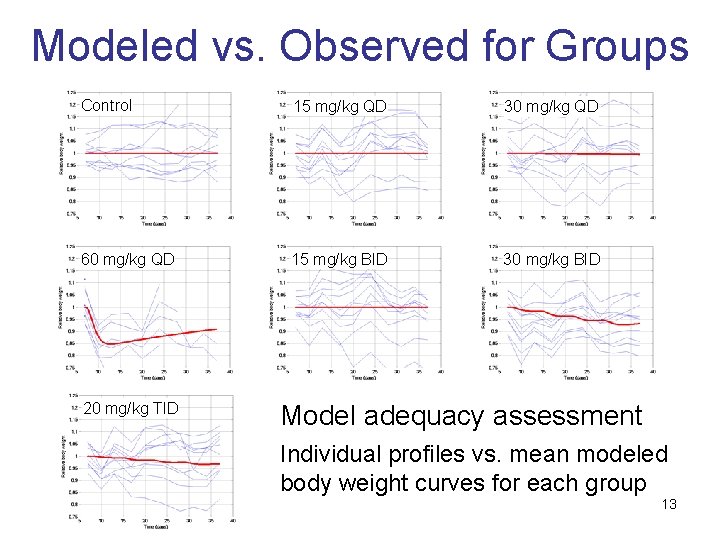
Modeled vs. Observed for Groups Control 15 mg/kg QD 30 mg/kg QD 60 mg/kg QD 15 mg/kg BID 30 mg/kg BID 20 mg/kg TID Model adequacy assessment Individual profiles vs. mean modeled body weight curves for each group 13
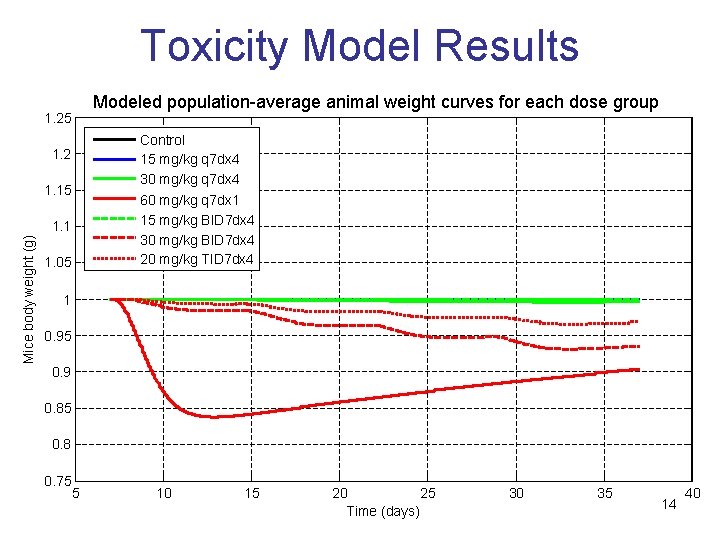
Toxicity Model Results Modeled population-average animal weight curves for each dose group 1. 25 Control 15 mg/kg q 7 dx 4 30 mg/kg q 7 dx 4 60 mg/kg q 7 dx 1 15 mg/kg BID 7 dx 4 30 mg/kg BID 7 dx 4 20 mg/kg TID 7 dx 4 1. 2 1. 15 Mice body weight (g) 1. 1 1. 05 1 0. 95 0. 9 0. 85 0. 8 0. 75 5 10 15 20 25 Time (days) 30 35 14 40

ML Parameter Estimation Computationally hard problem • • Numerical solution of system of differential equations Numerical integration due to random effects Numerical optimization of resulting likelihood function Three “heavy ”numerical problems nested in one another Implementation in Matlab • Relying on standard functions is unacceptably slow • Special problem-specific method was developed for ODE system solution and random effects integration • Numerical optimization was done by Matlab function 15

Regimens Simulation settings • Dosing was performed until day 28 as in original study • Doses from 1 to 30 mg/kg (QD, BID, TID) were used • Dosing interval was varied between 1 and 14 days Regimen evaluation • • • Efficacy and toxicity were computed for each regimen Efficacy was defined as overall tumor burden reduction Toxicity was defined as maximum relative weight loss Efficacy was plotted vs. toxicity for each simulation run Pareto-optimal solutions were identified for QD, BID, TID 16
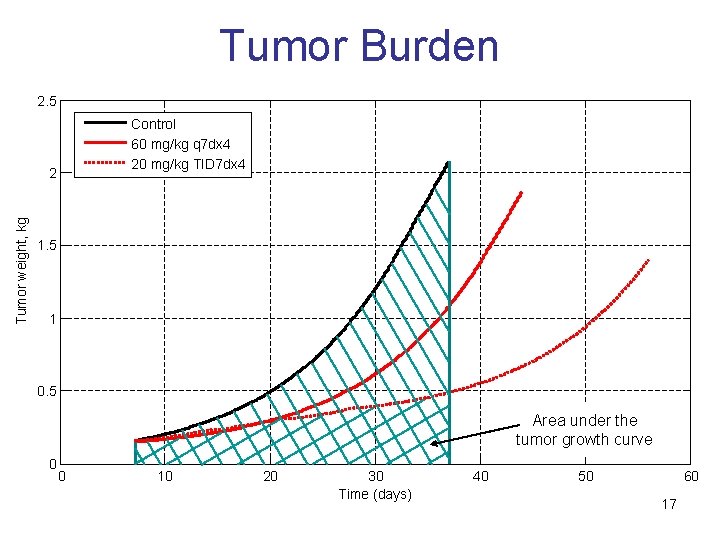
Tumor Burden 2. 5 Control 60 mg/kg q 7 dx 4 20 mg/kg TID 7 dx 4 Tumor weight, kg 2 1. 5 1 0. 5 Area under the tumor growth curve 0 0 10 20 30 Time (days) 40 50 60 17
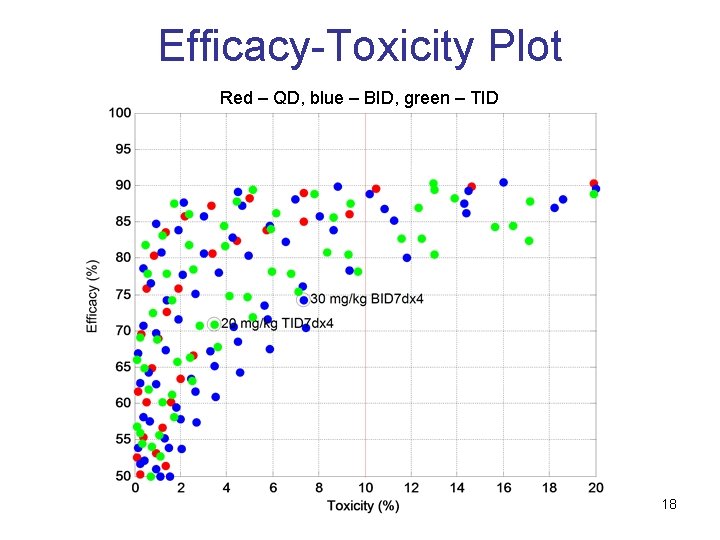
Efficacy-Toxicity Plot Red – QD, blue – BID, green – TID 18

Pareto-Optimal Solutions Red – QD, blue – BID, green – TID 19
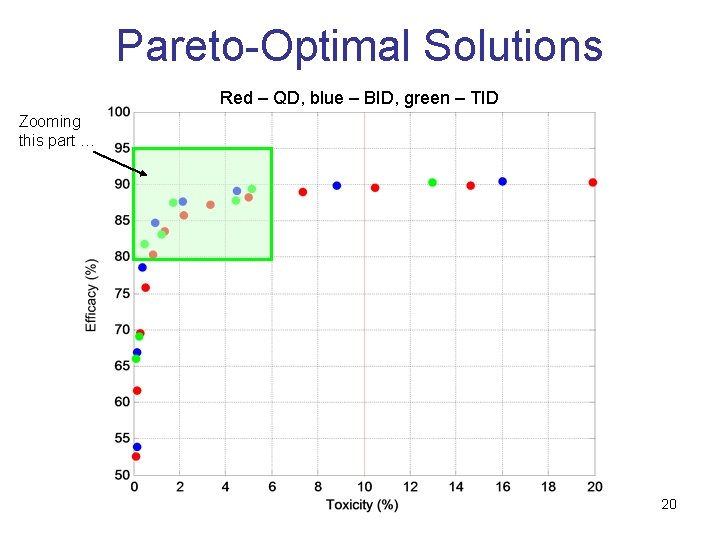
Pareto-Optimal Solutions Red – QD, blue – BID, green – TID Zooming this part … 20
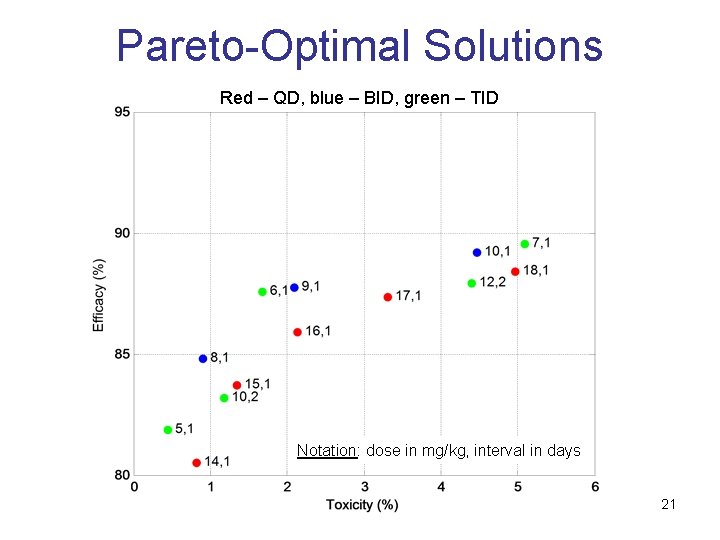
Pareto-Optimal Solutions Red – QD, blue – BID, green – TID Notation: dose in mg/kg, interval in days 21
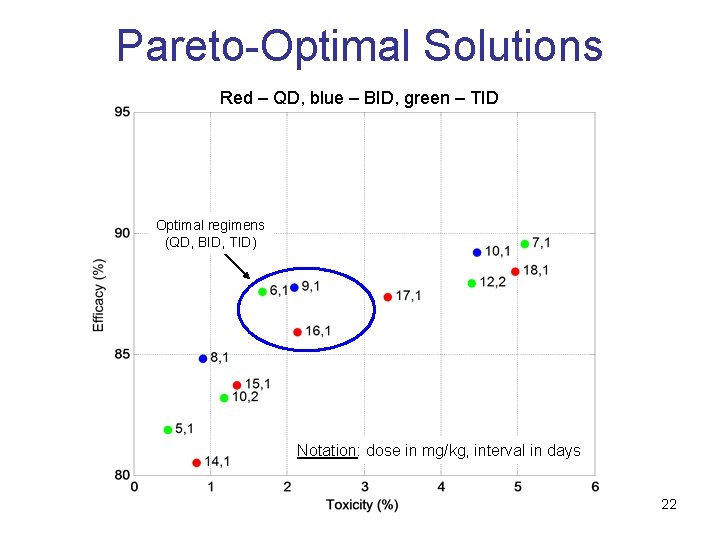
Pareto-Optimal Solutions Red – QD, blue – BID, green – TID Optimal regimens (QD, BID, TID) Notation: dose in mg/kg, interval in days 22
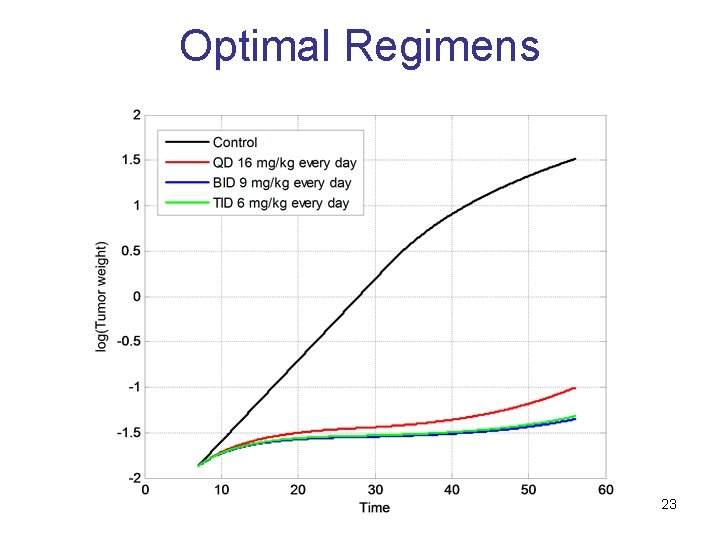
Optimal Regimens 23

Prediction Accuracy Methodology • Fisher’s information matrix computed numerically • Variance-covariance matrix for ML parameter estimates • Simulations performed to quantify prediction uncertainty Regimen: TID 6 mg/kg every day 24

Closer Look at QD Administration Notation: dose in mg/kg, interval in days 25
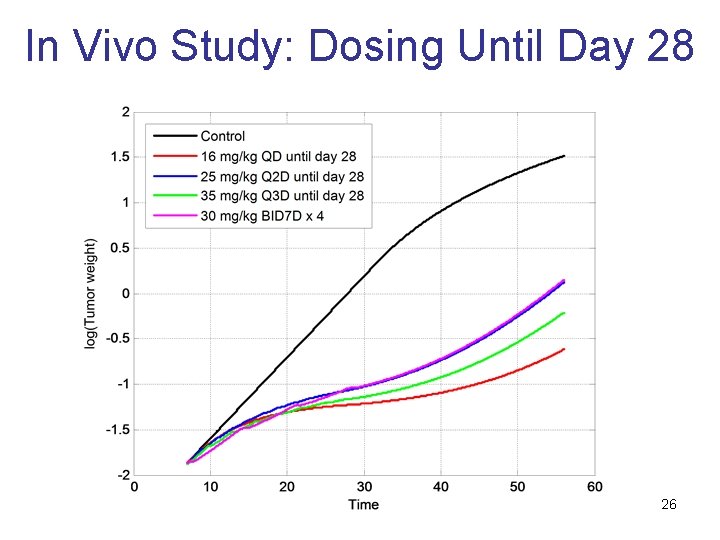
In Vivo Study: Dosing Until Day 28 26

Summary Methodological contribution • New multiobjective method for optimal regimen selection • Novel dynamic model for cancer tumor growth inhibition • Novel dynamic model for animal body weight loss Practical contribution • More efficacious and less toxic in vivo dosing regimens • Better understanding of compound potential pre-clinically Validation • Application of modeling results to in vivo study in progress 27

Acknowledgements Project collaborators • Philip Iversen • Harold Brooks Data generation • Robert Foreman • Charles Spencer 28
- Slides: 28