Math 8803 Discrete Mathematical Biology Mathematical Modeling of

Math 8803 – Discrete Mathematical Biology Mathematical Modeling of Cellular Behavior Ken Dupont Graduate Student (Bio) Mechanical Engineering

Introduction – Tissue Engineering n n n Tissue engineering (TE) aims to create, restore, and/or enhance function of biological tissues through a combination of engineering and biochemical techniques Bone TE Aim: regrow bone that has been lost due to causes such as trauma, congenital defect, or removal due to excision of tumors The basic method of TE is to implant a construct consisting of scaffold +/- cells +/- growth factors PLDL Scaffold, 4 mm D x 8 mm L (R Guldberg, GA Tech) Human mesenchymal stem cells (green) on PLDL scaffold (black struts), 20 X (K Dupont, GA Tech)

Introduction – Tissue Engineering - Cells n n Cells can either be seeded onto scaffolds ex vivo (outside the body) prior to implantation or can be enticed to infiltrate the scaffold in vivo (within the body) Stem cells can both differentiate into other cells and continue to proliferate (divide); mesenchymal stem cells are adult stem cells found in marrow cavities of long bones that can become muscle, cartilage, or bone cells

Introduction – Tissue Engineering – Modeling Mathematical/Computational modeling of cell dynamics has the potential to be a very useful tool in TE Advanced knowledge of the behavior of the cells on constructs could help to optimize TE construct design and limit the number of expensive and time-consuming empirical experiments

Introduction – Processes in TE Constructs Sengers has listed the many of the events happening at the cellular level in TE constructs: n n n n Proliferation – cells divide during mitosis Senescence/Death – cessation of division and later death Motility – cells adhere to and move throughout their environment due to a variety of guiding signals (taxis) Differentiation – stem cells turn into other cell types Nutrient transport/utilization – nutrient concentrations higher outside of constructs than inside, and cellular demands may vary Matrix changes - cells produce extracellular matrix proteins (i. e. collagen) and degradation of matrix may occur as well Cell-cell interactions – Cells can communicate with each other (such as during contact inhibition) NOTE – All of the processes can vary with space and time
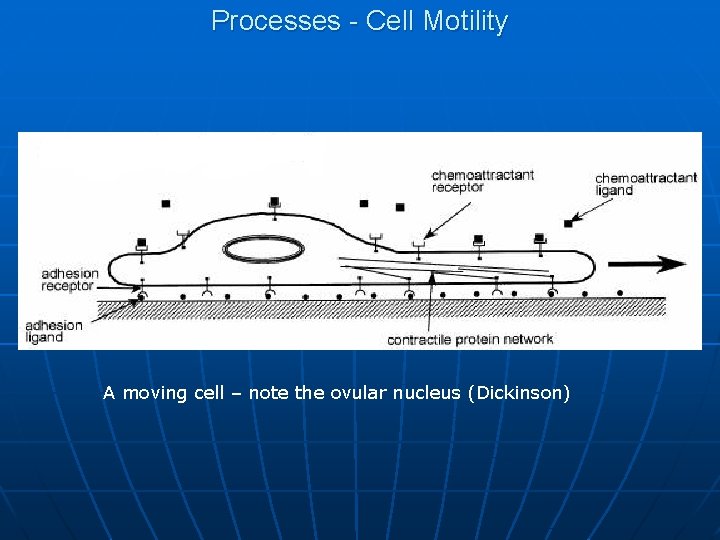
Processes - Cell Motility A moving cell – note the ovular nucleus (Dickinson)

Modeling – Cell Motility – Random Walk Background n Cell motion can be modeled as a random walk • Recall the Bridges of Konigsberg/random walks on graphs from class n n Random walk (RW) - stochastic process made up of a sequence of discrete steps of certain length(s). A random variable can determine the step length and/or walk direction A more formal description of a random walk is as follows: “Let X(t) define a trajectory that begins at position X(0) = X 0. A random walk is modeled by the following expression: X(t + τ) = X(t) + Φ(τ) , where Φ is the random variable that describes the probabilistic rule for taking a subsequent step and τ is the time interval between steps” (Wikipedia)

Modeling – Cell Motility – RW Background n A random walk is an example of a Markov chain, which is a “collection of random variables {Xt} (where the index t runs through 0, 1, …. . ) having the property that, given the present, the future is conditionally independent of the past” (Weisstein):
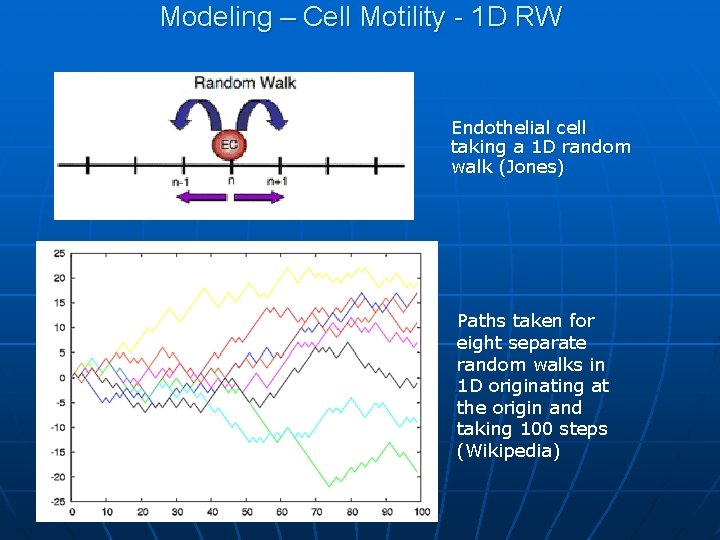
Modeling – Cell Motility - 1 D RW Endothelial cell taking a 1 D random walk (Jones) Paths taken for eight separate random walks in 1 D originating at the origin and taking 100 steps (Wikipedia)
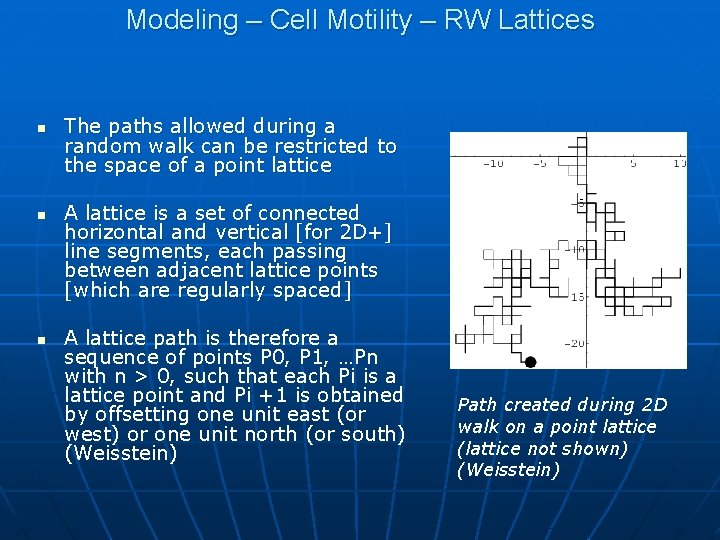
Modeling – Cell Motility – RW Lattices n n n The paths allowed during a random walk can be restricted to the space of a point lattice A lattice is a set of connected horizontal and vertical [for 2 D+] line segments, each passing between adjacent lattice points [which are regularly spaced] A lattice path is therefore a sequence of points P 0, P 1, …Pn with n > 0, such that each Pi is a lattice point and Pi +1 is obtained by offsetting one unit east (or west) or one unit north (or south) (Weisstein) Path created during 2 D walk on a point lattice (lattice not shown) (Weisstein)

Modeling – Cell Motility – RW Lattices Point lattice unit cells are generally in the shape of squares, such that the point lattices are sometimes referred to as grids or meshes n Square lattices helps to minimize memory use and computation times n Cells are far from squares or points, but their position in the mesh can be represented by the location of the cell’s nucleus n Rat mesenchymal stem cells on a 2 D cell culture dish with nuclei stained by Hoechst dye (K Dupont, GA Tech)
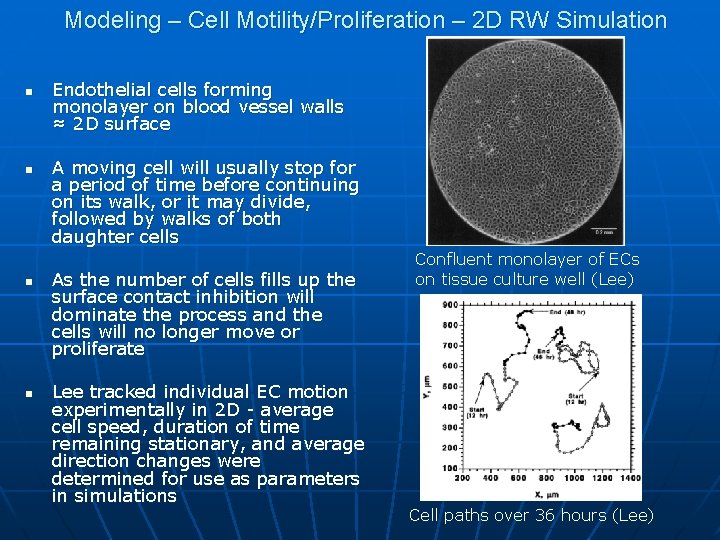
Modeling – Cell Motility/Proliferation – 2 D RW Simulation n n Endothelial cells forming monolayer on blood vessel walls ≈ 2 D surface A moving cell will usually stop for a period of time before continuing on its walk, or it may divide, followed by walks of both daughter cells As the number of cells fills up the surface contact inhibition will dominate the process and the cells will no longer move or proliferate Lee tracked individual EC motion experimentally in 2 D - average cell speed, duration of time remaining stationary, and average direction changes were determined for use as parameters in simulations Confluent monolayer of ECs on tissue culture well (Lee) Cell paths over 36 hours (Lee)

Modeling – Cell Motility/Proliferation – 2 D RW Simulation n Lee used a 2 D discrete cellular automaton model of the proliferation dynamics of populations of migrating cells n Assumed steady state nutrient concentrations and neglected cell loss n n These “discrete systems provide an alternative approach to continuous models that use ordinary and partial differential equation to describe the dynamics of systems evolving in space and time” (Lee) Discrete models can be used to describe movements of individual cells rather than looking at entire populations of cells 2 D lattice of square computational sites • Each site ≈ size of a cell (28 micron sides) • each site has a finite # of possible states and 8 nearest neighbors • The size of the total grid was made to simulate the size of one well of a 96 -well in vitro cell culture plate with diameter of seven millimeters (Jones)

Modeling – Cell Motility/Proliferation – 2 D RW Simulation n n At each time point a lattice site, automaton i, is in a certain state xi (xi = 0 means no cell present) If a cell is present, xi needs to specify if the cell is moving, the direction of locomotion, and the time remaining until a change of direction Time is viewed as discrete steps with uniform increments Δt The state xi of any automaton takes values from the set of 4 -digit integer numbers klmn • k is the direction that the cell is moving in; k can take any value from the set {0, 1, 2, … 8}, with 0 no motion, 1 motion east, 2 motion northeast, etc. . • l is the persistence counter that tells how much time is left until the next change of direction (tc = l * Δt) • mn is the cell phase counter, which tells the amount of time left until the next cell division (tr = (10 m + n) * Δt)

Modeling – Cell Motility/Proliferation – 2 D RW Simulation n Initial cell direction k assigned randomly n Experimental measurements of the cell trajectories were then used to assign initial values of l • The value of the counter decreasing by one after each iteration, with the cell direction changing when the counter reaches zero • The experimental data showed that cells generally change directions in a gradual fashion, so transition probabilities of a cell making a large angle change in direction are small n mn is assigned to each cell, again using the distribution obtained from experimental observations of real cell cycles • 64% of cells divided after 12 -18 h passed, 32% after 18 -24 h passed, and 4% after 24 -30 h passed • mn also decreases by one with each iteration and the cell divides when it reaches zero n l and mn are reset after each direction change and division, respectively

Modeling – Cell Motility/Proliferation – 2 D RW Simulation Example: • Assume a 2 D square lattice with N x N sites, with time step Δt = 0. 5 hours • Choosing an arbitrary automaton site i gives a value of x i = 3319 at to • This means that the site contains a cell moving north for three more iterations (1. 5 h) and that the cell will divide after 19 iterations (9. 5 h) • At time to + Δt, the cell will have moved to site i + N, located one site north of site i, and the value of xi + N = 3218 • The value of xi will then be equal to zero unless another cell moves into the site

Modeling – Cell Motility/Proliferation – 2 D RW Simulation n n n Each simulation run of the model starts by randomly distributing cells at varying densities throughout the 2 D space An algorithm is then begun to increment cell activity at each site with the motion of a cell stopping when it no longer has a free site in which to move If a cell tries to move into an occupied site during one iteration, it will stay in its current location until the next iteration If a cell divides during one iteration it will not move, and one daughter cell will remain in the current site and the other will be randomly assigned to one of the neighbor sites The rows and columns are scanned randomly for incrementation during each iteration to prevent artifacts due to scanning sites in one repeated order CPU time per run lasts between 50 -200 seconds on an IBMRS/6000 POWERStation 350 computer, with time varying based on grid size, initial density of cells, and spatial distribution of cells
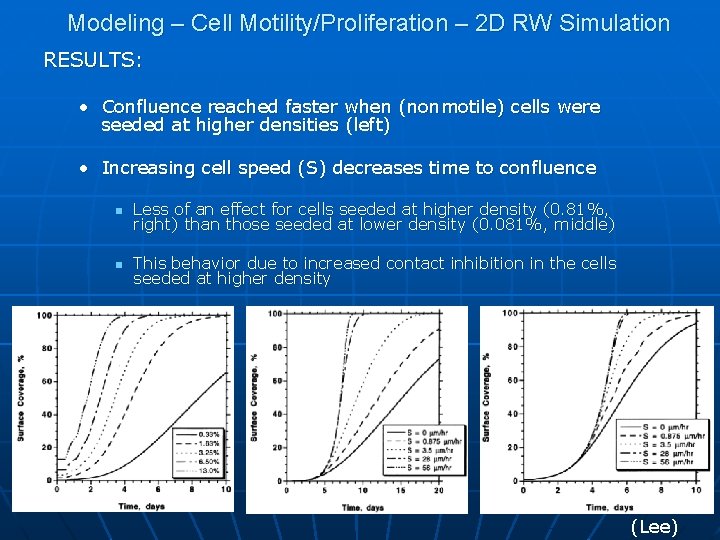
Modeling – Cell Motility/Proliferation – 2 D RW Simulation RESULTS: • Confluence reached faster when (nonmotile) cells were seeded at higher densities (left) • Increasing cell speed (S) decreases time to confluence n Less of an effect for cells seeded at higher density (0. 81%, right) than those seeded at lower density (0. 081%, middle) n This behavior due to increased contact inhibition in the cells seeded at higher density (Lee)
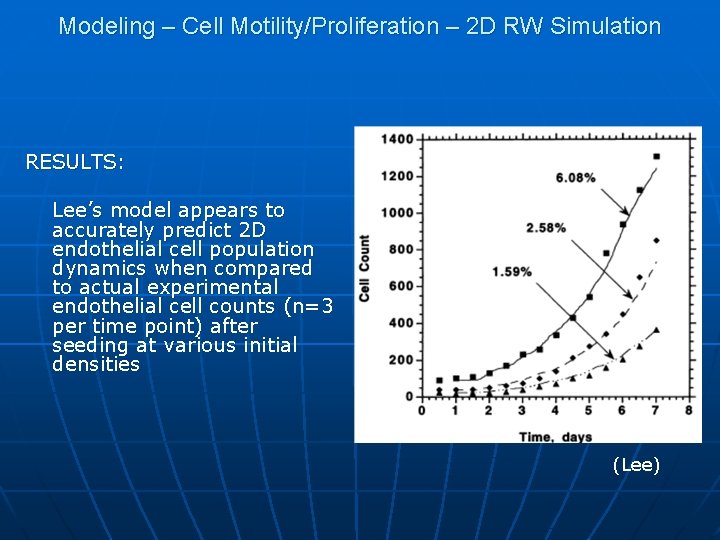
Modeling – Cell Motility/Proliferation – 2 D RW Simulation RESULTS: Lee’s model appears to accurately predict 2 D endothelial cell population dynamics when compared to actual experimental endothelial cell counts (n=3 per time point) after seeding at various initial densities (Lee)

Modeling – Cell Motility/Proliferation – 3 D RW Simulation n Cheng, from the same research group as Lee, investigated application of random walk model of cell motility in 3 D n Assumes highly porous scaffold • Allows unrestricted motion • A cell at one site can move to any of its 6 adjacent cubic faces n n The algorithm for 3 D motion is very similar to that of 2 D motion, again containing a migration index, cell division counter, direction persistence counter, waiting time, and varying transition probabilities to determine the new direction that a cell will move in after stopping, colliding, or dividing One additional feature of the model is that it incorporates a waiting time that a cell will remain stationary after colliding with another cell, which accounts for the tendency of cells to form clusters in 3 D
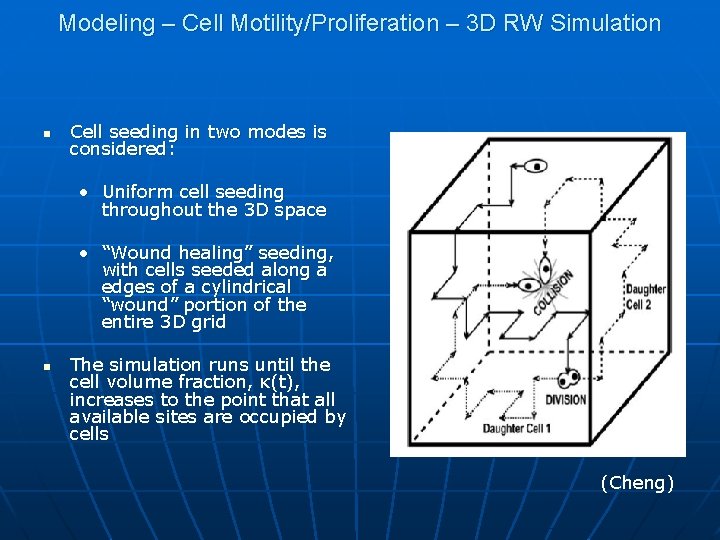
Modeling – Cell Motility/Proliferation – 3 D RW Simulation n Cell seeding in two modes is considered: • Uniform cell seeding throughout the 3 D space • “Wound healing” seeding, with cells seeded along a edges of a cylindrical “wound” portion of the entire 3 D grid n The simulation runs until the cell volume fraction, κ(t), increases to the point that all available sites are occupied by cells (Cheng)
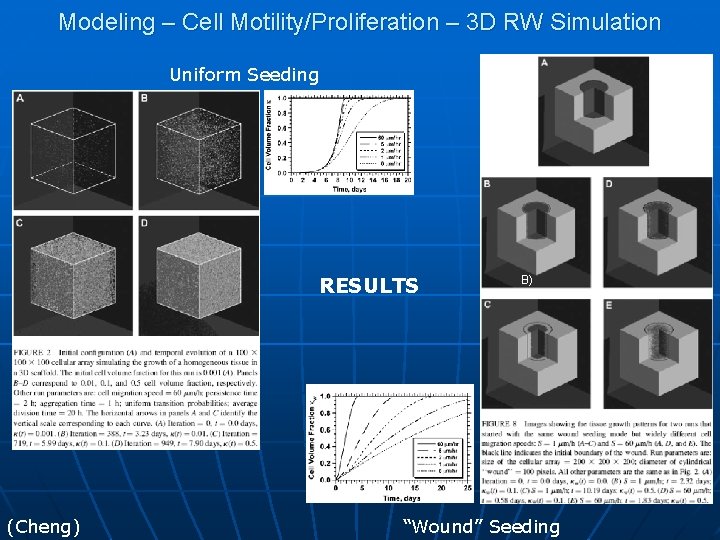
Modeling – Cell Motility/Proliferation – 3 D RW Simulation Uniform Seeding A) RESULTS (Cheng) B) “Wound” Seeding
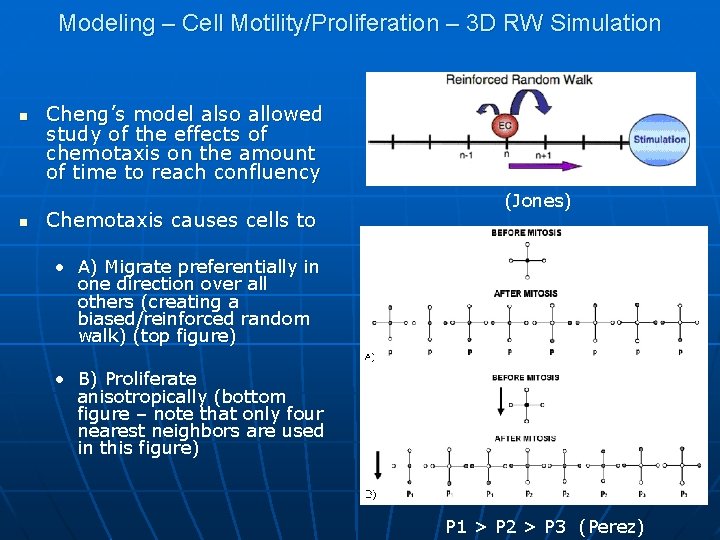
Modeling – Cell Motility/Proliferation – 3 D RW Simulation n n Cheng’s model also allowed study of the effects of chemotaxis on the amount of time to reach confluency Chemotaxis causes cells to (Jones) • A) Migrate preferentially in one direction over all others (creating a biased/reinforced random walk) (top figure) • B) Proliferate anisotropically (bottom figure – note that only four nearest neighbors are used in this figure) P 1 > P 2 > P 3 (Perez)
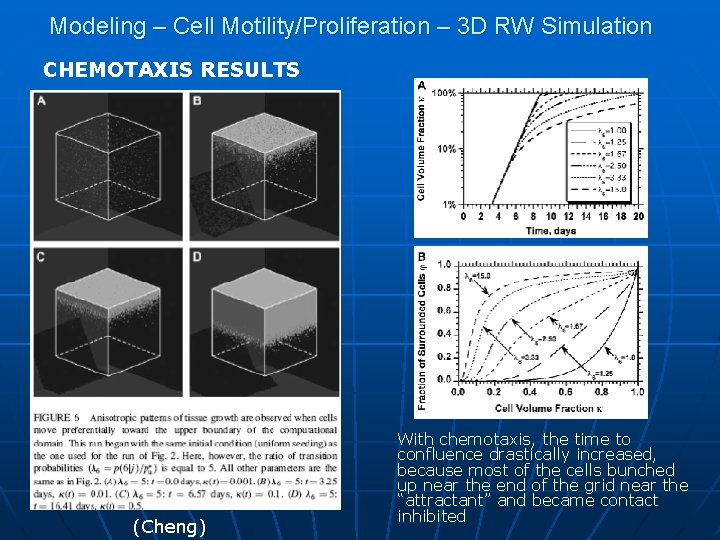
Modeling – Cell Motility/Proliferation – 3 D RW Simulation CHEMOTAXIS RESULTS (Cheng) With chemotaxis, the time to confluence drastically increased, because most of the cells bunched up near the end of the grid near the “attractant” and became contact inhibited

Conclusion n The list of individual phenomena occurring during tissue repair is a long one even without considering the specific spatial and temporal interactions between them Currently, no model can completely describe the tissue growth process, because there are still too many unknowns regarding the process itself Application of discrete models of cell behavior and treatment of cells as individual stochastic objects can be advantageous compared to continuous models because the complex behavior of cells can be broken down into constituent elements • In the words of Jones: “by modeling crucial steps as discrete processes, it is then possible to develop individual areas independently of the rest of the model” n n Caution must be used in applying models to living systems because “theoretical understanding is required as a check on the great risk of error in software and to bridge the enormous gap between computational results and insight or understanding” (Cohen) Until more of the basic biology is known, as well as the math to represent that biology, models will serve as fair predictors for simplified cases of cell dynamics and tissue growth

References n Key Publication References: • • n Biological/Pub. Med only • • • n Byrne DP, Lacroix D, Planell JA, Kelly DJ, Prendergast PJ. Simulation of tissue differentiation in a scaffold as a function of porosity, Young's modulus and dissolution rate: application of mechanobiological models in tissue engineering. Biomaterials. 2007 Dec: 28(36): 5544 -54 Cohen JE. Mathematics Is biology’s next microscope, only better; biology is mathematics’ next physics, only better. PLo. S Biology 2004 Dec: 2(12): e 439. Deasy BM, Jankowski RJ, Payne TR, Cao B, Goff JP, Greenberger JS, Huard J. Modeling stem cell population growth: incorporating terms for proliferative heterogeneity. Stem Cells 2003: 21: 536 -545. Jones PF, Sleeman BD. Angiogenesis - understanding the mathematical challenge. Angiogenesis. 2006: 9(3): 127 -38. Perez MA, Prendergast PJ. Random-walk models of cell dispersal included in mechanobiological simulations of tissue differentiation. Journal of Biomechanics 2007: 40: 2244 -2253. Mathematical/Math. Sci. Net only • • • n Cheng G, Youssef BB, Markenscoff P, Zygourakis K. Cell population dynamics modulate the rates of tissue growth processes. Biophys J. 2006 Feb 1; 90(3): 713 -24. Epub 2005 Nov 18. Lee Y, Kouvroukoglou S, Mc. Intire LV, Zygourakis K. A cellular automaton model for the proliferation of migrating contact-inhibited cells. Biophys J. 1995 Oct; 69(4): 1284 -98. Mac. Arthur BD, Please CP, Taylor M, Oreffo RO. Mathematical modelling of skeletal repair. Biochem Biophys Res Commun. 2004 Jan 23; 313(4): 825 -33. Sengers BG, Taylor M, Please CP, Oreffo RO. Computational modelling of cell spreading and tissue regeneration in porous scaffolds. Biomaterials. 2007 Apr; 28(10): 1926 -40. Epub 2006 Dec 18. Cavalli F, Gamba A, Naldi G, Semplice M. Approximation of 2 D and 3 D models of chemotactic cell movement in vasculogenesis. Math Everywhere: deterministic and stochastic modeling in biomedicine, economics and industry. Springer, Berlin, 2007. Pp. 179 -191. Sherratt JA. Cellular growth control and traveling waves of cancer. SIAM J. Appl. Math. 1993 Dec: 53(6): 1713 -1730. Sleeman BD, Wallis IP. Tumour Induced Angiogenesis as a Reinforced Random Walk: Modelling Capillary Network Formation without Endothelial Cell Proliferation. Mathematical and Computer Modelling. 2002: 36: 339 -358. Jointly Referenced/Other: • • Dickinson RB. A generalized transport model for biased cell migration in an anisotropic environment. . J. Math. Biol. 2000: 40: 97 -135. Perumpanani AJ, Simmons DL, Gearing AJH, Miller KM, Ward G, Norbury J, Schneemann M, Sherratt JA. Extracellular Matrix-Mediated Chemotaxis Can Impede Cell Migration. Proceedings: Biological Sciences 1998 Dec 22: 265(1413) 2347 -2352. “Random Walk”. Wikipedia. 6 April 2008. http: //en. wikipedia. org/wiki/Random_walk Weisstein, EW. “Random Walk”. From Math. World – A Wolfram Web Resource. http: //mathworld. wolfram. com/Random. Walk. html
- Slides: 26