Master Seminar Deep Learning for Medical Applications Unsupervised









- Slides: 21

Master Seminar: Deep Learning for Medical Applications Unsupervised domain adaptation for medical imaging segmentation with self-ensembling Perone CS, Ballester B, Barros RC, Cohen-Adad J. Neuro. Image, 2019 05. 12. 2019 Christoph Clement Supervisor: Roger Soberanis

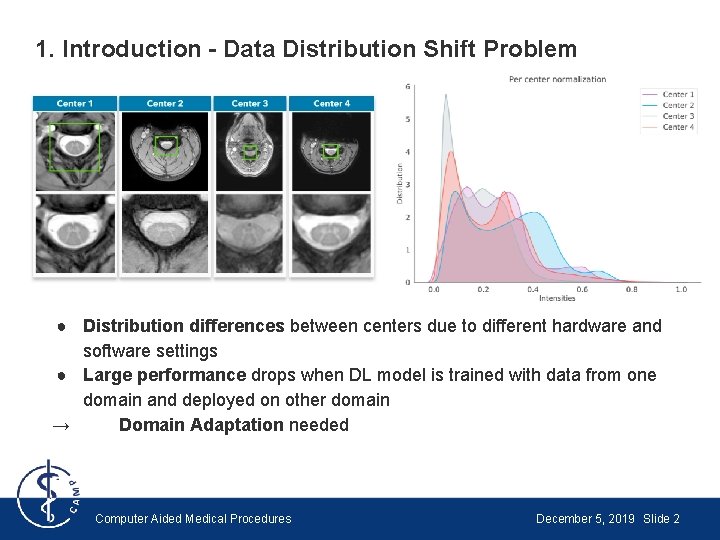
1. Introduction - Data Distribution Shift Problem ● Distribution differences between centers due to different hardware and software settings ● Large performance drops when DL model is trained with data from one domain and deployed on other domain → Domain Adaptation needed Computer Aided Medical Procedures December 5, 2019 Slide 2

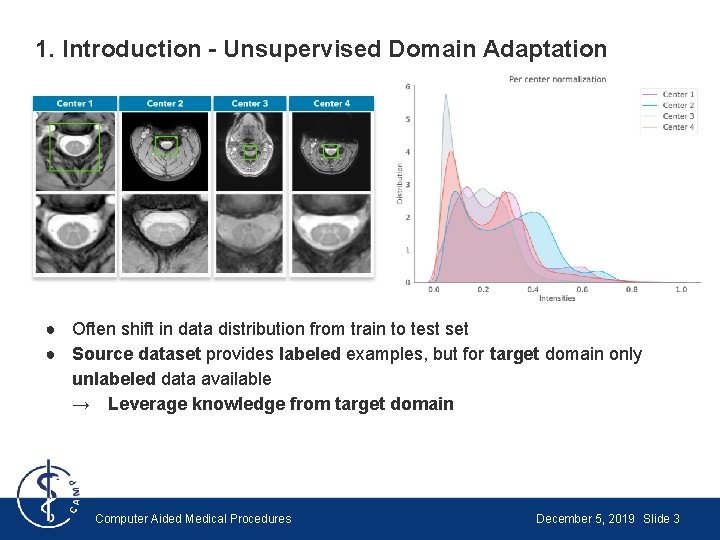
1. Introduction - Unsupervised Domain Adaptation ● Often shift in data distribution from train to test set ● Source dataset provides labeled examples, but for target domain only unlabeled data available → Leverage knowledge from target domain Computer Aided Medical Procedures December 5, 2019 Slide 3

1. Introduction - Main Contributions 1. Extend unsupervised domain adaptation to semantic segmentation 1. Evaluate performance using a realistic small MRI dataset [1] 1. Perform ablation experiment to show that unlabeled data is responsible for performance improvement 1. Visually analyze of how domain adaptation affects prediction space [1] Prados F, Ashburner J, Blaiotta C, Brosch T, Carballido-Gamio J, Cardoso MJ. Spinal cord grey matter segmentation challenge, Neuroimage, 2017 Computer Aided Medical Procedures December 5, 2019 Slide 4

1. Introduction - Related Work ● U-net by Ronneberger et al. , 2015 ● Deep Domain Adaptation (DDA) ○ GANs to build domain-invariant feature spaces [1] ○ Optimizing higher-order statistics to change parameters of neural network layers [2] ○ Explicit minimization of discrepancy between source and target domains [3] ○ Self-ensembling methods based on the Mean Teacher network [4] [1] Hoffman J, Tzeng E, Park T, Zhu JY, Isola, P, Saenko, K. Cycada: Cycle-Consistent Adversarial Domain Adaptation, ar. Xiv preprint, 2017 [2] Li Y, Wang N, Shi J, Liu J, Hou X. Revisiting Batch Normalization for Practical Domain Adaptation, ar. Xiv preprint, 2016 [3] Tzeng E, Ho_man J, Zhang N, Saenko K, Darrell T. Deep Domain Confusion: Maximizing for Domain Invariance, ar. Xiv preprint, 2014 [4] Tarvainen A, Valpola H. Mean teachers are better role models: weight-averaged consistency targets improve semi-supervised deep learning results, Advances in Neural Information Processing Systems, 2017 Computer Aided Medical Procedures December 5, 2019 Slide 5

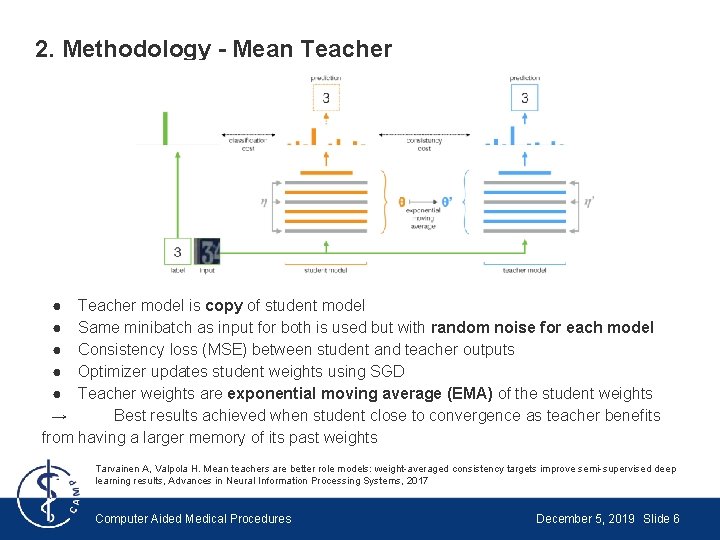
2. Methodology - Mean Teacher ● Teacher model is copy of student model ● Same minibatch as input for both is used but with random noise for each model ● Consistency loss (MSE) between student and teacher outputs ● Optimizer updates student weights using SGD ● Teacher weights are exponential moving average (EMA) of the student weights → Best results achieved when student close to convergence as teacher benefits from having a larger memory of its past weights Tarvainen A, Valpola H. Mean teachers are better role models: weight-averaged consistency targets improve semi-supervised deep learning results, Advances in Neural Information Processing Systems, 2017 Computer Aided Medical Procedures December 5, 2019 Slide 6

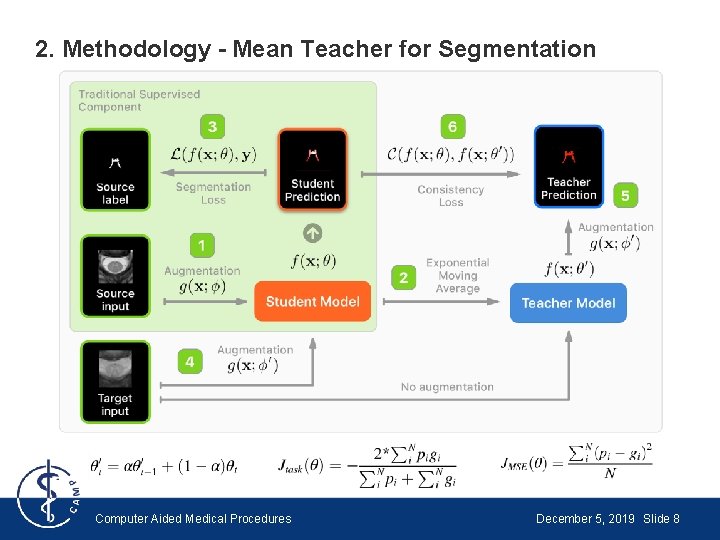
2. Methodology - Mean Teacher for Segmentation ● Use Dice Loss instead of Classification Cost ● Use random spatial transformation instead of random noise ● Spatial transformation applied to inputs of student model and outputs of teacher model → Solves problem of inconsistency when random spatial transformation applied to both inputs of the teacher and student models Computer Aided Medical Procedures December 5, 2019 Slide 7

2. Methodology - Mean Teacher for Segmentation Computer Aided Medical Procedures December 5, 2019 Slide 8

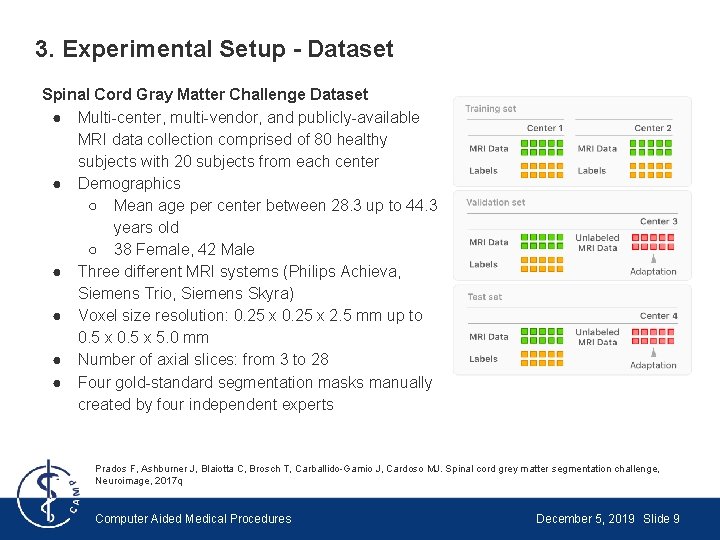
3. Experimental Setup - Dataset Spinal Cord Gray Matter Challenge Dataset ● Multi-center, multi-vendor, and publicly-available MRI data collection comprised of 80 healthy subjects with 20 subjects from each center ● Demographics ○ Mean age per center between 28. 3 up to 44. 3 years old ○ 38 Female, 42 Male ● Three different MRI systems (Philips Achieva, Siemens Trio, Siemens Skyra) ● Voxel size resolution: 0. 25 x 2. 5 mm up to 0. 5 x 5. 0 mm ● Number of axial slices: from 3 to 28 ● Four gold-standard segmentation masks manually created by four independent experts Prados F, Ashburner J, Blaiotta C, Brosch T, Carballido-Gamio J, Cardoso MJ. Spinal cord grey matter segmentation challenge, Neuroimage, 2017 q Computer Aided Medical Procedures December 5, 2019 Slide 9

3. Experimental Setup - Baseline ● U-Net architecture and training hyper-parameters for baseline same as for teacher and student models ● Group norm instead of Batch norm used ● Difference between baseline and proposed method: train baseline model in standard supervised learning fashion with no additional unlabeled data Computer Aided Medical Procedures December 5, 2019 Slide 10

3. Experimental Setup - Training Setup ● Extensive hyperparameter search ○ Mini-batch size: 12 ○ Dropout rate: 0. 5 ○ Adam optimizer ○ Learning rate: sigmoid learning rate ramp-up strategy until epoch 50 followed by a cosine ramp-down until epoch 350 ● Trained network with centers 1 and 2 in a supervised fashion → adapted to centers 3 and 4 separately Computer Aided Medical Procedures December 5, 2019 Slide 11

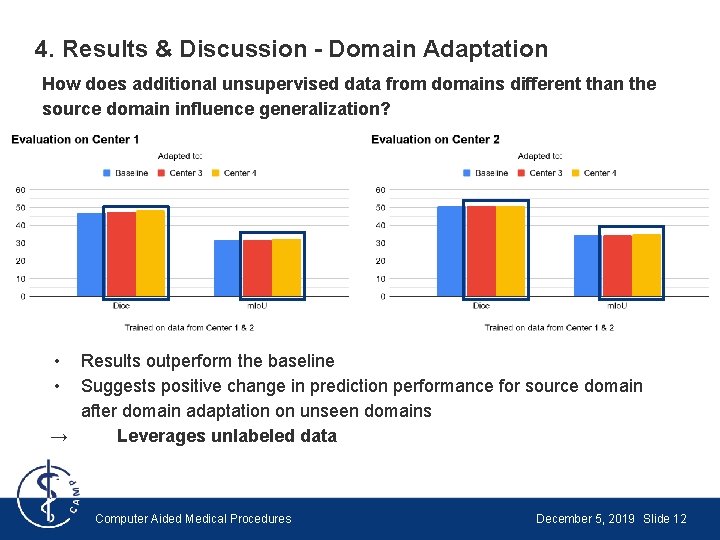
4. Results & Discussion - Domain Adaptation How does additional unsupervised data from domains different than the source domain influence generalization? • • Results outperform the baseline Suggests positive change in prediction performance for source domain after domain adaptation on unseen domains → Leverages unlabeled data Computer Aided Medical Procedures December 5, 2019 Slide 12

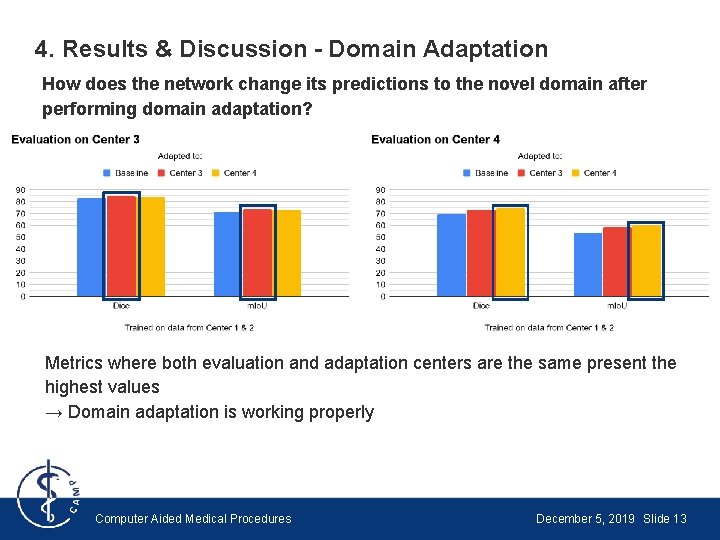
4. Results & Discussion - Domain Adaptation How does the network change its predictions to the novel domain after performing domain adaptation? Metrics where both evaluation and adaptation centers are the same present the highest values → Domain adaptation is working properly Computer Aided Medical Procedures December 5, 2019 Slide 13

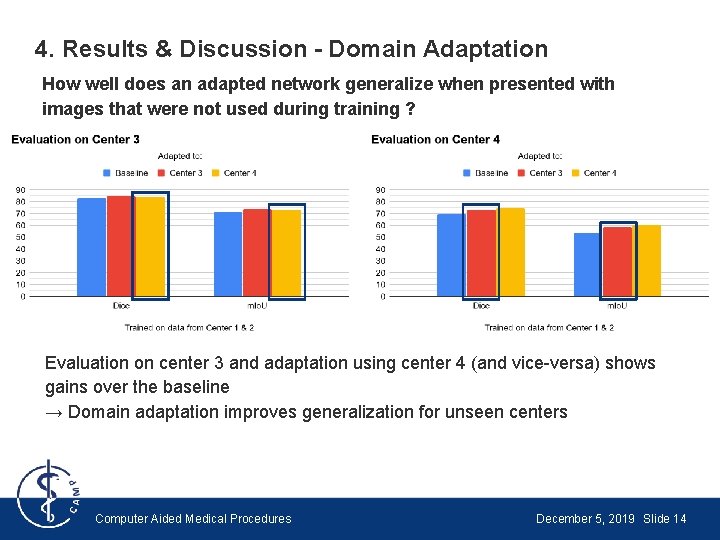
4. Results & Discussion - Domain Adaptation How well does an adapted network generalize when presented with images that were not used during training ? Evaluation on center 3 and adaptation using center 4 (and vice-versa) shows gains over the baseline → Domain adaptation improves generalization for unseen centers Computer Aided Medical Procedures December 5, 2019 Slide 14

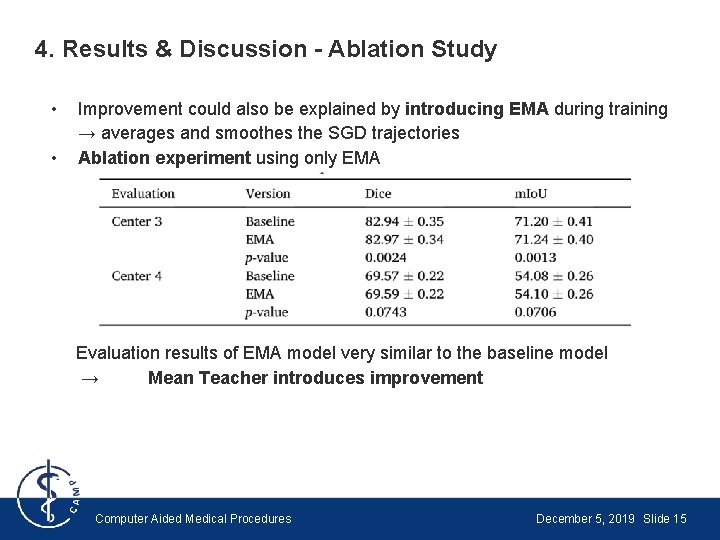
4. Results & Discussion - Ablation Study • • Improvement could also be explained by introducing EMA during training → averages and smoothes the SGD trajectories Ablation experiment using only EMA Evaluation results of EMA model very similar to the baseline model → Mean Teacher introduces improvement Computer Aided Medical Procedures December 5, 2019 Slide 15

5. Conclusion • Unsupervised domain adaptation is an effective way to increase performance of ML models across multiple centers • Self-ensembling methods improve generalization on unseen domains through the leverage of unlabeled data from multiple domains • Improvements come through introduction of unlabeled data and not only through exponential moving average Computer Aided Medical Procedures December 5, 2019 Slide 16

5. Conclusion - Limitations & Future Work Limitations • No evaluation of adversarial training methods for domain adaptation as comparison • Single task evaluation of gray matter segmentation • Limited amount of centers Future Work • Reassess importance of proper multidomain evaluation in studies and medical imaging challenges → rarely provide test set from different domains • Try on different datasets Computer Aided Medical Procedures December 5, 2019 Slide 17

5. Conclusion - Own View Done well: • In general, understandably and well written • Ensured fair comparison between baseline and proposed method • Repeated experiments over 10 runs • Open sourced their code Could be improved: • Explanation of Mean Teacher could be more detailed • Deeper analysis of their method would be nice Computer Aided Medical Procedures December 5, 2019 Slide 18

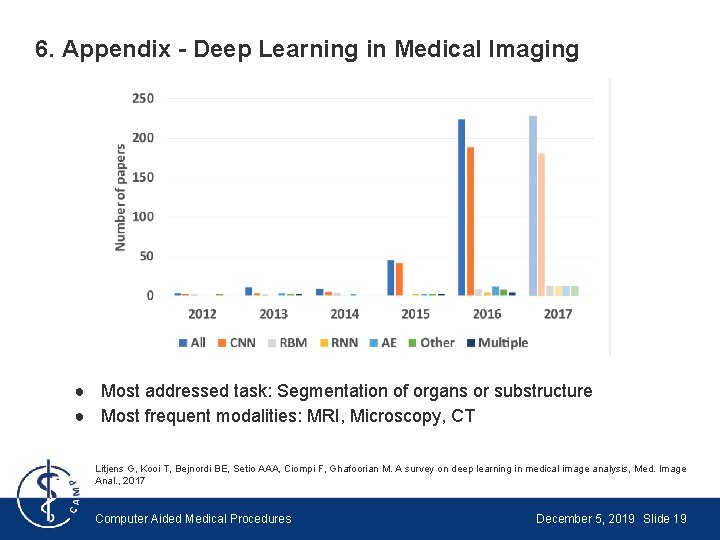
6. Appendix - Deep Learning in Medical Imaging ● Most addressed task: Segmentation of organs or substructure ● Most frequent modalities: MRI, Microscopy, CT Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M. A survey on deep learning in medical image analysis, Med. Image Anal. , 2017 Computer Aided Medical Procedures December 5, 2019 Slide 19

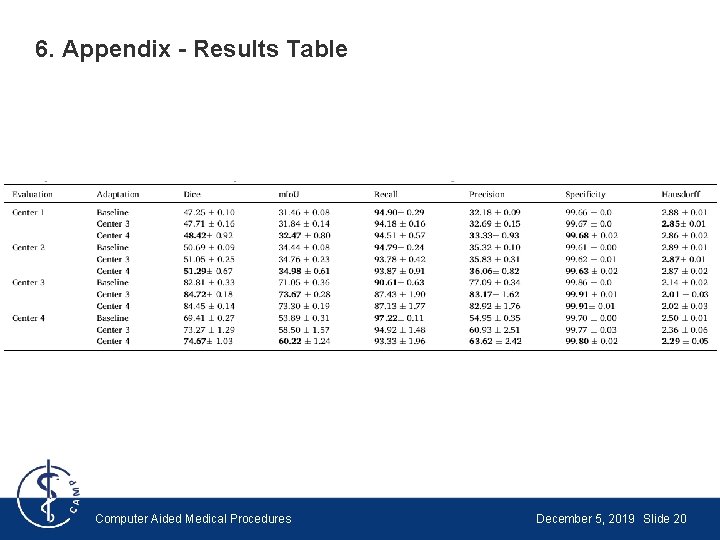
6. Appendix - Results Table Computer Aided Medical Procedures December 5, 2019 Slide 20

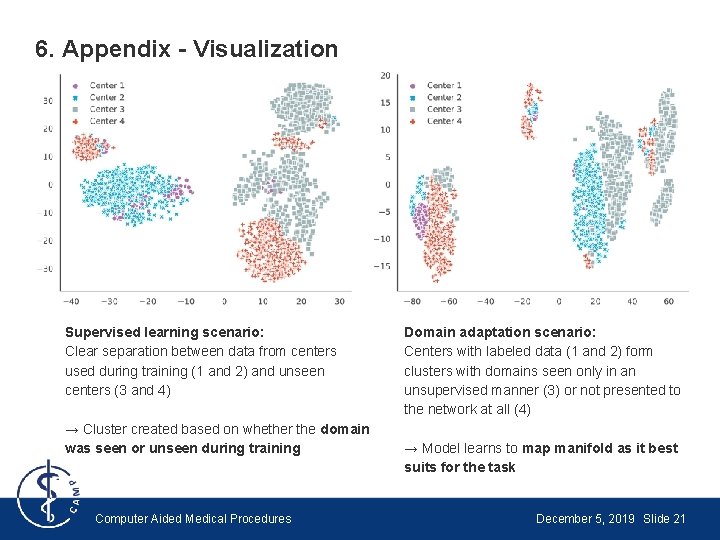
6. Appendix - Visualization Supervised learning scenario: Clear separation between data from centers used during training (1 and 2) and unseen centers (3 and 4) → Cluster created based on whether the domain was seen or unseen during training Computer Aided Medical Procedures Domain adaptation scenario: Centers with labeled data (1 and 2) form clusters with domains seen only in an unsupervised manner (3) or not presented to the network at all (4) → Model learns to map manifold as it best suits for the task December 5, 2019 Slide 21